👀
20.06.2025 22:03 — 👍 0 🔁 0 💬 0 📌 0👀
20.06.2025 22:03 — 👍 0 🔁 0 💬 0 📌 0
I’m hiring for a microscopy image analysis role at Calico!
www.calicolabs.com/careers/?gh_...
We’re looking for someone at ~recent PhD lvl who has previous experience with biological samples and analyzing microscopy datasets.
Feel free to reach out with any questions!

Analyzing organelles is HARD. They're complex, but critical to understanding biology. Current tools are manual, organelle-specific, or can't handle large 3D datasets.
With an easy install and a click of a button, Nellie solves ALL these problems!
nature.com/articles/s41...
🧵2/N
This is definitely my favorite response yet haha, my favorite movie of the year!
10.03.2025 21:25 — 👍 1 🔁 0 💬 0 📌 0Thank you!!
10.03.2025 20:58 — 👍 0 🔁 0 💬 0 📌 0Thanks a lot Nico!
10.03.2025 20:58 — 👍 1 🔁 0 💬 0 📌 0Thanks again Felipe!
10.03.2025 20:43 — 👍 1 🔁 0 💬 0 📌 0Already some cool applications for Nellie: If you're used to MitoGraph data and want some of the same outputs from Nellie, check out Gav's Python code.
10.03.2025 20:37 — 👍 7 🔁 0 💬 1 📌 0Thanks for the post! Hope everyone finds it useful. And don’t hesitate to reach out if you have any questions or issues!
06.03.2025 17:28 — 👍 1 🔁 0 💬 0 📌 0🤟🏽
06.03.2025 17:27 — 👍 1 🔁 0 💬 0 📌 0
From Lefebvre and colleagues, Nellie is a comprehensive automated pipeline for studying the structure and intracellular dynamics of diverse organelles that offers accurate segmentation, tracking and feature extraction on both 2D and 3D data.
www.nature.com/articles/s41...
Great collection. Thanks for including Nellie!!
03.03.2025 06:46 — 👍 2 🔁 0 💬 0 📌 0We spend a lot of time thinking about segmenting cells, but organelles represent a similarly important challenge that is tricky due to their diversity in shapes and sizes. Please check out Nellie, a generalist tool for segmenting AND TRACKING organelles!
28.02.2025 02:13 — 👍 82 🔁 10 💬 1 📌 1Thanks a lot Darren!
27.02.2025 21:33 — 👍 0 🔁 0 💬 0 📌 0
Try and share Nellie today!
We've included some sample data if you want to explore what Nellie can do.
Code: github.com/aelefebv/nel...
Paper: nature.com/articles/s41...
And please share your results with us! We're excited to see how you'll use it in your research.
🧵Fin/N



We also compare Nellie to existing state of the art methods for both segmentation and tracking across imaging resolution, structure widths, and structure lengths, as well as the pipeline's speed on different sized images
🧵14/N

Now we can generate flow fields and get cool metrics, like if fields are diverging from (fission?) or converging to (fusion?) a point, or whether a branch is rotating along its long axis (?? dancing?). All of this, and much much more, linked to our organellar hierarchy.
🧵13/N

From these beacons, we can use a distance and cost-weighted interpolation to generate tracks for any coordinate within our organelles! Even at a sub-voxel level. We can use this to link voxels (and thus objects) between temporal frames too.
🧵12/N

We generate internal motion capture markers (think Hollywood) and pattern match these markers between frames to create beacons that point the way for the actual tracking. This means that we don't have to rely on finicky segmentations or skeletons for tracking.
🧵11/N

Then Nellie breaks down the organelles into connected components, and uses its skeleton to break it down even further into branches, which break down into individual nodes, which are variable-range containers for surrounding voxels. A big hierarchical organelle tree.
🧵10/N

So how does Nellie actually work? First Nellie figures out your image's metadata. Then it applies an automated structural-enhancement regime (modified Frangi filter + others) to make those organelles pop.
🧵9/N
Before deep-diving into Nellie, shout out my co-authors: @gavsturm.bsky.social , for help on all the paper, experiments, and testing, and to the other co-authors @kayleyhake.bsky.social, Ben, Emily, Magdalena, and Molly.
And of course @ritastrack.bsky.social for being a great editor!
🧵8/N
And just for fun, here's me walking around, masked by Meta's Segment Anything Model and tracked via Nellie's tracking pipeline
🧵7/N
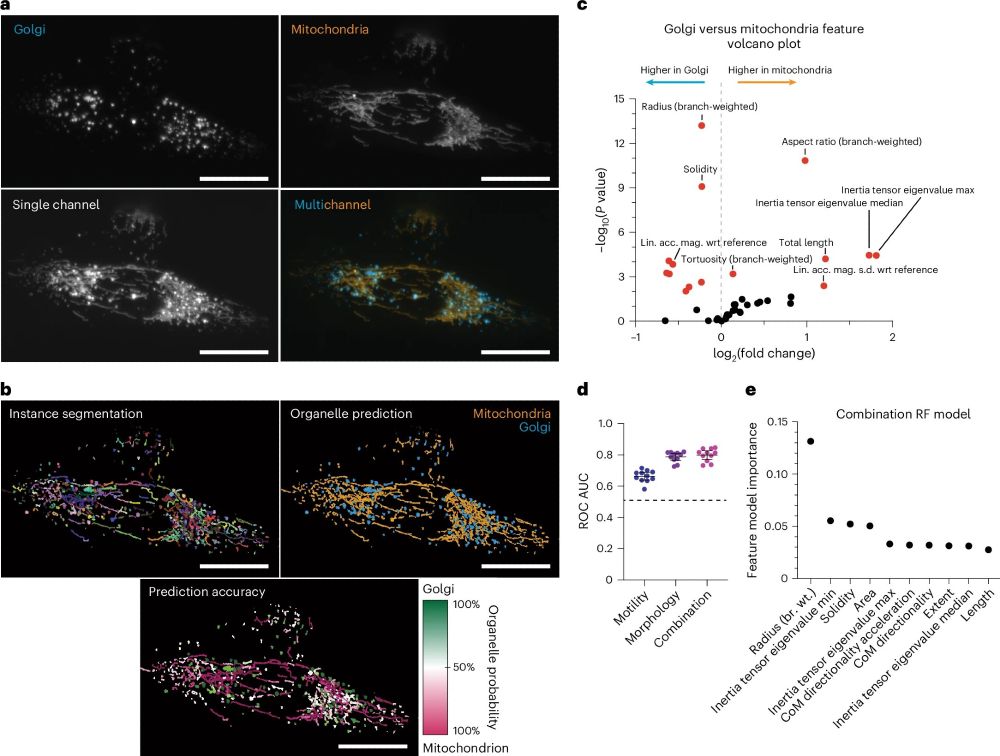
We used Nellie to unmix multiple organelles from a single fluorescence channel with high accuracy using just their morphology and motility patterns with a standard (random forest) classifier.
More fluorescence channels for your experiments!
🧵6/N

We also analyzed the intracellular ER highways using Nellie's outputs, allowing us to understand organellar rearrangements using its topology from the perspective of graph theory
🧵5/N

Inspired by Google Deepmind's weather forecasting model GraphCast, we also developed an organelle graph autoencoder using Nellie's outputs to analyze mitochondrial networks' responses to ionomycin treatment, revealing complex dynamics invisible to traditional analysis
🧵4/N
Nellie is FULLY automated with NO parameters to tune, and works on ALL structures - from mitochondria to ER to microtubules and beyond
(credit to @andrewgyork.bsky.social and @amsikking.bsky.social 's Snouty SOLS microscope for the data, and @alisterburt.bsky.social for the Animation plugin)
🧵3/N

Analyzing organelles is HARD. They're complex, but critical to understanding biology. Current tools are manual, organelle-specific, or can't handle large 3D datasets.
With an easy install and a click of a button, Nellie solves ALL these problems!
nature.com/articles/s41...
🧵2/N
Do you LOVE organelles??
Well I am THRILLED to announce that Nellie has been published in @naturemethods.bsky.social!
Nellie a fully automated pipeline for organelle segmentation, tracking, and hierarchical feature extraction in 2D, 3D, timelapse, multichannel live-cell microscopy
🧵1/N