Really excited to share the latest work from my PhD with @giuliotesei.bsky.social and @lindorfflarsen.bsky.social!
09.02.2026 12:55 — 👍 6 🔁 4 💬 1 📌 0Really excited to share the latest work from my PhD with @giuliotesei.bsky.social and @lindorfflarsen.bsky.social!
09.02.2026 12:55 — 👍 6 🔁 4 💬 1 📌 0New preprint with @invemichele.bsky.social, Sandro Bottaro, Kamil Tamiola, and @lindorfflarsen.bsky.social on transiently structured states of IDPs sampled in atomistic simulations with enhanced sampling and integrated with experimental data.
21.01.2026 18:01 — 👍 13 🔁 4 💬 0 📌 1
Integrative structural biology by the sea
The Jan 31st deadline for our FEBS advanced course on integrative structural biology is getting closer, so get your applications in for Lost In Integrative Vol. 2
probingbiomolecules2026.febsevents.org
network.febs.org/posts/integr...

Figure illustrating the role of machine learning in IDP design, showing both the training data, machine learning and applications
Looking for something to read over the weekend?
Our review on computational design of intrinsically disordered proteins is now published in final form 🍝
With @giuliotesei.bsky.social & @fpesce.bsky.social
doi.org/10.1016/j.sb...
First preprint of the year, led by Junjie Zhu from Haifeng Chen’s lab
Extending Conformational Ensemble Prediction to Multidomain Proteins and Protein Complex
doi.org/10.64898/202...

New preprint!🚨 We're finally able to sample secondary structure in coarse-grained simulations of IDPs! 🍝🍝🍝 With an AI-based prediction of NMR chemical shifts in CG simulations, we correct dynamics on the fly, making secondary structure emerge as dictated by expts😍! www.biorxiv.org/content/10.6... 🧵👇
07.01.2026 07:03 — 👍 16 🔁 5 💬 1 📌 1
We (@sobuelow.bsky.social) developed AF-CALVADOS to integrate AlphaFold and CALVADOS to simulate flexible multidomain proteins at scale
See preprint for:
— Ensembles of >12000 full-length human proteins
— Analysis of IDRs in >1500 TFs
📜 doi.org/10.1101/2025...
💾 github.com/KULL-Centre/...
Registration now open for the '3rd Annual Danish Workshop on Advanced Molecular Simulation'
Join us in beautiful Aarhus for a 2-day workshop on biomolecular simulations!
9-10 December 2025
Link for registration: aias.au.dk/events/show/...

Check out our latest, where we investigate the molecular origin of differences between Lys and Arg and their roles in biomolecular #condensates
@dipcehu.bsky.social
www.biorxiv.org/content/10.1...

Figure 1 from the review. Caption: Comparison of a schematic example showing static, time-dependent, and time-resolved experiments illustrated by a protein folding process. (a) A static experiment measuring the observable O$_{\text{exp}}$ is shown, which can be modelled as a distribution of simulated values, O$_{\text{calc}}$, representing a conformational ensemble of folded and unfolded states. (b) Shows a time-dependent experiment, where the equilibrium dynamics of reversible folding gives rise to measured transition times $\tau_1$ and $\tau_2$. These can be modelled as equilibrium dynamics, illustrated by a free energy (FE) surface along a chosen degree of freedom (D.O.F.) (c) A time-resolved experiment probes a non-equilibrium process, where the system begins at $t_{0}$ in the folded state. During the observation time $t$ the protein unfolds until $t_{\text{max}}$. At each time point, a distinct ensemble average, O$_{\text{exp}}$, can be observed, reflecting the proteins changing structure. This evolution can be modelled as distributions of O$_{\text{calc}}$ at each time point. These are shown together with a FE surface.
Integrative modelling of biomolecular dynamics
Time-dependent and -resolved experiments combined with computation provide a view on molecular dynamics beyond that available from static, ensemble-averaged experiments
Review w @dariagusew.bsky.social & Carl G Henning Hansen
doi.org/10.48550/arX...

Snapshot of a condensate near a lipid membrane with Swedish Research Council and Malmö University logos.
I'm hiring for a PhD position at Malmö University, Sweden!
The project will focus on molecular modelling of proteins, lipids, and biomolecular condensates at cell membranes.
More details and application form: tinyurl.com/4zm92365
Please feel free to share!
@vetenskapsradet.bsky.social | @mau.se

Why are transcription factors disordered? Come join us in Oxford as a postdoc and we'll find out together!
Help publish 3 mature projects, AND develop cool new single molecule fluorescence binding assays!
biophysics
transcription
protein:DNA interactions
my.corehr.com/pls/uoxrecru...
Excited to see our review now on arXiv, written together with @fpesce.bsky.social and @lindorfflarsen.bsky.social
doi.org/10.48550/arX...
Our paper on:
A coarse-grained model for simulations of phosphorylated disordered proteins
(aka parameters for phospho-serine and -threonine for CALVADOS)
is now published in Biophysical Journal
authors.elsevier.com/a/1lTcE1SPTB...
@asrauh.bsky.social @giuliotesei.bsky.social & Gustav Hedemark
Arriën & Giulio's paper on
A coarse-grained model for disordered proteins under crowded conditions
(that is the CALVADOS PEG model) is now published in final form:
dx.doi.org/10.1002/pro....
@asrauh.bsky.social @giuliotesei.bsky.social
AlphaFold is amazing but gives you static structures 🧊
In a fantastic teamwork, @mcagiada.bsky.social and @emilthomasen.bsky.social developed AF2χ to generate conformational ensembles representing side-chain dynamics using AF2 💃
Code: github.com/KULL-Centre/...
Colab: github.com/matteo-cagia...
Happy to share a walkthrough of the applications of our package for simulations using CALVADOS!
Big thanks to @sobuelow.bsky.social, @lindorfflarsen.bsky.social, and the whole team for making this possible.
Thrilled to mark this as my first last-author paper!
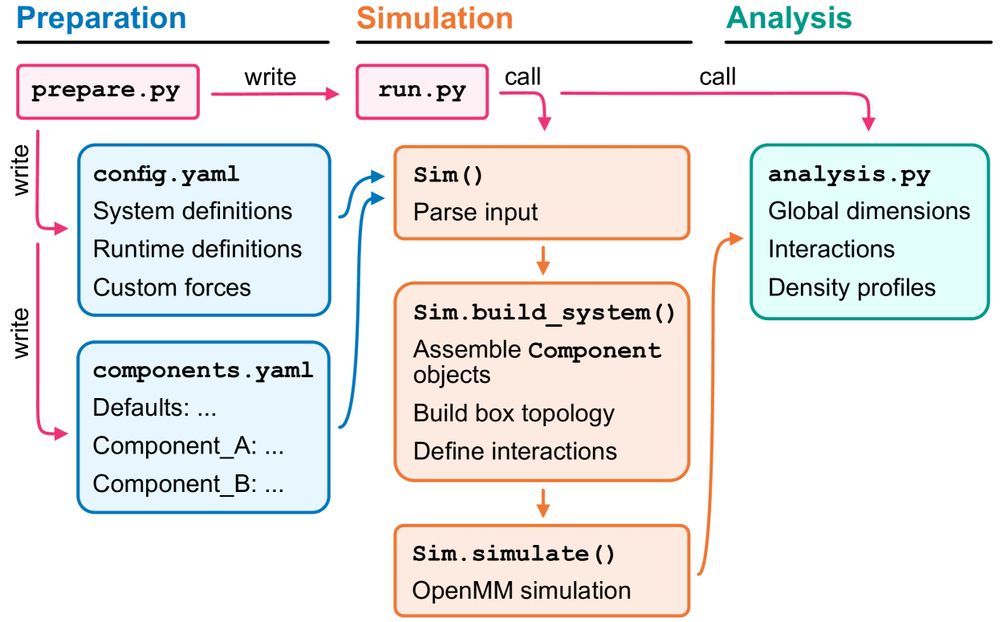
Figure showing the architecture of the CALVADOS package.
Do you like CALVADOS but are not quite sure how to make it?
We’ve got your back!
@sobuelow.bsky.social & @giuliotesei.bsky.social—together with the rest of the team—describe our software for simulations using the CALVADOS models incl. recipes for several applications. 1/5
doi.org/10.48550/arX...
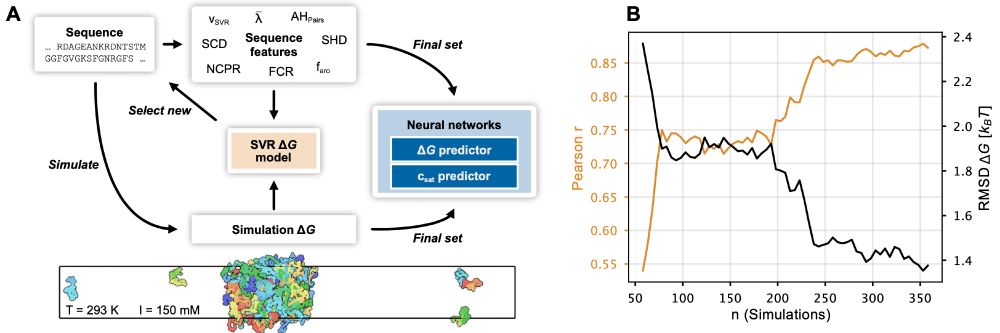
Our paper on prediction of phase-separation propensities of disordered proteins from sequence is now published:
www.pnas.org/doi/10.1073/...
The paper has been substantially updated compared to the preprint including new experimental data and using the neural network to finetune CALVADOS. 1/n
CALVADOS now has parameters for phosphorylated amino acids
@asrauh.bsky.social @giuliotesei.bsky.social and Gustav Hedemark used a top-down approach in which we targeted experimental data to derive parameters or phosphorylated serine and threonine doi.org/10.1101/2025...
Excited to share our PEG model for disordered proteins in CALVADOS!
If you are interested in exploring the effects of a crowder on the global dimensions of an IDP or want to explore the phase separation behaviour of a more weakly PS-prone IDP, have a look at our preprint and give it a try.
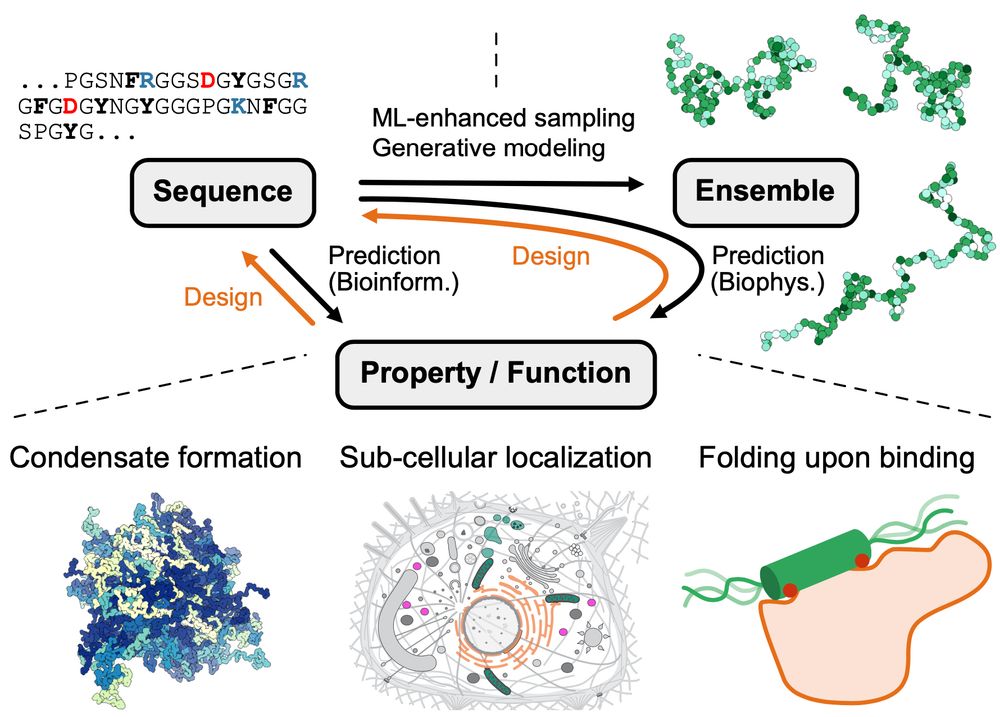
Figure from the paper illustrating sequence–ensemble–function relationships for disordered proteins. ML prediction (black) and design (orange) approaches are highlighted on the connecting arrows. Prediction of properties/functions from sequence (or vice versa, design) can include biophysics approaches via structural ensembles, or bioinformatics approaches via other hetero- geneous sources. The lower panels show examples of properties and functions of IDRs for predictions or design targets. ML, machine learning; IDRs, intrinsically disordered proteins and regions.
Our review on machine learning methods to study sequence–ensemble–function relationships in disordered proteins is now out in COSB
authors.elsevier.com/sd/article/S...
Led by @sobuelow.bsky.social and Giulio Tesei