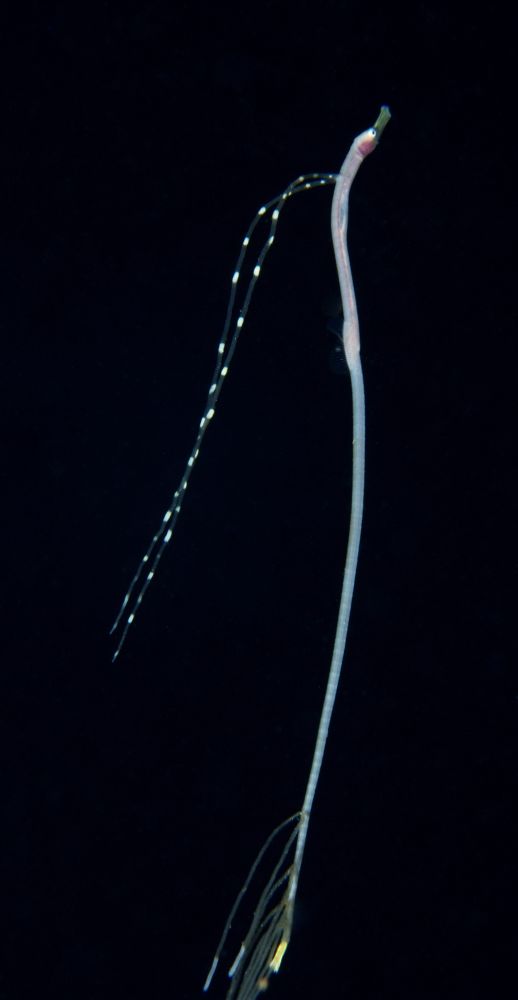
A very long, thin fish. Two dorsal spines are extended into long filaments that extend over the back; there are more filaments extending from the tail.
#UnderwaterPhotography #Macro #Blackwater #Wildlife #Nature #NaturePhotography #WildlifePhotography
A juvenile pipefish. I believe the long filaments probably mimic siphonophores, but I'm no expert on this stuff. Pic from Balayan Bay, Anilao
#MarineLife 🐟🌿
09.09.2025 00:10 —
👍 112
🔁 24
💬 0
📌 0

Population structure of the endangered Siberian flying squirrel Pteromys volans revealed by genomic and mitochondrial data vist.ly/32xin #Conservation #GeneFlow
11.08.2025 10:20 —
👍 8
🔁 2
💬 0
📌 0
Does genetic rescue disrupt local adaptation? An experimental test using thermally adapted Tribolium castaneum lines https://www.biorxiv.org/content/10.1101/2025.08.07.669094v1
11.08.2025 05:32 —
👍 6
🔁 2
💬 0
📌 0

Macrosyntenic evolution of clitellates.
An episodic burst of massive genomic rearrangements and the origin of non-marine annelids 🧪
www.nature.com/articles/s41...
Suggests genomic landscape of Clitellata resulted from a rare burst of genomic changes that ended a long period of stability that persists across large phylogenetic distances.
07.08.2025 16:40 —
👍 6
🔁 2
💬 0
📌 0

Phylogeny and nucleotide diversity of the hamlet radiation.
Radiation with reproductive isolation in the near-absence of phylogenetic signal 🧪
www.science.org/doi/10.1126/...
Shows phenotypic diversification and reproductive isolation—2 major attributes of species—may unfold in near-absence of phylogenetic signal, both genome-wide and at the gene-tree level
07.08.2025 17:00 —
👍 4
🔁 1
💬 2
📌 0

Logo for the Genomic History Inference Strategies Tournament.
If you're new to demographic history inference from population genomics, try this webapp I created to illustrate how dadi fits bottleneck models to site frequency spectra: ryangutenkunst-dadi-two-epoch.hf.space . It even outputs files for submitting to the GHIST competition! ghi.st
05.08.2025 18:48 —
👍 39
🔁 21
💬 1
📌 2

Phenotypic analysis of body standard length and gut length data. a) Phenotypic survey of individuals raised under common diet and housing conditions supports a genetic basis of normalized intestine length variation by species. Letters in parentheses indicate trophic level of species (C, carnivore; H, herbivore; O, omnivore). Inset fish images are representative individuals of parental species for hybrid cross. b) Binning intestine length data by trophic level of species shows that carnivores have significantly shorter intestines than omnivores or herbivores, but omnivores and herbivores are not significantly different from each other. The hybrid F2 mapping population (MmAkF2) exhibits intermediate normalized intestine length distinct from both carnivore and omnivore–herbivore species bins (ANOVA with Tukey's HSD, different letters indicate significance at P < 0.05). c) Standard body length and d) gut length in the parental species (N = 10 each) of the hybrid cross, and female (N = 218) and male (N = 230) individuals of the hybrid F2 population (t-test, ***P < 0.001, **P < 0.01). e) Scatter plot of standard length vs gut length for individuals in the F2 mapping population (circles), and the parental samples (squares: M. mbenjii; triangles: A. koningsi). The linear regression line of the model GL∼SL with confidence interval is shown for the F2 individuals. f) Residuals of the model GL∼SL are not significantly different by sex in the F2 hybrids. All sampled F2 individuals (N = 495) are presented in a) and b). Only the F2 individuals with unambiguous sex calls and used for mapping (N = 448) are included in c–e).
Carmona Baez, A., et. al. 2025. Gut length evolved under sexual conflict in Lake Malawi cichlids, Genetics, Volume 230, Issue 3, July 2025, iyaf102. 🐟🧪
doi.org/10.1093/gene...
10.07.2025 13:28 —
👍 7
🔁 3
💬 0
📌 1

Dia taking notes while examining a herbarium sheet of Aconitum reclinatum.
Woot! @diaathena.bsky.social submitted her final MS Thesis to Bucknell today!
Assessing Genetic Diversity and Population Structure in the Imperiled #Aconitum reclinatum (white monkshood), an Endemic Wildflower of Central Appalachia.
Great work that will help to protect a cool plant.
18.07.2025 17:37 —
👍 26
🔁 2
💬 0
📌 0
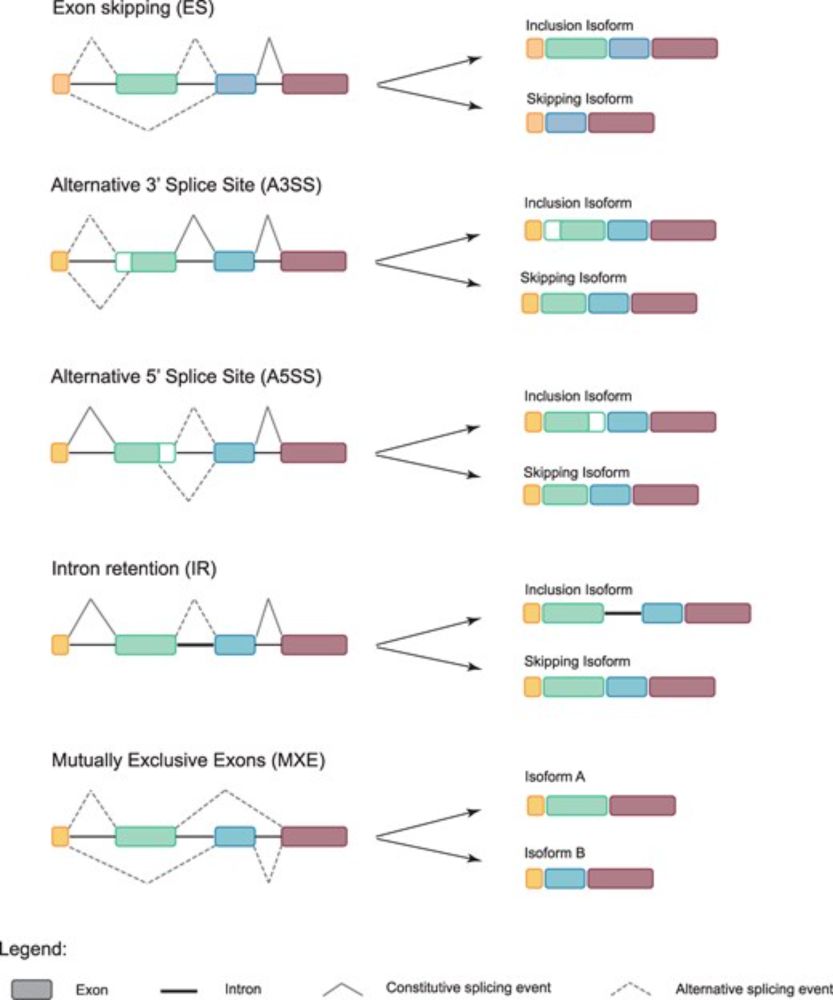
The Role of Alternative Splicing in Marine–Freshwater Divergence in Threespine Stickleback
Abstract. Alternative splicing regulates which parts of a gene are kept in the messenger RNA and has long been appreciated as a mechanism to increase the d
To test whether alternative splicing contributes to adaptation, Rodríguez-Ramírez & Peichel analyzed gill RNA-seq data from marine and freshwater sticklebacks, and identified differentially spliced genes enriched in regions under divergent selection.
🔗 doi.org/10.1093/gbe/evaf105
#genome #RNAseq
17.06.2025 14:39 —
👍 13
🔁 4
💬 0
📌 0


Presenting a poster at 2nd Biodiversity Interdisciplinary Meeting in Mallorca, Spain.
I present my MSc thesis, Population genetics of Pygmy seahorse (Hippocampus bargibanti) from West Papua, Indonesia: Insights from mitochondrial DNA and RAD sequencing.
22.05.2025 13:54 —
👍 0
🔁 0
💬 0
📌 0
Combining fossil taxa with and without morphological data improves dated phylogenetic analyses https://www.biorxiv.org/content/10.1101/2025.04.07.646048v1
09.04.2025 15:32 —
👍 13
🔁 10
💬 0
📌 3
Maybe I'm too much of a negativist, but I've always disliked the 'de-extinction' field. It all looks like one big grift that attracts charlatans and pretenders, and which is forever making false claims and promises that distract people from what's actually interesting about extinct animals.
08.04.2025 10:31 —
👍 425
🔁 49
💬 29
📌 4

We de-extincted dire wolves.
Be honest.
We created a dire wolf.
Be honest.
It's a genetically modified puppy dog.
Thank you.
08.04.2025 15:57 —
👍 2700
🔁 695
💬 23
📌 26

Hemen gara jada!
¡Ya estamos aquí!
@upvehu.bsky.social @ztffct.bsky.social
19.03.2025 16:23 —
👍 12
🔁 5
💬 0
📌 1

👥 Esta semana mantuvimos nuestro encuentro anual con @cienciagob.bsky.social para hacer seguimiento de las acciones pasadas y futuras y seguir impulsando la investigación marina.
@embrcspain.bsky.social @embrc-eu.bsky.social @ulpgc.es @uvigo.bsky.social @pie-upvehu.bsky.social @upvehu.bsky.social
21.03.2025 11:47 —
👍 13
🔁 4
💬 0
📌 0
@caraotametalera.bsky.social maybe something you would love to check
12.03.2025 16:38 —
👍 1
🔁 0
💬 1
📌 0

Ostracod: Thalassocypria cumangulus n. sp.: 4976/08, holotype, female. Scale bar 100 μm
https://www.tandfonline.com/doi/full/10.1080/08912963.2017.1340471

Copepod: Ectinosomatidae (IHNFG–4899/Cop1); A1 = antennule, A2 enp = antennary exopod, A2 enp = antennary endopod, GDS = genital double-somite, AS = anal somite. Scale bar 200 μm. Photo from Francisco Vega

Isopod: Palaeospherarmadillo rotundus gen. nov. sp. nov., adult holotype (IHNFG-5302); General habitus, lateral view. Scale bar = 200 µm.
http://rmcg.geociencias.unam.mx/index.php/rmcg/article/view/639

Crab: Sesarmidae. Dorsal view of cephalothorax, specimen IHNFG-4991. LCh = left cheliped, P2-P5 = pereiopods, RCh = right cheliped. Scale 3 mm
http://boletinsgm.igeolcu.unam.mx/bsgm/vols/epoca04/6801/(6)Serrano.pdf
Aquatic crustaceans of MANY types are fossilized in amber from 23 mya Mexico, but amber comes from tree sap - how did crustaceans get there??? We think the fossils may come from estuarine mangroves. Shown here are ostracods, copepods, isopods, and even CRAB.
#Crustmas 🧪🦑
Paper list in next skeets
22.12.2023 16:12 —
👍 156
🔁 35
💬 10
📌 6
Genomics of experimental adaptive radiation in the cryptic coloration of feather lice https://www.biorxiv.org/content/10.1101/2024.12.20.629508v1
23.12.2024 02:33 —
👍 0
🔁 1
💬 0
📌 0
Genomic signals of local adaptation in Eleginops maclovinus from Northern Chilean Patagonia https://www.biorxiv.org/content/10.1101/2024.12.20.629640v1
23.12.2024 02:33 —
👍 0
🔁 1
💬 0
📌 0

A time-calibrated salamander phylogeny including 765 species and 503 genes
Recent time-calibrated amphibian phylogenies agree on the family-level relationships among extant salamanders but had disparate sampling regimes and i…
Do you need a salamander phylogeny? You're in luck! We put together the largest salamander phylogeny to date based on molecular markers and used more fossil calibrations than any other currently available trees. 765 salamander species! Check it out!
www.sciencedirect.com/science/arti...
18.12.2024 23:54 —
👍 194
🔁 72
💬 7
📌 5

Learn about #Biodiversity #Genomics #Europe -BGE Project- Case Studies here biodiversitygenomics.eu/citizen-scie... ... many of which participated in the @ergabiodiv.bsky.social Genome Applications Symposium www.erga-biodiversity.eu/post/genome-... - which you can watch on YouTube
15.12.2024 11:36 —
👍 11
🔁 5
💬 0
📌 0
This gorgeous little goober is a polychaete (bristle worm) larva from the family Serpulidae. I adore the big red eyes. For some reason they always remind me of the Stay Puft Marshmallow Man!
🦑
06.12.2024 19:40 —
👍 61
🔁 4
💬 2
📌 0


Not for sale, I guess, but this is a beautiful #nudibranch mobile by Tristan A. F. Long
#Phylogeny: onlinelibrary.wiley.com/doi/full/10....
07.12.2024 10:31 —
👍 15
🔁 5
💬 3
📌 0

Uneven distribution of prokaryote-derived horizontal gene transfer in fungi: a lifestyle-dependent phenomenon (mBio)
journals.asm.org/doi/full/10....
Fungi with different lifestyles have different levels of bacterial HGT - mycoparasites with the most and ectomycorrhizal fungi with the fewest:
09.12.2024 05:40 —
👍 59
🔁 23
💬 0
📌 1
Do you care about DEI in our Evolutionary Biology Community? We are looking for new members for our @sse-evolution.bsky.social diversity committee. Come join us!
05.12.2024 19:31 —
👍 11
🔁 11
💬 0
📌 0

Lineage of Crustacea - PhyloPic
Illustrated evolutionary lineage of Crustacea Brünnich 1772.
You can also celebrate with the evolutionary lineage of crustaceans. See where they came from! www.phylopic.org/nodes/a70684...
06.12.2024 22:20 —
👍 16
🔁 4
💬 1
📌 0

Diagnosing the affinities of early stem group crab fossils. Artistic reconstructions by Franz Anthony.
(A) Potential positions of early fossils following the favored phylogenomic hypothesis. Abbreviations p2, p3 refer to the second and third pereopods. Silhouettes from PhyloPic (phylopic.org), with fossils based on recent publications.
(B) Artistic reconstruction of partly carcinized Eoprosopon klugi as a facultative scavenger.
(C) Artistic reconstruction of uncarcinized Platykotta akaina as a dweller of Triassic bivalve reefs.
Since it's both #Crustmas and #FossilFriday, let's talk about the earliest crabs 220-190 million years ago. Unfortunately only 3 fossil species known. Probably their ancestors were not fully crabby, making carcinization more remarkable 🧪
onlinelibrary.wiley.com/doi/abs/10.1...
@franzanth.bsky.social
06.12.2024 16:33 —
👍 119
🔁 27
💬 3
📌 0

Long-lost ocean worms photobomb tiny seahorses, surprising scientists | CNN
Photographs of seahorses taken by scuba divers revealed evidence of a long-lost species of marine worm that hasn’t been seen since the mid-1950s, scientists say.
Tiny marine worms that live inside corals are tough to find and study in the ocean, so it's a good thing that their neighbors are incredibly photogenic. A type of bristle worm that hadn't been scientifically observed since 1956 was recently found in abundance—lurking in photos of pygmy seahorses 🧪
16.11.2024 16:02 —
👍 83
🔁 14
💬 2
📌 2



























