I can tell Huabio is a real company!😀
04.03.2026 07:09 — 👍 0 🔁 0 💬 0 📌 0I can tell Huabio is a real company!😀
04.03.2026 07:09 — 👍 0 🔁 0 💬 0 📌 0Thanks! I see, then anti-NRXN3α is impossible to detect NRXN3β, but anti-NRXN3β has a possibility to detect NRXN3α.
04.03.2026 07:08 — 👍 0 🔁 0 💬 1 📌 0Great tools! For anti-NRXN3a/b antibodies, does they recognize each other? I saw the test data, and most likely a does not cross with NRXNa families, and b does not cross with NRXNb families. It would be great to know does anti-NRXN3a antibody detect NRXN3b and vice versa.
02.03.2026 02:24 — 👍 1 🔁 0 💬 1 📌 0Saw a familiar face. Congratulations Rohith!
02.03.2026 02:14 — 👍 0 🔁 0 💬 0 📌 0
Check out our newest work! This is a story on how to get selectivity in binders - both isoform and site selectivity. Read the paper or enjoy this brief Skytorial of what we did!
www.biorxiv.org/content/10.1...
1/n
How cool it is!!
19.11.2025 01:19 — 👍 0 🔁 0 💬 1 📌 0How do cells adapt morphology to function? In a 🔥 preprint by @zjmaggiexu.bsky.social , with @dudinlab.bsky.social and @amyweeks.bsky.social , we identify a self-organizing single-cell morphology circuit that optimizes the feeding trap structure of the suctorian P. collini. 🧵 tinyurl.com/4k8nv926
18.11.2025 16:15 — 👍 131 🔁 55 💬 4 📌 11
A parts list of promoters and gRNA scaffolds for mammalian genome engineering and molecular recording - @jshendure.bsky.social @troymcdiarmid.bsky.social @uwgenome.bsky.social go.nature.com/49eTPCu
11.11.2025 16:36 — 👍 28 🔁 12 💬 0 📌 2Fun to see this thread answer some questions I had, thank you.
01.10.2025 04:55 — 👍 2 🔁 0 💬 0 📌 0Cool! This could have a lot of influence on sensors, pull down, signaling rewiring...🤔
01.10.2025 04:43 — 👍 1 🔁 0 💬 0 📌 0I see👍 thank you!
27.09.2025 05:21 — 👍 1 🔁 0 💬 0 📌 0I would like to hear some suggestions on why you think we should not use it?
26.09.2025 12:33 — 👍 0 🔁 0 💬 1 📌 0
🚀 Our new paper is out @natmethods.nature.com!
Kuffer & Marzilli engineered conditionally stable MS2 & PP7 coat proteins (dMCP & dPCP) that degrade unless bound to RNA, enabling ultra–low-background, single-mRNA imaging in live cells.
🔗 www.nature.com/articles/s41...
🧬 www.addgene.org/John_Ngo/
Thank you, looks very cool! Can I message you about the cell marker you are using here?
25.08.2025 03:00 — 👍 1 🔁 0 💬 1 📌 0Very cool! What kind of cell types are used here, and what kind of microscope is used? How long does the imaging take?
22.08.2025 01:40 — 👍 1 🔁 0 💬 1 📌 0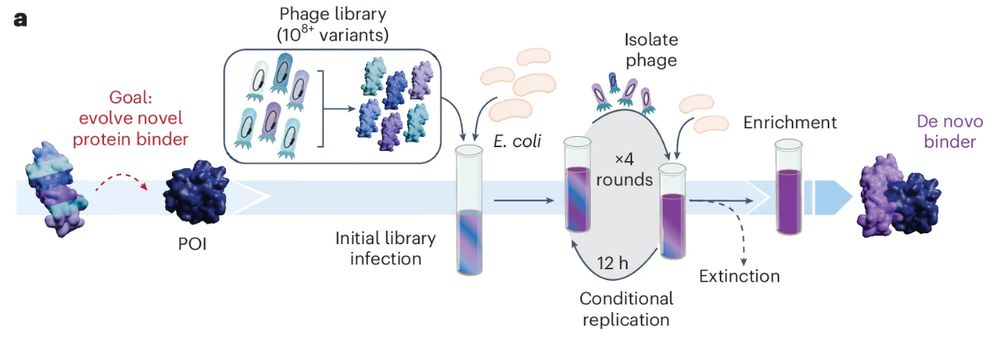
PANCS-Binders (phage-assisted noncontinuous selection of protein binders) screens multiple high-diversity protein libraries against a panel of dozens of targets for high-throughput binder discovery. @chembiobryan.bsky.social @mstyles-chembiol.bsky.social
www.nature.com/articles/s41...
That was so cool!
19.06.2025 00:41 — 👍 1 🔁 0 💬 0 📌 0Humanized Caffeine-Inducible Systems for Controlling Cellular Functions https://www.biorxiv.org/content/10.1101/2025.06.13.659463v1
14.06.2025 05:04 — 👍 7 🔁 5 💬 0 📌 0That was cool👍
13.06.2025 09:58 — 👍 1 🔁 0 💬 0 📌 0Cool, intresting findings, could be invented as tool for transcription control?
12.06.2025 12:43 — 👍 0 🔁 0 💬 0 📌 0Very cool study!
05.06.2025 01:53 — 👍 1 🔁 0 💬 0 📌 0I have a lot of recommendations if you are in China😀
01.06.2025 07:26 — 👍 1 🔁 0 💬 0 📌 0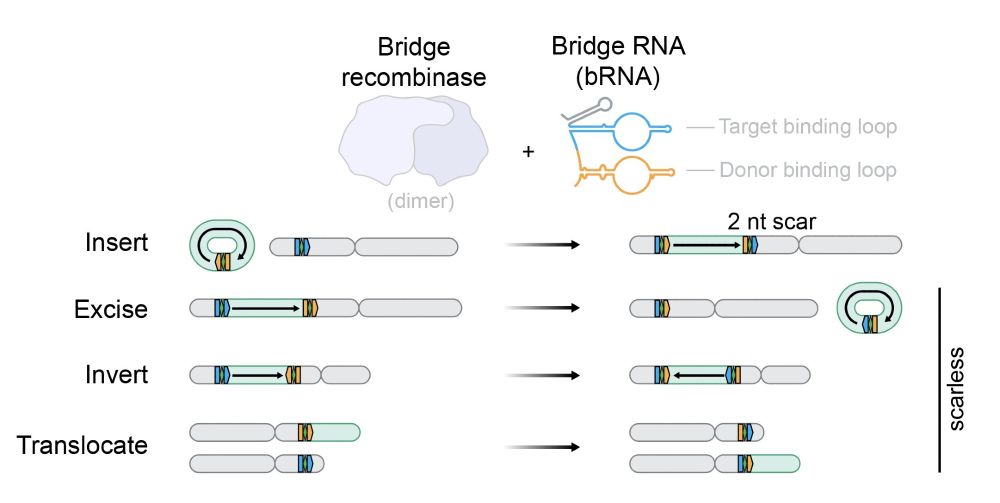
Genomes encode biological complexity, which is determined by combinations of DNA mutations across millions of bases
In new work @arcinstitute.org, we report the discovery and engineering of the first programmable DNA recombinases capable of megabase-scale human genome rearrangement
Congrats! Useful tools!
24.04.2025 01:47 — 👍 5 🔁 0 💬 0 📌 0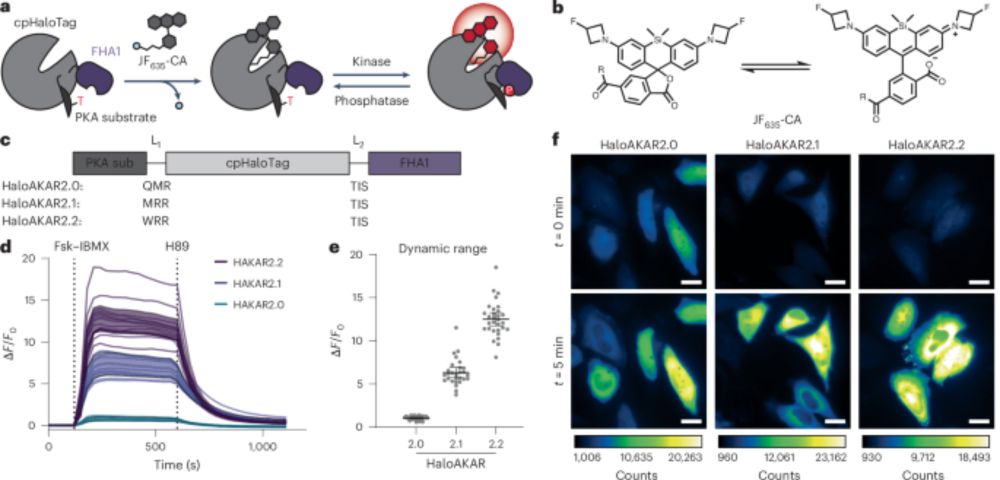
Far-red chemigenetic kinase biosensors enable multiplexed and super-resolved imaging of signaling networks - @michellefrei17.bsky.social @jinzhanglab.bsky.social go.nature.com/3GfqrzL
21.04.2025 14:45 — 👍 35 🔁 7 💬 1 📌 0Smart idea! Bring phos-sensor to transcription activation?
22.04.2025 05:39 — 👍 1 🔁 0 💬 0 📌 0Wow, I heard the report before. Congratulations!
22.04.2025 05:34 — 👍 1 🔁 0 💬 0 📌 0Wow, congratulations!
22.04.2025 05:31 — 👍 0 🔁 0 💬 0 📌 0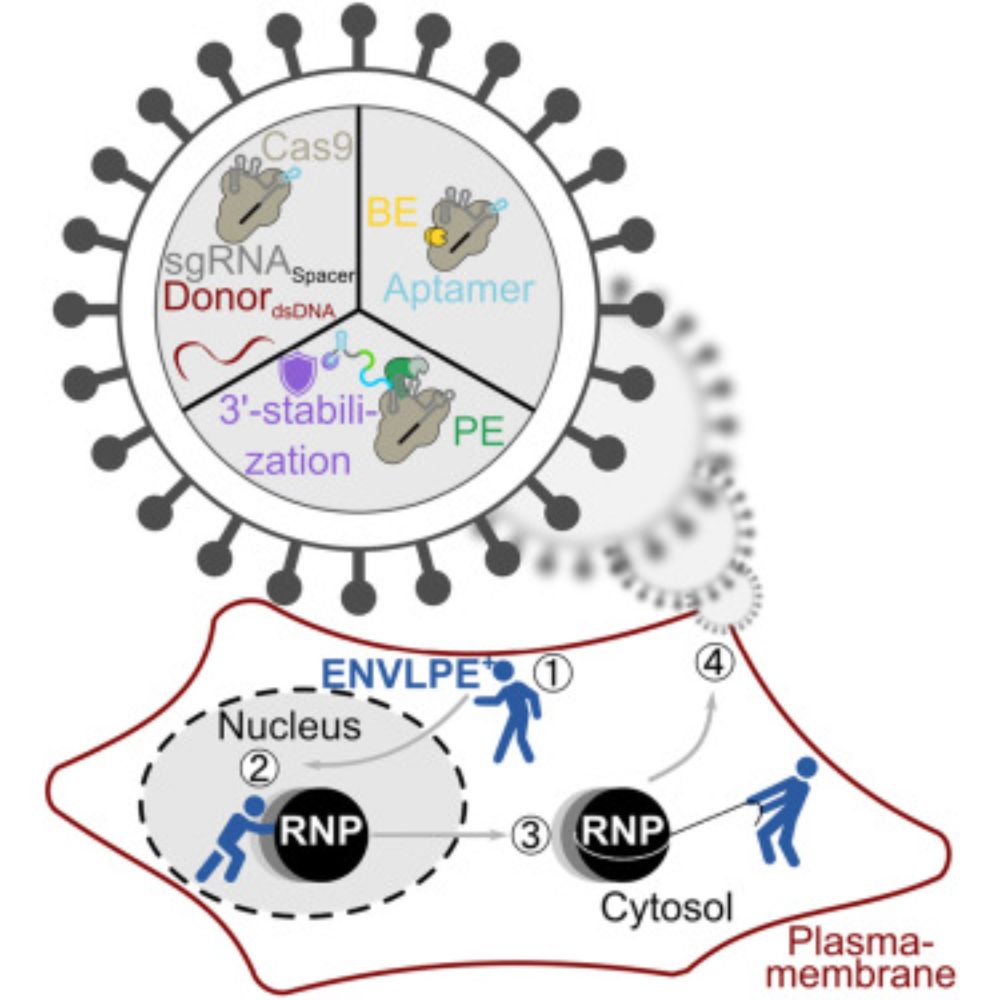
www.cell.com/cell/fulltex...
ENVLPE, our system for VLP-based delivery of gene editors, is now published in @cellcellpress.bsky.social. We showed efficient delivery of base- and prime-editors in primary T cells & in vivo retinal models. All core constructs are available from @addgene.bsky.social.
Great work and very impressive!
13.04.2025 02:49 — 👍 1 🔁 0 💬 0 📌 0