
Στην Αττική τα data centers πολλαπλασιάζονται, χωρίς όμως να είναι σαφές από πού θα εξασφαλίζουν το νερό που χρειάζονται.
✍️ @akaradimitri.bsky.social
#insidestory_gr
insidestory.gr/article/ta-d...

Στην Αττική τα data centers πολλαπλασιάζονται, χωρίς όμως να είναι σαφές από πού θα εξασφαλίζουν το νερό που χρειάζονται.
✍️ @akaradimitri.bsky.social
#insidestory_gr
insidestory.gr/article/ta-d...
Closing out my year with a journal editor shocker 🧵
Checking new manuscripts today I reviewed a paper attributing 2 papers to me I did not write. A daft thing for an author to do of course. But intrigued I web searched up one of the titles and that's when it got real weird...

Three schematic diagrams. The first illustrates selective publishing of internal resection, the second selective causal focus, and the third selective access and funding for researchers.
1. We ( @jbakcoleman.bsky.social, @cailinmeister.bsky.social, @jevinwest.bsky.social, and I) have a new preprint up on the arXiv.
There we explore how social media companies and other online information technology firms are able to manipulate scientific research about the effects of their products.

Yet again, machine learning — even gussied up via the transformer architecture — encodes and reinforces societal biases.
This study reveals that LLM-based peer review relies heavily on author institution in its decisions.
arxiv.org/abs/2509.15122

Want to be able to start micro-editing on Wikipedia in a few minutes – and know why you could be very glad you did? Then my new post @plos.org is just what you need!
absolutelymaybe.plos.org/2025/09/21/w...
#Wikipedia
The new -outfmt 20 in BLAST 😍. No more frustrated googling to remember which column is which.....
06.09.2025 14:54 — 👍 68 🔁 19 💬 4 📌 2
Mindblowing research out of the wonderful @isemevol.bsky.social (where I'm delighed to be based for the next three months).
Opening line: "Living organisms are assumed to produce same-species offspring [1,2]. Here, we report a shift from this norm".
1. Darwin (1859)
2. Mayr (1942).

What was antibiotic resistance like before we ever used antibiotics? How did we change what antibiotic resistance genes looked like over 100 years?
Our paper looking at resistance genes from a century of NCTC historical isolates now out in mGen:
www.microbiologyresearch.org/content/jour...
True, however there is this interesting exception. Heard about it in the Night Science episode with Fischbach explorecourses.stanford.edu/search?view=...
30.08.2025 18:40 — 👍 1 🔁 0 💬 0 📌 0
xkcd comic about replication crisis https://xkcd.com/3117
@xkcd.com again provides for your meta-science slides (link xkcd.com/3117). I do wish we'd stop talking about a "crisis", bc it's been here for generations. Ppl used to pipette by mouth ffs. Enrico Fermi won a Nobel prize for a false result. But that doesn't mean we can't do better going forward.
20.07.2025 05:27 — 👍 114 🔁 17 💬 1 📌 0
OrthoFinder just dropped a major update
It’s faster, more accurate, and ready for thousands of genomes
Let’s break it down (1/10)
github.com/OrthoFinder/...
www.biorxiv.org/content/10.1...
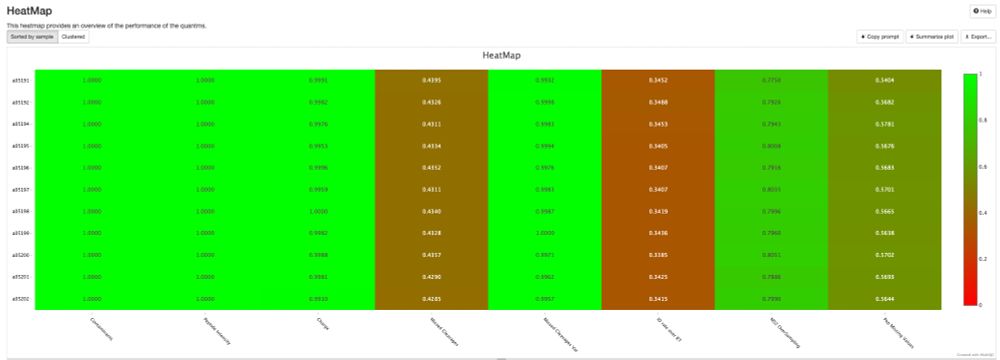
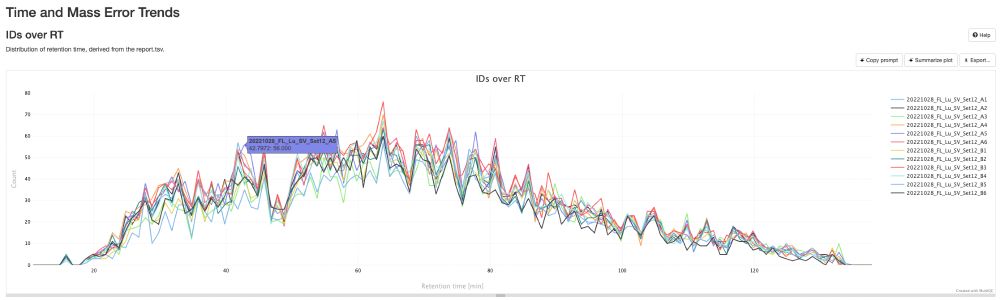
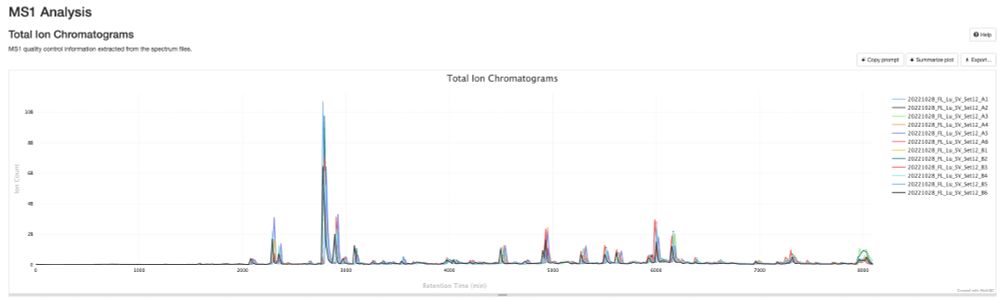
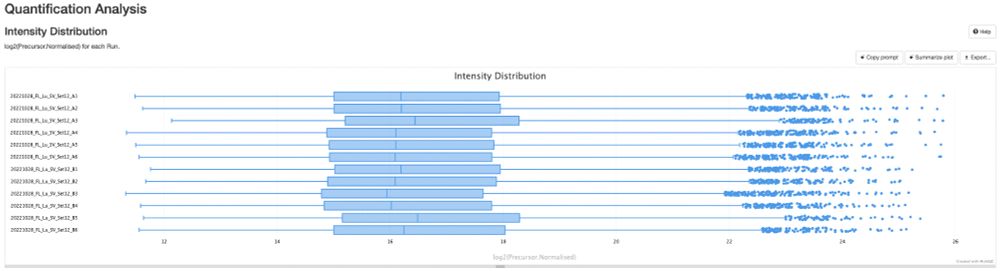
🚀 Attention DIA/DDA proteomics users! Whether you're using #DIA-NN, #MaxQuant, #quantms, or any tool that outputs mzIdentML and mzML, the NEW pmultiqc v0.0.29 is here!
💡 Create stunning, shareable HTML reports for your collaborators in seconds.
✨ Try pmultiqc.quantms.org Examples👇
#Proteomics #QC
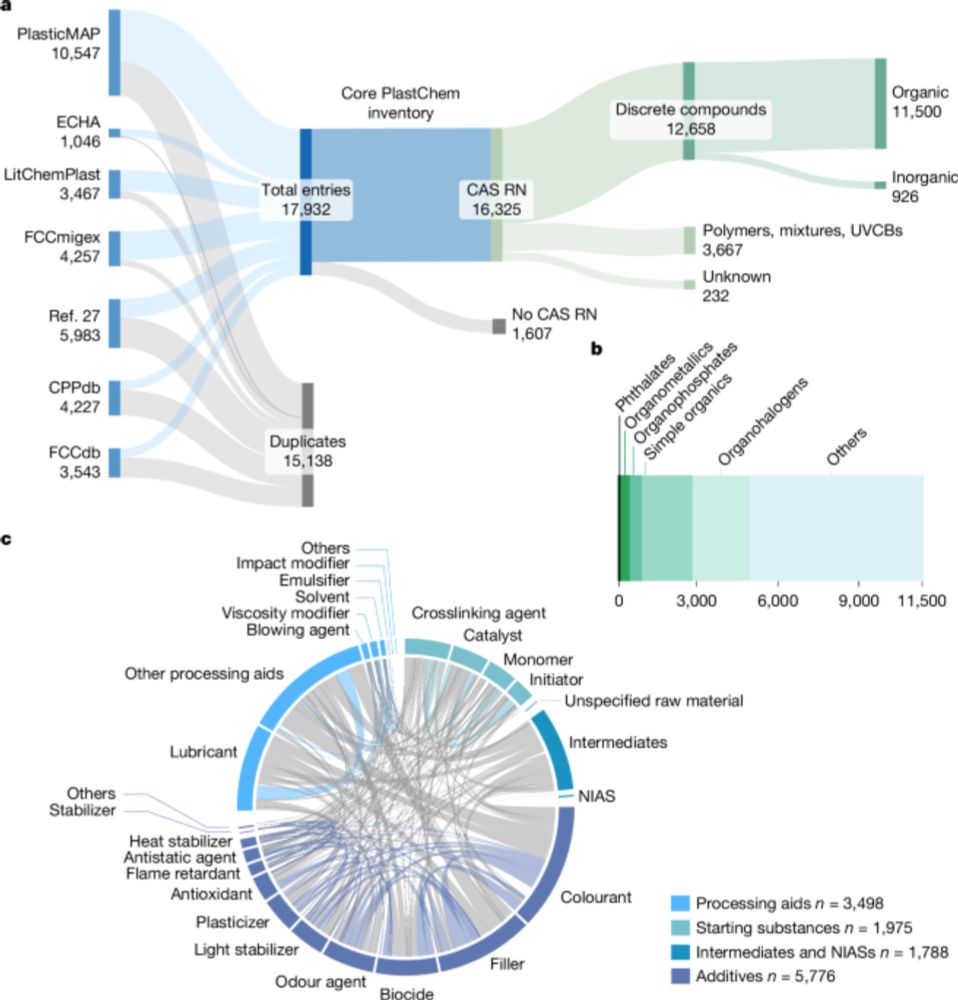
Over the past few years, our PlastChem team has worked tirelessly to map the chemical complexity of #plastics. Today, we publish the findings in @nature.com. We hope this evidence propels the development of safer plastics to protect human health and environment. 1/4
www.nature.com/articles/s41...
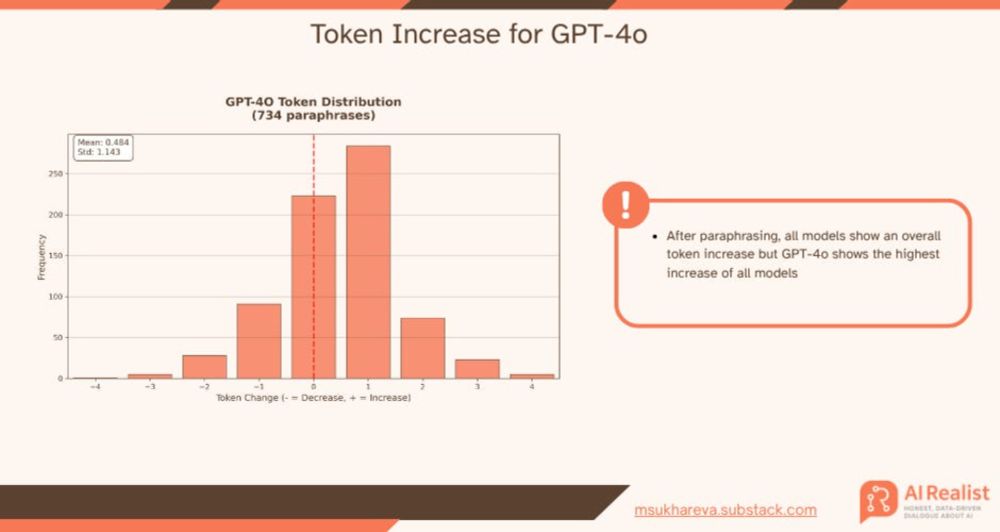
This is the best explanation I've seen yet for _why_ language models prefer em-dashes (—). In brief: The tokenization scheme mean em-dashes result in a smaller loss than other, equivalent punctuation options.
msukhareva.substack.com/p/the-myster...
DIA, DOA, DUI, DDA, etc. Here is a comparisons of some quantitative proteomics methods from a POV you might not have seen before:
github.com/pwilmart/qua...
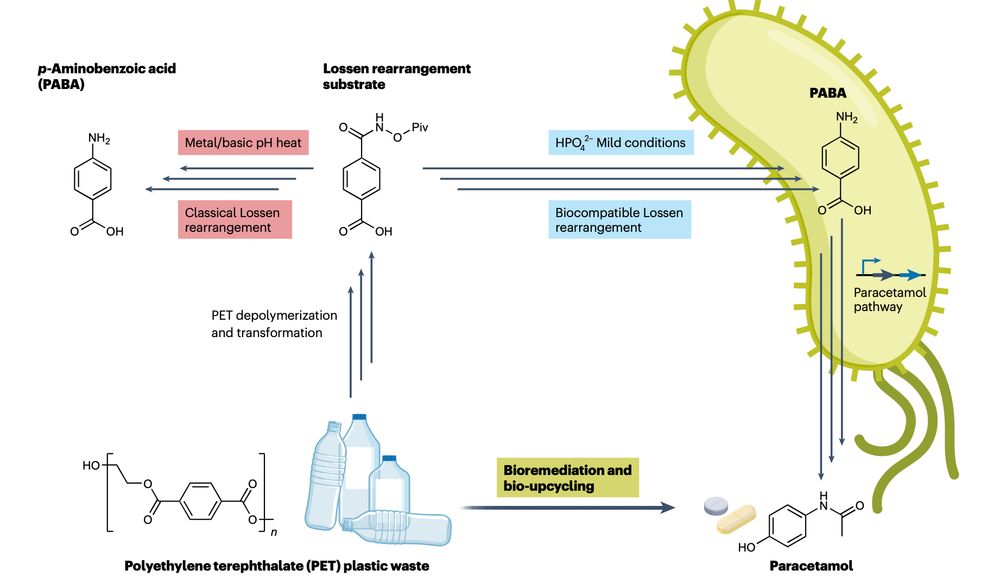
This is wild!
Engineering E. coli bacteria to turn plastic waste into paracetamol (Tylenol)
www.nature.com/articles/s41...
www.nature.com/articles/s41...
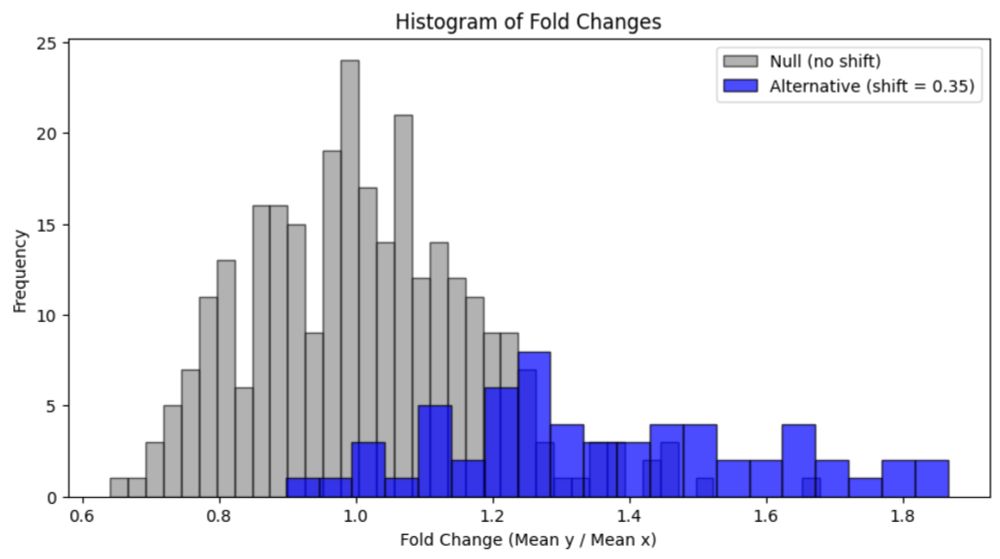
I wrote a review of a recent paper on false discovery and multiple testing correction. liorpachter.wordpress.com/2025/06/16/r...
16.06.2025 21:31 — 👍 144 🔁 41 💬 7 📌 7While *ideal* impact factor is well known to be a bad proxy of article quality, it is less well known that *actual* impact factor is negotiated and gamed by journals. You think Nature earned that impact factor? You think that's air you're breathing? (see e.g. journals.plos.org/plosmedicine...)
19.06.2025 11:35 — 👍 64 🔁 23 💬 1 📌 3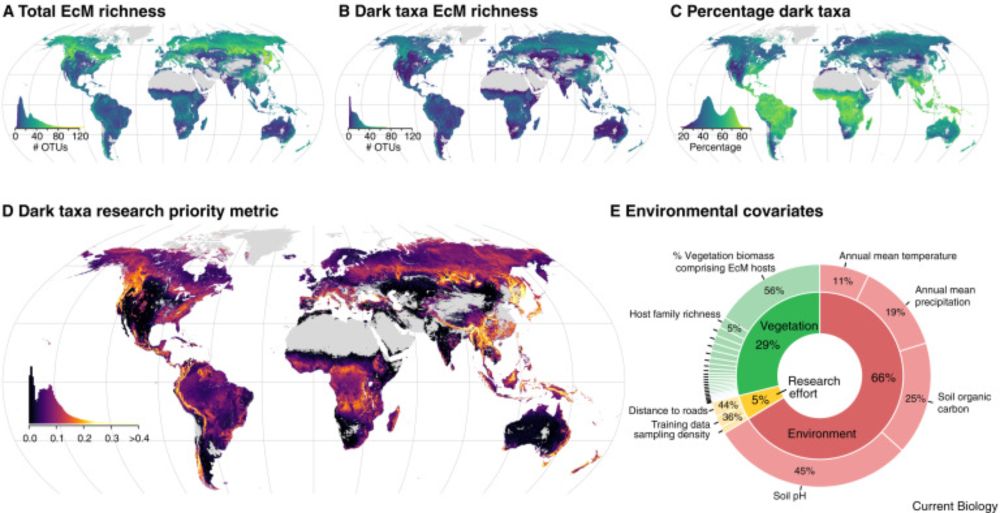
Our paper shows that most ectomycorrhizal fungal species (83% of OTUs) are "dark taxa": species we detect in DNA, but can't match to known species names. We map global "darkspots" - the parts of the world most in need of more research. @spun.earth @ethz.ch www.cell.com/current-biol...
10.06.2025 00:39 — 👍 92 🔁 41 💬 4 📌 4
Just a nice reminder! 🧪🧬🖥️ I found this GitHub very useful: github.com/cxli233/Frie...
25.05.2025 07:05 — 👍 38 🔁 15 💬 2 📌 0
Just out work lead by @mycomile.bsky.social @fungage-lab.bsky.social in A. fumigatus that demonstrate how large scale gene cluster movement is driving heterogeneity in this fungus -- Giant transposons promote strain heterogeneity in a major fungal pathogen pubmed.ncbi.nlm.nih.gov/40353686/
19.05.2025 18:34 — 👍 19 🔁 8 💬 1 📌 0
🧵5 Top Free Alternatives to BioRender for Scientific Illustrations!
These five websites offer free scientific illustrations for biologists. Great for presentations, research papers and other research communication needs.
Save and share the post!

I am very happy (and anxious) to share with you our most recent work in which we evaluated four of the most popular long-read assemblers,
www.biorxiv.org/content/10.1...
and tell you just a little bit about it in the following 🧵
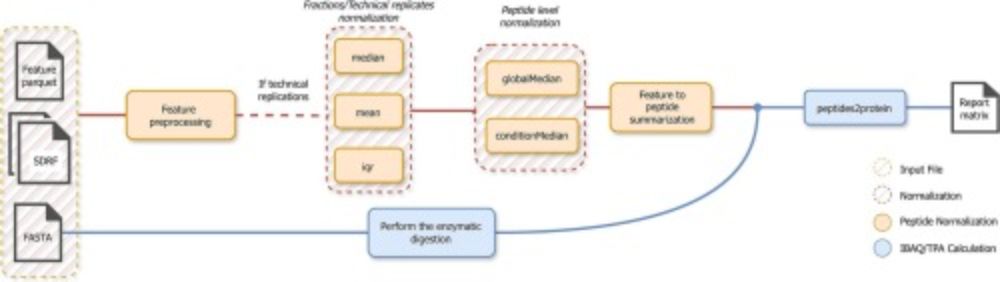
ibaqpy: A scalable #Python package for baseline quantification in proteomics leveraging #SDRF metadata www.sciencedirect.com/science/arti.... We continue our effort to provide tools and algorithms that leverage SDRF metadata to perform quantitative proteomics analysis.
25.04.2025 12:22 — 👍 12 🔁 9 💬 1 📌 0YouTube was a life saver in the beginning (thank you MaxQuant and May playlists). Doing PhD in microbial Biotechnology and my lab got free proteomics samples from the EPIC-XS consortium so someone had to analyze the data. Was familiar with HPLC an MS concepta
20.03.2025 14:25 — 👍 2 🔁 0 💬 0 📌 0
Lungfish xkcd.com/3064
17.03.2025 16:53 — 👍 11921 🔁 1308 💬 96 📌 87
Did you know that Greece is one of the most biodiverse countries in #Europe? 🇬🇷
Greek partners institutions in @biogeneurope.bsky.social are using #genomic data to help study and protect these unique species 🧬
Learn more: www.erga-biodiversity.eu/post/navigat... @iboleurope.bsky.social
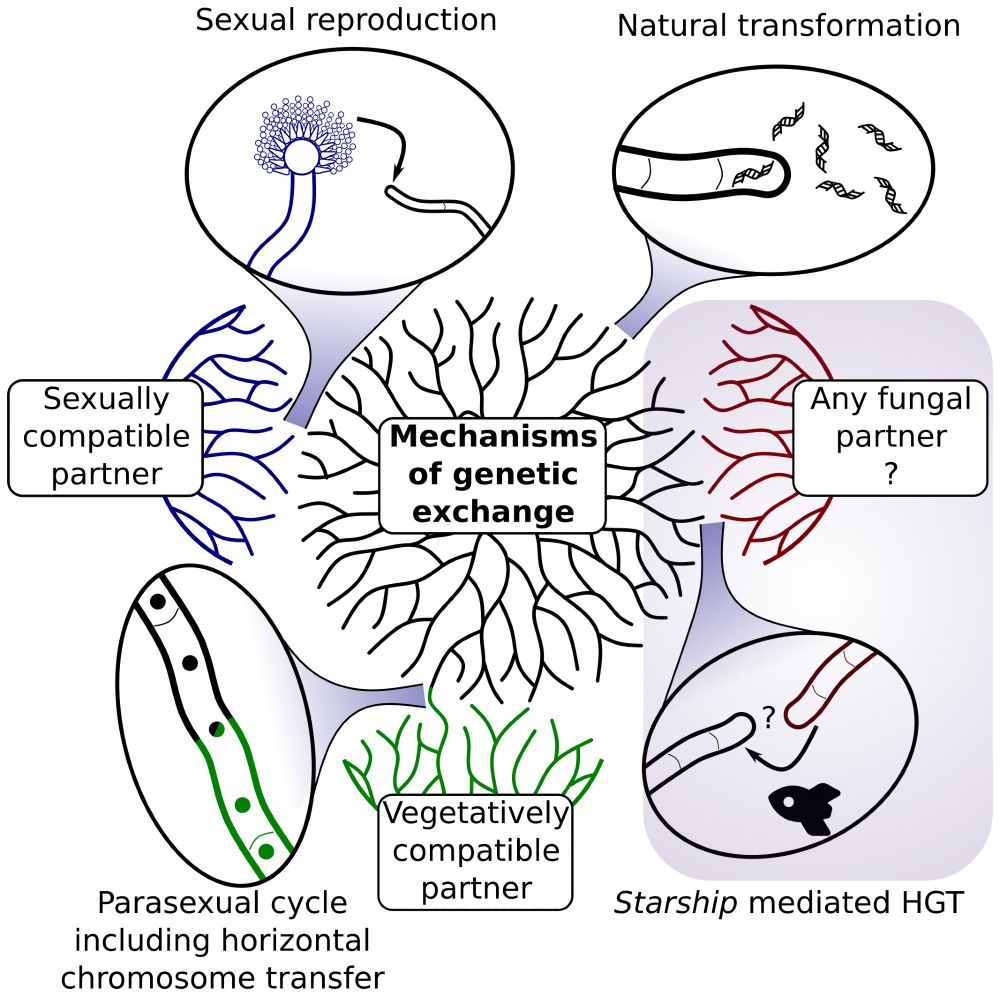
Illustration depicting the ways in which fungi can exchange genetic material with each other. Starship mediated HGT is confirmed experimentally in this preprint.
We did it! We caught Starship #transposons moving between #fungal species in the lab, including between species separated by ~100my. We think Starships are a mediator of HGT in fungi, akin to conjugative elements in bacteria. Check out the preprint. www.biorxiv.org/content/10.1...
07.03.2025 06:18 — 👍 255 🔁 137 💬 3 📌 10
Once upon a time, in the late 1800s, people in Japan got really into breeding mice.
Coloured mice. Patterned mice. Even mice that danced.
They became known as Japanese Fancy Mice, and that caught the attention of researchers in Europe and America, who imported them for study.
2/n