Looking for suggestions for concentrating multiple low concentration, replicate 50 uL DNA and RNA extractions ranging from 0-3 ng/uL. Column or mag bead preference? Or SpeedVac? TIA!! Please share.
28.02.2026 01:38 —
👍 1
🔁 1
💬 0
📌 0

We show that losing the N-term slows growth under temp/pH stress, while adding archaeal C-term tails boosts fitness at high temp + anaerobiosis.
Take home: Evolution of a conserved IF2 core with stress-tuning evolutionary add-ons.
Study led by Evrim Fer @uwmadisonmdtp.bsky.social! #translation
11.02.2026 18:21 —
👍 0
🔁 1
💬 0
📌 0

Latest work! 🧬
We uncover how evolution of translation initiation factor 2 (IF2) extensions links translation to bacterial stress response. We map 7 structural architectures & show how terminal extensions are enriched in intrinsic disorder & phase-separation features.
Link: doi.org/10.64898/202...
11.02.2026 18:18 —
👍 25
🔁 13
💬 2
📌 0

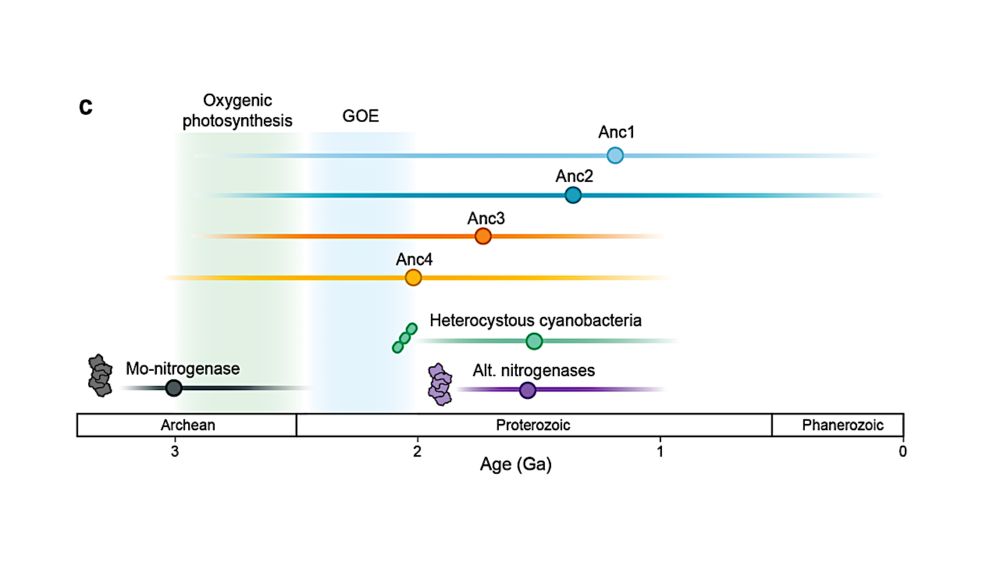
Ecological constraints and evolutionary trade-offs shape nitrogen fixation across habitats
Abstract. From its earliest beginnings, life’s expansion into new habitats has been profoundly shaped by its reciprocal interactions with Earth’s changing
New paper! Why are some Nitrogen fixing microbes more complex?
@msobol.bsky.social et al. find that microbes with more N2-fixation genes have larger, more versatile genomes, showing how changing environments shaped this key metabolism!
> academic.oup.com/ismecommun/a... @isme-microbes.bsky.social
06.02.2026 16:44 —
👍 12
🔁 11
💬 0
📌 0

We show that losing the N-term slows growth under temp/pH stress, while adding archaeal C-term tails boosts fitness at high temp + anaerobiosis.
Take home: Evolution of a conserved IF2 core with stress-tuning evolutionary add-ons.
Study led by Evrim Fer @uwmadisonmdtp.bsky.social! #translation
11.02.2026 18:21 —
👍 0
🔁 1
💬 0
📌 0

Latest work! 🧬
We uncover how evolution of translation initiation factor 2 (IF2) extensions links translation to bacterial stress response. We map 7 structural architectures & show how terminal extensions are enriched in intrinsic disorder & phase-separation features.
Link: doi.org/10.64898/202...
11.02.2026 18:18 —
👍 25
🔁 13
💬 2
📌 0
Thank you!
10.02.2026 14:00 —
👍 1
🔁 0
💬 0
📌 0
Fabulous work! 👏
10.02.2026 04:32 —
👍 2
🔁 1
💬 1
📌 0
Me pregunto qué pensaría Susumu Ohno de que los eventos más antiguos que podemos rastrear mediante métodos filogenéticos sean procesos de duplicación génica.
07.02.2026 02:44 —
👍 4
🔁 1
💬 0
📌 1
a daring approach: looking at LUCA's ancestors, i.e. pre-darwinian evolution. but not surprising it's coming from @kacarlab.bsky.social 👏
glad to see Iwabe et al. (1989) among the references (blew my mind when it came out >30 y ago)
06.02.2026 22:42 —
👍 21
🔁 10
💬 1
📌 0
Bookmarking, #bookmark
06.02.2026 17:21 —
👍 1
🔁 1
💬 0
📌 0
Thank you!
06.02.2026 17:12 —
👍 1
🔁 0
💬 0
📌 0
Very nice work!!
06.02.2026 16:42 —
👍 2
🔁 1
💬 1
📌 0

Ecological constraints and evolutionary trade-offs shape nitrogen fixation across habitats
Abstract. From its earliest beginnings, life’s expansion into new habitats has been profoundly shaped by its reciprocal interactions with Earth’s changing
New paper! Why are some Nitrogen fixing microbes more complex?
@msobol.bsky.social et al. find that microbes with more N2-fixation genes have larger, more versatile genomes, showing how changing environments shaped this key metabolism!
> academic.oup.com/ismecommun/a... @isme-microbes.bsky.social
06.02.2026 16:44 —
👍 12
🔁 11
💬 0
📌 0

NASA
NASA.gov brings you the latest news, images and videos from America's space agency, pioneering the future in space exploration, scientific discovery and aeronautics research.
Our latest paper on ancient microbes and nitrogen is featured on the NASA website today! 🚀🔬Way to go, @hollyrucker.bsky.social!
Resurrecting Ancient Enzymes in NASA’s Search for Life Beyond Earth
www.nasa.gov
Paper link: www.nature.com/articles/s41... #astrobiology @uwmadscience.bsky.social
02.02.2026 16:02 —
👍 9
🔁 4
💬 0
📌 0
I wonder if some of the Archean fossils I've found were using these molecules?
01.02.2026 16:46 —
👍 7
🔁 2
💬 0
📌 0
Thank you!
31.01.2026 01:49 —
👍 1
🔁 0
💬 0
📌 0

Resurrecting Ancient Enzymes in NASA's Search for Life Beyond Earth - NASA Science
NASA-supported scientists have resurrected an enzyme first used by organisms on Earth 3.2-billion years ago and, in the process, have validated a chemical
Check out NASA’s new press release on how we recreated a molecule used by some of Earth’s earliest life forms, helping us understand how life began here and how we might spot signs of life on other planets! Big congrats to first author, @hollyrucker.bsky.social!
science.nasa.gov/science-rese...
30.01.2026 21:16 —
👍 8
🔁 3
💬 0
📌 0