Excited to share the new Steering Committee of PAASTA 🎉
An ECR-driven community working to foster an open, inclusive, and collaborative space for palaeoproteomics researchers worldwide.
Looking forward to building and growing this community together 🧬
30.01.2026 17:54 —
👍 17
🔁 9
💬 0
📌 0
Have fun Zandra 🥳
12.12.2025 12:17 —
👍 1
🔁 0
💬 0
📌 0

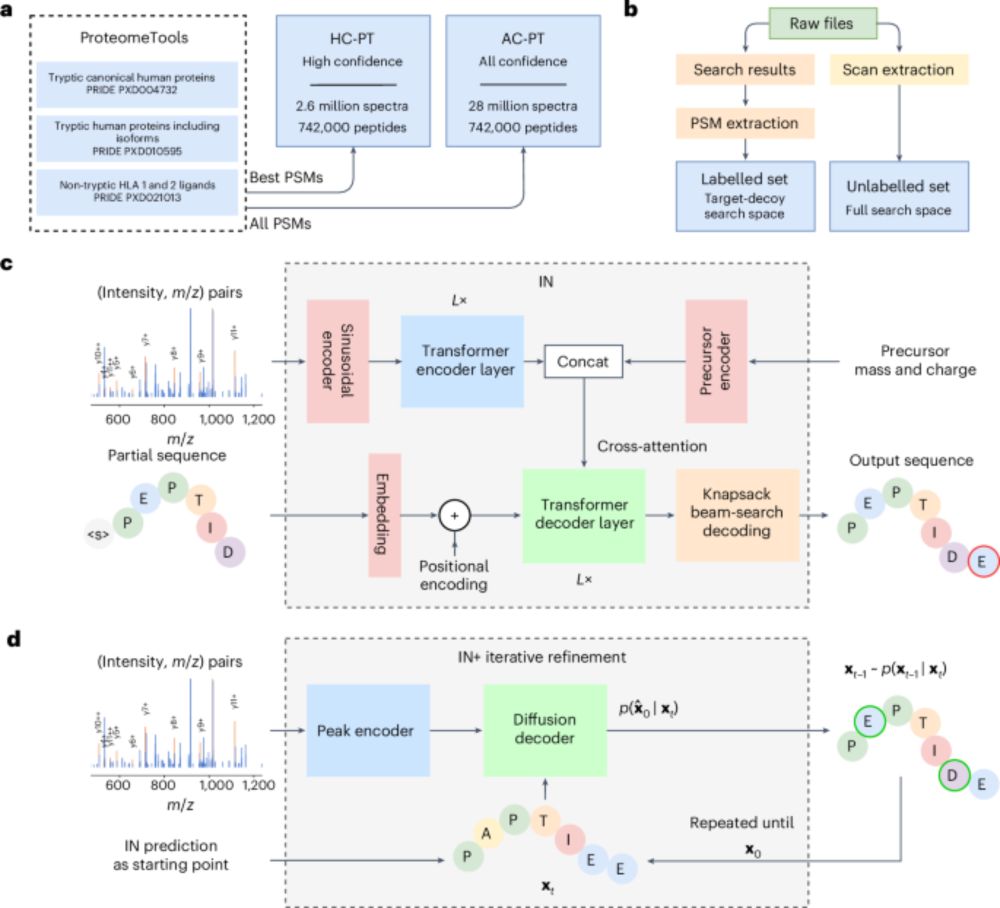
Foundation model for mass spectrometry proteomics
Mass spectrometry is the dominant technology in the field of proteomics, enabling high-throughput analysis of the protein content of complex biological samples. Due to the complexity of the instrument...
Interested in prediction tasks involving peptide mass spectra? Our foundation model uses pre-trained spectrum representations learned by a de novo sequencing model to solve many tasks better and with less data, from recognizing chimeras to separating N- and O-glycopeptides. arxiv.org/abs/2505.10848
22.05.2025 11:49 —
👍 12
🔁 7
💬 0
📌 1

SHARK: web server for alignment-free homology assessment for intrinsically disordered and unalignable protein regions academic.oup.com/nar...
---
#proteomics #prot-paper
21.05.2025 19:20 —
👍 2
🔁 1
💬 0
📌 0

Exciting news: We have released #FragPipe 23, and it's one of our biggest updates ever. Windows installer, support for TMT on Astral and timsTOF, TMT35, PTM site reports for DIA, improved Astral data handling in #MSFragger, improved diaTracer for diaPASEF data, better Skyline integration, and more!
05.05.2025 04:06 —
👍 80
🔁 15
💬 4
📌 1
I'm so happy to see this paper out, as it has a special place in my heart! A weird, silly idea that worked out brilliantly. If you want a serious thread about the results, read the one by @welkergroup.bsky.social. I will instead provide you with a thread about involving your dog in your research:
24.04.2025 13:16 —
👍 22
🔁 7
💬 1
📌 0

Fast and memory efficient searching of large-scale mass spectrometry data using Tide www.biorxiv.org/cont...
---
#proteomics #prot-preprint
07.04.2025 16:01 —
👍 6
🔁 3
💬 0
📌 1

Laboratory Manager Codicum Project
1 of 4 We're hiring a Laboratory Manager for the CODICUM project at @UniCopenhagen to help explore medieval manuscript fragments using cutting-edge biomolecular methods!
Apply by April 13, 2025!
Link: employment.ku.dk/staff?show=1...
#LabJobs #BiomolecularScience #AncientDNA
👇
06.04.2025 20:38 —
👍 36
🔁 33
💬 1
📌 1
Google colab has free GPUs (T4). Instanovo now has a Hugging Face space for online predictions! CPUs still work for Casa & Insta (just slower🐢)
03.04.2025 07:44 —
👍 3
🔁 1
💬 1
📌 0
Chat platform · ISBA
In following suit with @isbarchaeology.bsky.social, we have moved from our former chat platform home of Slack over to Element. More details *and* how to make the switch can be found here: www.isbarch.org/chat and from the ISBA space, you can readily find us (paasta:archaeo.social)!
01.04.2025 13:14 —
👍 3
🔁 4
💬 0
📌 1
Im teaching here about DIANN (DIA analysis), OpenMS and quantms analysis for DDA. We have lectures about MaxQuant, PeptideShaker, ProteomeXchange, data reuse, standards, @pride-ebi.bsky.social One week fully packed with lectures about the fundamentals of computational proteomics.
27.03.2025 12:57 —
👍 19
🔁 8
💬 0
📌 0
“We confirmed the workflow its exceptional stability through analysis of 1,500 quality control samples systematically interspersed among 36,000 plasma samples measured continuously over 311 days.” 🤯 🫠
27.03.2025 09:20 —
👍 7
🔁 2
💬 0
📌 0
Dino!!! Yay time to go to colab🙈
25.03.2025 06:44 —
👍 0
🔁 0
💬 0
📌 0

These are some examples of #tidyplots built-in color schemes 🌈
Feel free to create your own!
#rstats #dataviz #phd
17.03.2025 20:29 —
👍 16
🔁 4
💬 1
📌 1
Cheers @matthewcollins.bsky.social 🎉 It’s been fun and thanks for the custom Lego mini with an Orthrus script jumper 🐾
16.03.2025 19:56 —
👍 4
🔁 0
💬 0
📌 1
Thread 5/5: 🧬 Amber's preservation capabilities for proteins remain debated.
While promising, paleoproteomics requires exceptional methodology to distinguish genuine ancient proteins from contamination. Let's wait for peer review and independent verification. #paleoproteomics #scienceskepticism 🙂
07.03.2025 07:47 —
👍 11
🔁 1
💬 0
📌 0
If you're about to order an Exploris 480, Astral, or even an EasySpray source, I would either:
A) Wait for the clearance deals in the coming 6 weeks and take advantage of the deals
B) Wait until after ASMS and then update the order
06.03.2025 08:51 —
👍 14
🔁 1
💬 1
📌 0
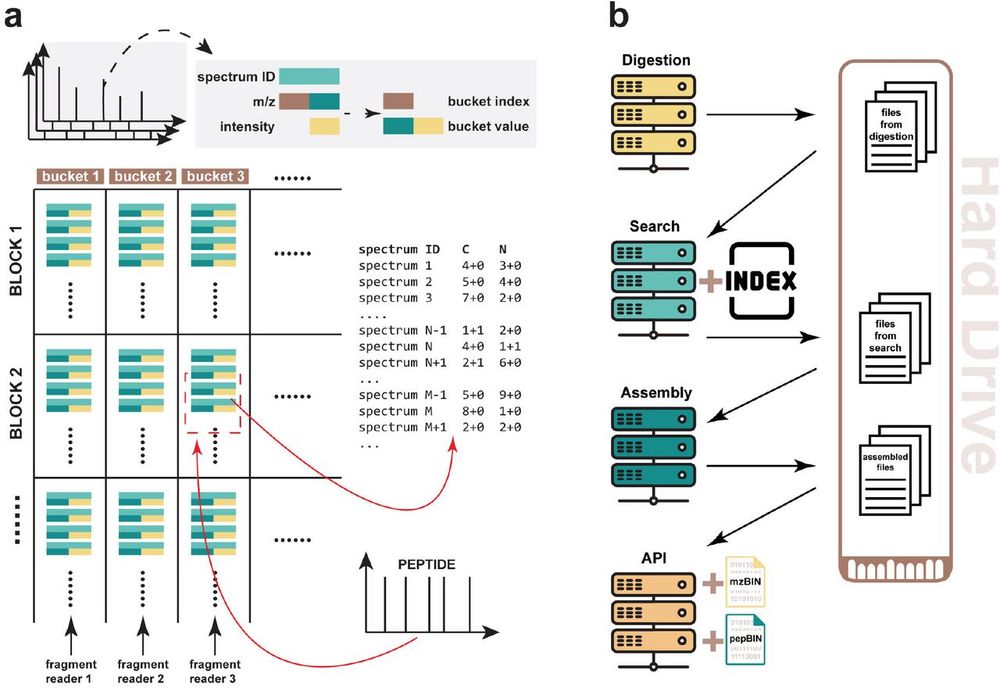
One of the most significant and challenging projects of my career so far. PepCentric: a scalable computational platform utilizing novel 2-D fragment indexing for rapid peptide-centric searches, enabling proteogenomics searches against billions of spectra in seconds. www.biorxiv.org/content/10.1...
04.03.2025 09:09 —
👍 72
🔁 23
💬 3
📌 2

New #tidyplots cheatsheet 🤩
tidyplots.org/cheatsheet
#rstats #dataviz #phd
03.03.2025 17:30 —
👍 38
🔁 8
💬 0
📌 1
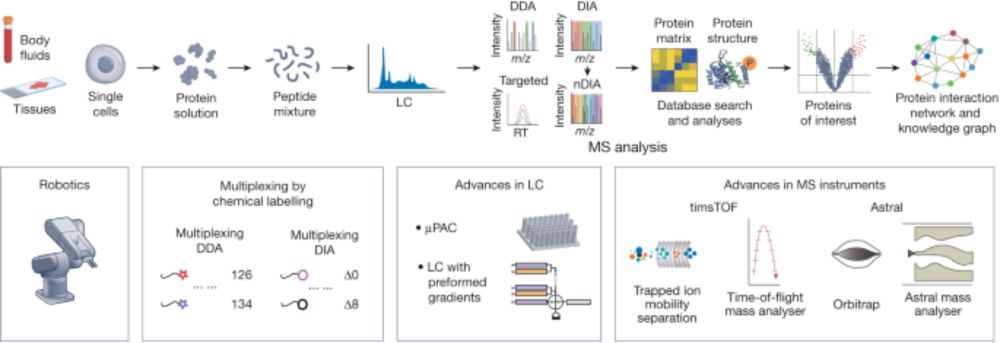
Mass-spectrometry-based proteomics: from single cells to clinical applications - Nature
This Review summarizes advances in mass-spectrometry-based proteomics and explores the potential applications of these technologies in the clinic.
In our latest @Nature review with Tiannan Guo & Judith Steen, we explore how technological breakthroughs are revolutionizing MS-based proteomics: From enhanced sensitivity enabling single-cell analysis to high-throughput plasma proteomics & AI-based data interpretation
www.nature.com/articles/s41...
26.02.2025 16:22 —
👍 14
🔁 9
💬 1
📌 0

Protein degradation and growth dependent dilution substantially shape mammalian proteomes
Cellular protein concentrations are maintained through a balance of synthesis and clearance. Clearance occurs through both protein degradation and growth-dependent dilution. At slow growth, clearance ...
Our study reconfigures our understanding of the mechanism cells use regulate gene expression.
What was previously reported to have little global impact now is reported to explain up to half of the variation in how protein levels are controlled. Protein degradation matters!
shorturl.at/5wPiS
25.02.2025 17:44 —
👍 5
🔁 3
💬 0
📌 0
Yeah have been banging my head against the wall trying to ID some ancient complex mixtures with limited peps and bad ion coverage…maybe time to go back to the drawing board and write more codes 🙈💻☠️
17.02.2025 18:50 —
👍 2
🔁 0
💬 0
📌 0