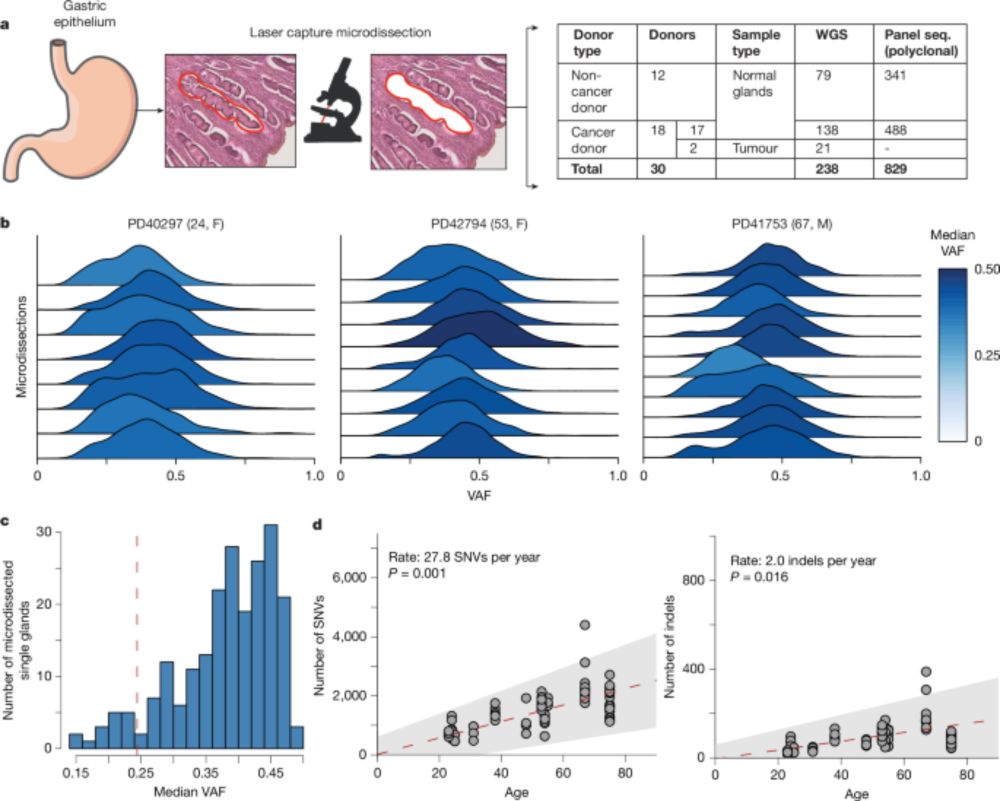
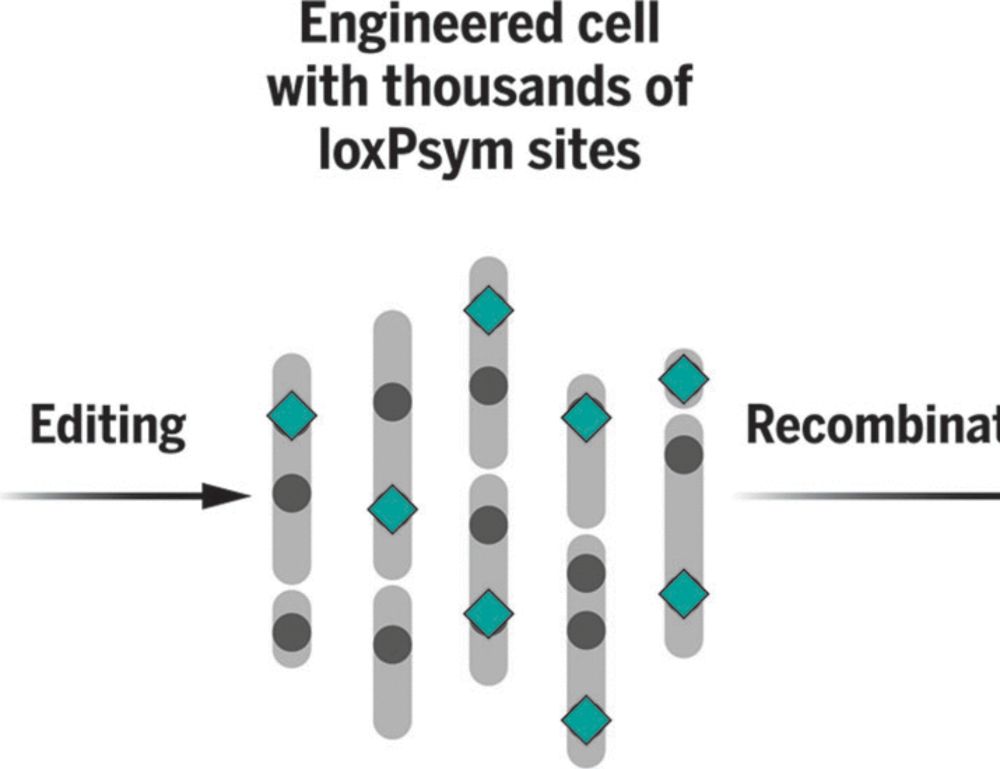
Very happy to share our venture into the wild world of imaging is now out on biorxiv. It has been the start of a very interesting journey with lots of interesting learnings along the way. I for sure will never make the mistake of considering a Jurkat a proper T cell again :)
12.02.2026 10:22 —
👍 4
🔁 1
💬 0
📌 0
See some pretty T cells in our preprint
www.biorxiv.org/content/10.6...
12.02.2026 10:00 —
👍 0
🔁 0
💬 0
📌 0
Thanks to co‑first @obbakker.bsky.social and lead from @gosiatrynka.bsky.social and co‑authors: Madeline Ohl, Francesco Cisternino, Andrea Manrique Rincón, Anke Husmann, Florence Lichou, Tong Li, Kwasi Kwakwa, Anneliese Speak, Craig Glastonbury, Omer Ali Bayraktar, Carla Jones, Melina Claussnitzer.
12.02.2026 09:59 —
👍 0
🔁 0
💬 1
📌 0
🤝 We’re excited to get the community involved and see TGlow (and these pipelines) applied to all sorts of crazy — and not‑so‑crazy — biological questions.
12.02.2026 09:55 —
👍 0
🔁 0
💬 1
📌 0
💾 🤖 For the imaging enthusiasts: our open‑source Nextflow pipeline and R toolkit work across imaging assays — not just T cells or TGlow — and enable reproducible processing, visualisation, and single‑cell statistical analysis, all while preserving that beautiful cellular heterogeneity.
12.02.2026 09:55 —
👍 0
🔁 0
💬 1
📌 0
🧠 The result: we’ve profiled >400,000 primary human CD4⁺ and CD8⁺ T cells with TGlow, uncovering activation-, drug-, CRISPR-, and exhaustion‑specific morphological programs — from mitochondrial remodelling to cytoskeletal collapse to lipid biogenesis.
12.02.2026 09:55 —
👍 0
🔁 0
💬 1
📌 0
🔬 Inspired by image‑based profiling approaches like Cell Painting, we tailored the experimental and computational pipelines to the immune system: integrating cyclic imaging to include functional markers (activation, cell cycle, metabolism), and pushing deeper in z‑space to capture 3D architecture.
12.02.2026 09:54 —
👍 0
🔁 0
💬 1
📌 0
💡 TGlow bridges this gap: a high-dimensional phenotyping platform for lymphocytes with omics-level scale and resolution, offering a complementary lens on downstream cellular behaviours.
12.02.2026 09:54 —
👍 0
🔁 0
💬 1
📌 0
❓ With single-cell omics, we can now profile immune cells at scale, down to every RNA and protein. But some questions still persists: how, or even if, do these molecular changes translate to function? what unfolds inside the cell afterward?
12.02.2026 09:54 —
👍 1
🔁 0
💬 1
📌 0
Thrilled to share TGlow, our high‑content imaging platform for scalable single‑cell phenotyping of lymphocytes. It complements single‑cell omics by revealing what happens next inside the cell and does so at scale. Details in the thread & link to our preprint: www.biorxiv.org/content/10.6...
12.02.2026 09:53 —
👍 3
🔁 0
💬 1
📌 1
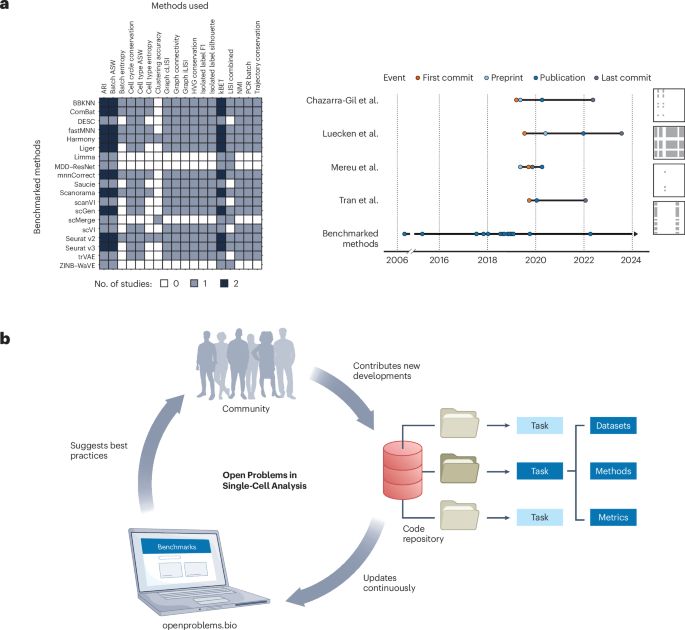
Defining and benchmarking open problems in single-cell analysis - Nature Biotechnology
Nature Biotechnology - Defining and benchmarking open problems in single-cell analysis
New OpenProblems paper out! 📝
Led by Malte Lücken with Smita Krishnaswamy, we present openproblems.bio – a community-driven platform benchmarking single-cell analysis methods.
Excited about transparent, evolving best practices for the field!
🔗 www.nature.com/articles/s41...
03.07.2025 06:35 —
👍 23
🔁 9
💬 0
📌 0
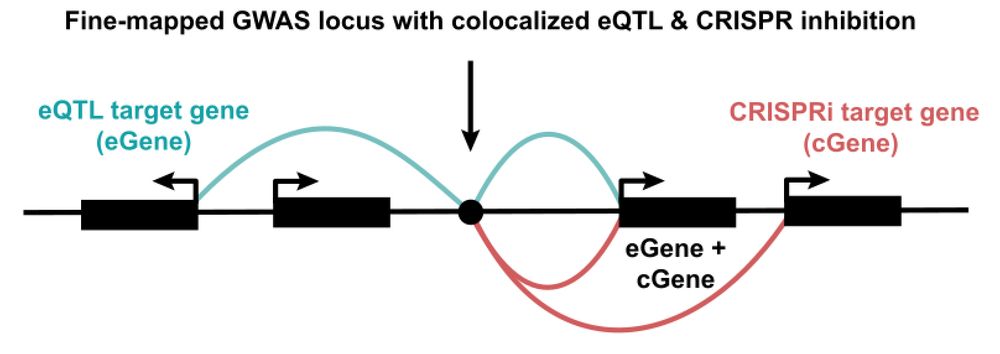
Our new contribution to the quest to find causal GWAS genes! Sam Ghatan from my lab at @nygenome.org led a systematic comparison of eQTLs and CRISPRi+scRNA-seq screens. TL;DR: they provide highly complementary insights, with ortogonal pros and cons. 🧵👇
www.biorxiv.org/content/10.1...
06.05.2025 17:00 —
👍 98
🔁 42
💬 1
📌 2
Out today in Nature Biotechnology!
This Open Targets project created an atlas of tissue-specific protein associations for 11 human tissues, furthering our understanding of cell-type and tissue-specific function, and helping to prioritise more specific—and potentially safer—drug targets 🧬🖥️
02.05.2025 11:02 —
👍 11
🔁 3
💬 0
📌 0

Genome-scale models in human metabologenomics | Nature Reviews Genetics
Metabologenomics integrates metabolomics with other omics data types to comprehensively study the genetic and environmental factors that influence metabolism. These multi-omics data can be incorporated into genome-scale metabolic models (GEMs), which are highly curated knowledge bases that explicitly account for genes, transcripts, proteins and metabolites. By including all known biochemical reactions catalysed by enzymes and transporters encoded in the human genome, GEMs analyse and predict the behaviour of complex metabolic networks. Continued advancements to the scale and scope of GEMs — from cells and tissues to microbiomes and the whole body — have helped to design effective treatments and develop better diagnostic tools for metabolic diseases. Furthermore, increasing amounts of multi-omics data are incorporated into GEMs to better identify the underlying mechanisms, biomarkers and potential drug targets of metabolic diseases. Metabologenomics integrates multi-omics data into geno
Explore human metabologenomics with GEMs, integrating 1000s of genes, proteins, and metabolites to uncover metabolism's mysteries! PMID:39300314, Nat Rev Genet 2025, @NatureRevGenet https://doi.org/10.1038/s41576-024-00768-0 #Medsky #Pharmsky #RNA #ASHG #ESHG 🧪
02.05.2025 09:10 —
👍 2
🔁 1
💬 0
📌 0


Tomorrow is the last day to apply to the Leena Peltonen School of Human Genomics summer program (Wellcome Campus, UK) for scientists nearing the completion of their PhD. 1:1 mentorship from many fantastic tutors!
lpshg.com/how-to-apply/
07.03.2025 02:50 —
👍 19
🔁 16
💬 0
📌 1
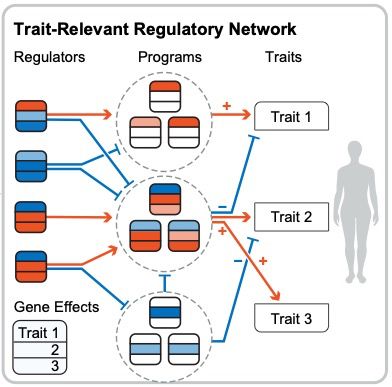
Modern GWAS can identify 1000s of significant hits but it can be hard to turn this into biological insight. What key cellular functions link genetic variation to disease?
I'm very excited to present our new work combining associations and Perturb-seq to build interpretable causal graphs! A 🧵
26.01.2025 00:13 —
👍 320
🔁 118
💬 6
📌 10
"The main question when reviewing a paper should be whether its conclusions are likely to be correct, not whether it would be important if it were true. Real advances are built with bricks, not straw."
A perpetual must-read: www.nature.com/articles/545...
18.01.2025 00:58 —
👍 36
🔁 11
💬 1
📌 0
The Week in Bio #34. Research highlights from the week including randomisation of a super-enhancer, catalytic LYTACs from Lycia Therapeutics and de novo design of binders for snake venom. Link below.
20.01.2025 16:04 —
👍 5
🔁 4
💬 1
📌 0
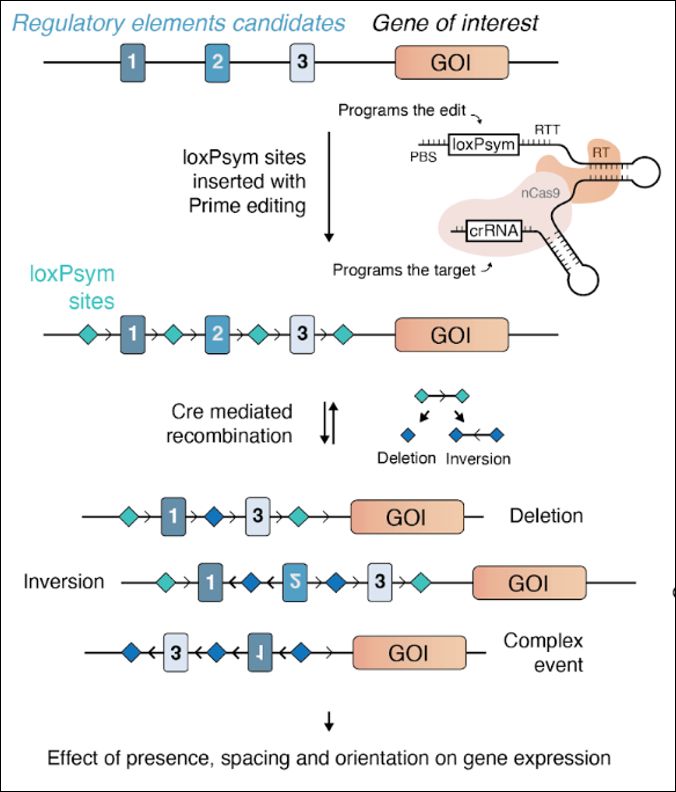
Enhancer scrambling strategy
We are happy to share our enhancer scramble story, a strategy to create hundreds of stochastic deletions, inversions, and duplications within mammalian gene regulatory regions and associate these new architectures with gene expression levels 🧵
www.biorxiv.org/content/10.1...
15.01.2025 20:32 —
👍 183
🔁 77
💬 3
📌 2

The Week in Bio #33
Research highlights from the week
The Week in Bio #33. Research highlights from the week including saturation mutagenesis of 500 human protein domains from the Lehner lab, Profluent Bio’s latest deep learning model and predicting RNA-seq from DNA with Borzoi. weekinbio.substack.com/p/the-week-i...
14.01.2025 13:13 —
👍 6
🔁 2
💬 0
📌 0