Amazonian blackwater lake (left), rainforest, and whitewater river. The whitewater river is laden with sediments from erosion of the Andes.
Photo Rhett A. Butler for Mongabay

Miocene fossil palm pollen grain (40 µm across) from the Pebas Wetland System. This species and others found with it are characteristic of lacustrine-estuarine conditions with wetland vegetation, peripheral forest, and a diverse aquatic fauna.

The Amazon pink dolphin Inia geoffrensis is derived from a marine species.
Michel Viard

Modern plants that are still distributed in the former marine incursion pathway which is thought to have left a geochemical imprint in the soils on top of the Solimões/Pebas formation. It has been shown that the bedrock and soils of the Pebas/Amazonas are much richer due either due to general chemistry or chemistry related to marine incursions.
🧪⚒️ Just posted: Carina Hoorn on the evolution of the Amazon basin. It was the rise of the Andes and the subsidence induced by the subduction under S. America's W coast that drove diversification. Listen and find out why. #geology #earth #Amazon
25.12.2025 10:10 —
👍 17
🔁 3
💬 0
📌 1
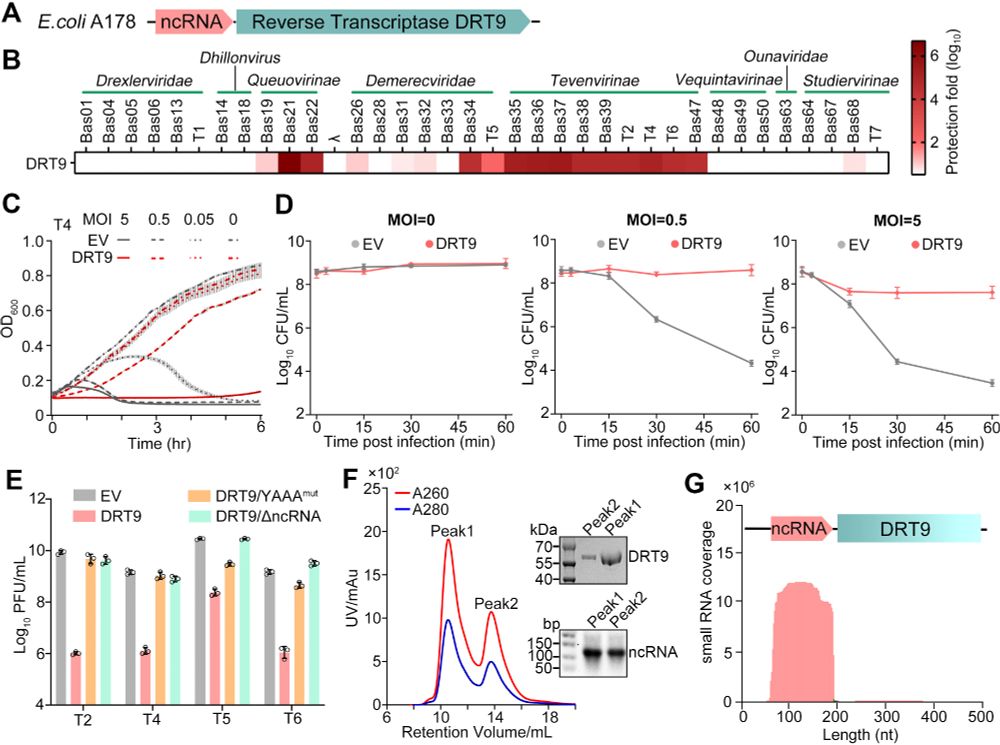
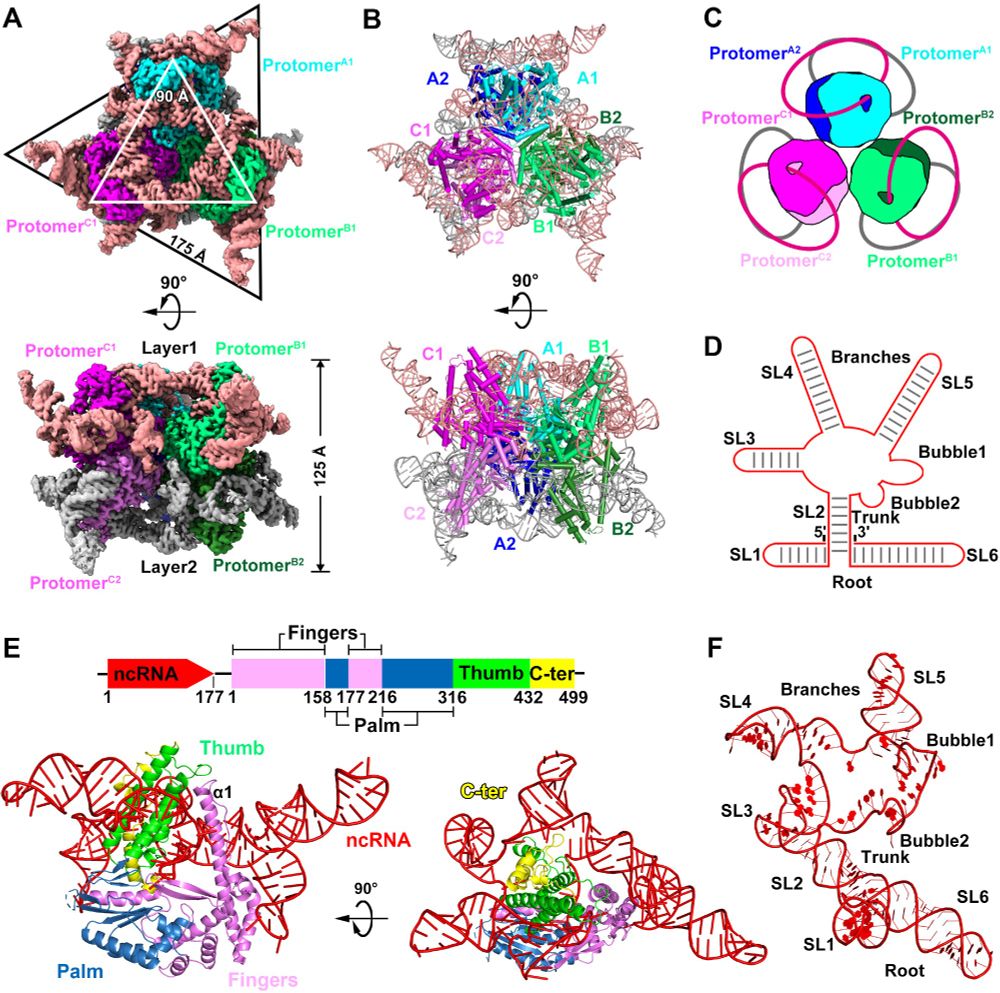
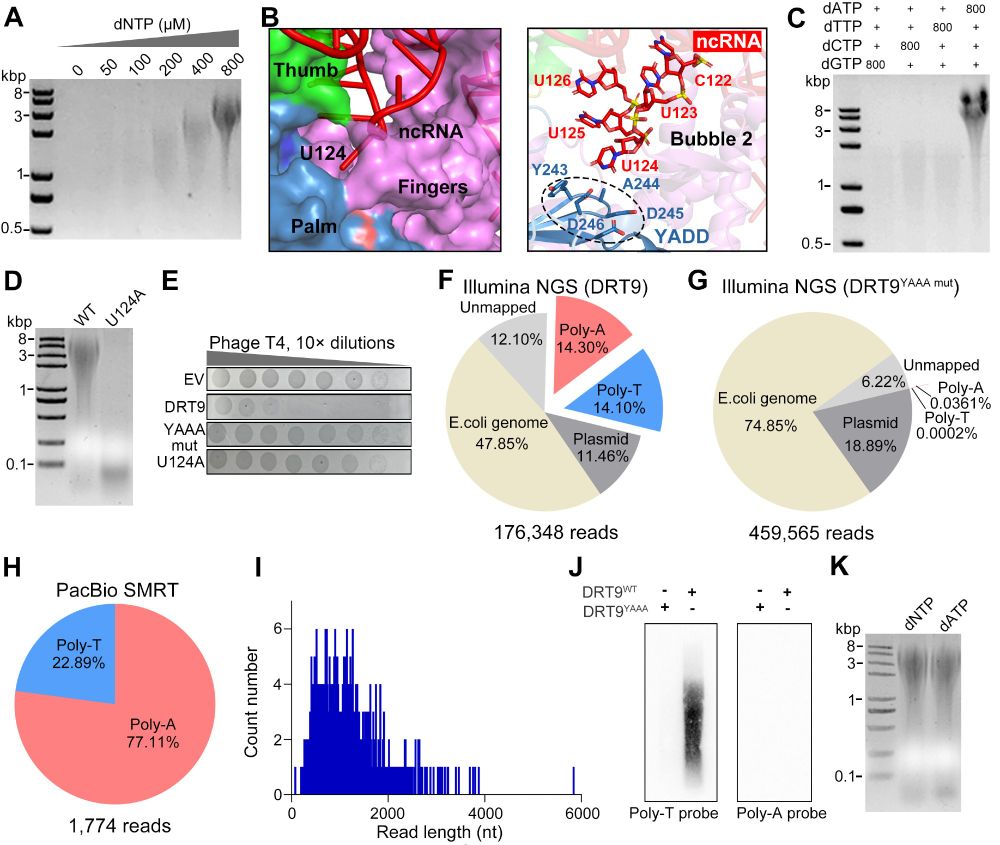
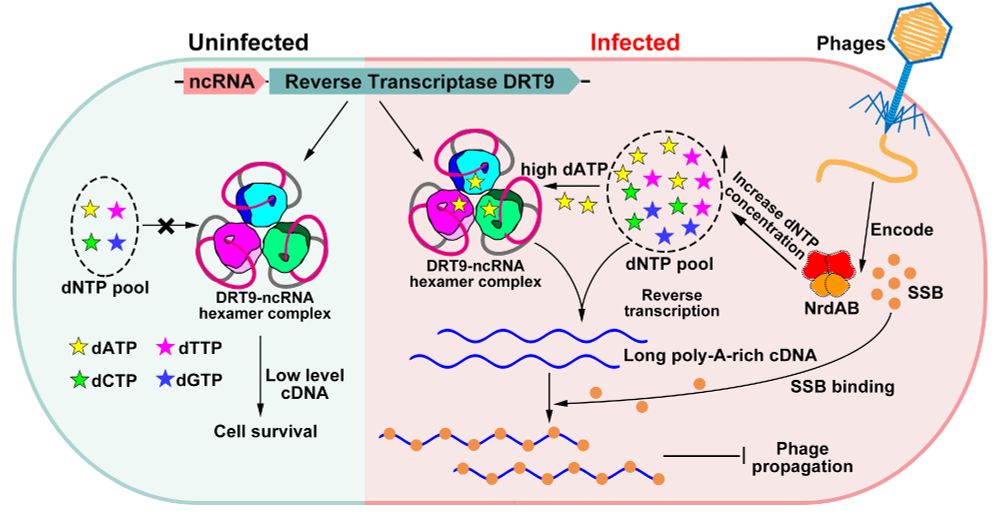
Fantastic work from Weiwei Yang in collaboration with Shuang yong Xu, Nan Dai and Ivan Correa
24.10.2025 12:56 —
👍 0
🔁 0
💬 0
📌 0
We also discovered new protein families potentially involved in dZ biosynthesis: DNA polymerases with enhanced dZ incorporation efficiency and nucleoside 2-deoxyribosyltransferases (NDTs) confirmed biochemically to catalyze dZ production.
24.10.2025 12:56 —
👍 0
🔁 0
💬 1
📌 0
Top hits matched known dZ biosynthesis enzymes, validating our hypothesis-free approach. Residue-level analysis pinpointed both known and novel amino acids in PurZ & YfbR-like dATPases, shedding light on substrate specificity.
24.10.2025 12:56 —
👍 0
🔁 0
💬 1
📌 0
Using metaGPA, we linked protein families to the dZ modification phenotype. This analysis revealed 116 protein families significantly associated with phages predicted to contain dZ in their DNA.
24.10.2025 12:56 —
👍 0
🔁 0
💬 1
📌 0
While representative dZ biosynthetic pathway have been studied in lab isolates, the diversity & distribution of dZ phages in natural environments is largely unknown. We therefore developed a restriction–based enrichment method to isolate dZ-containing phages directly from aquatic metagenomes.
24.10.2025 12:56 —
👍 0
🔁 0
💬 1
📌 0
Excited to share our new study using metagenomes to reveal key players in 2,6-Diaminopurine (dZ) Biosynthesis : We explore how some bacteriophages swap dA for 2-aminoadenine (dZ) in their genomes to evade bacterial defenses. #PhageBiology #Metagenomics #bacteriophage www.biorxiv.org/content/10.1...
24.10.2025 12:56 —
👍 3
🔁 1
💬 1
📌 0
A wonderful collaboration with Dr. Jackson Buss, Dr. Kyle Brumfield and Dr. Rita R. Colwell
13.10.2025 22:50 —
👍 2
🔁 0
💬 0
📌 0
This study demonstrates that it is possible to elicit and track functional responses directly within a microbiome, and to discover new enzymes in a hypothesis-free, reference-independent manner
13.10.2025 22:50 —
👍 1
🔁 0
💬 1
📌 0
By searching for protein domains that mirrored this behavior (rather than relying on homology), she found GH18 to be the most significantly associated family — the very domain found in known chitinases.
13.10.2025 22:50 —
👍 1
🔁 0
💬 1
📌 0
Chitinases were expected to increase during the bait phase and be rapidly downregulated after the switch — providing a measurable, dynamic phenotype within the native microbiome.
13.10.2025 22:50 —
👍 2
🔁 0
💬 1
📌 0
To make this possible, Colleen developed a “Bait-and-Switch” strategy:
first, she triggered a community succession toward chitin degraders (the bait), and then shifted the nutrient source to simple sugars (the switch).
13.10.2025 22:50 —
👍 1
🔁 0
💬 1
📌 0

So happy to share our new study
“A Bait-and-Switch strategy links phenotypes to genes coding for Polymer-Degrading Enzymes in Intact Microbiomes.”
In this work, we discovered novel chitinases directly on microbiome; a study brilliantly led by Dr. Colleen Yancey
www.biorxiv.org/content/10.1...
13.10.2025 22:50 —
👍 12
🔁 4
💬 2
📌 1
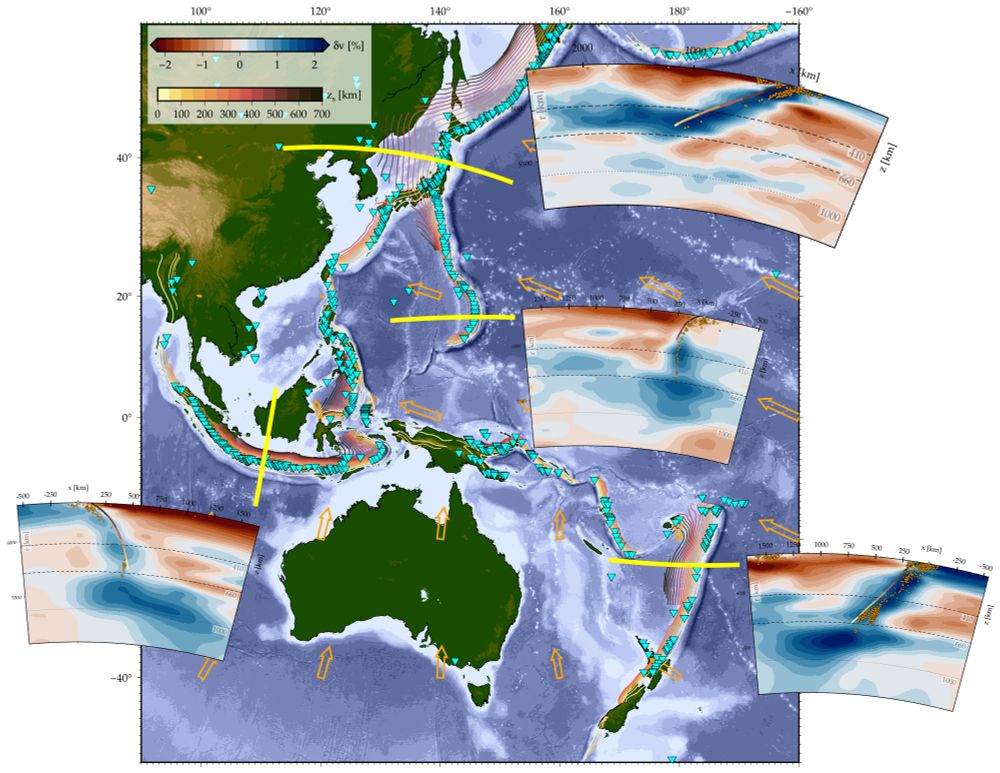
The figure shows sections of P-wave tomography anomalies along the yellow lines on the map (Fukao & Obayashi, 2013), seismicity (orange dots on profiles; Engdahl et al., 1998), plate velocities (orange vectors; Argus et al., 2011), and volcanoes (cyan inverted triangles; Siebert & Simkin, 2023). The Western Pacific and Japan subduction zones (the two most northerly zones) do not appear to penetrate the lower mantle, unlike those of Indonesia and Kermadec (north of New Zealand). Whether a slab penetrates the lower mantle depends on factors such as the duration of subduction and the angle at which the slab reaches the 660-km discontinuity, with steeper angles favoring descent.
Becker, T and Faccenna, C. (2025), Tectonic Geodynamics, Princeton University Press
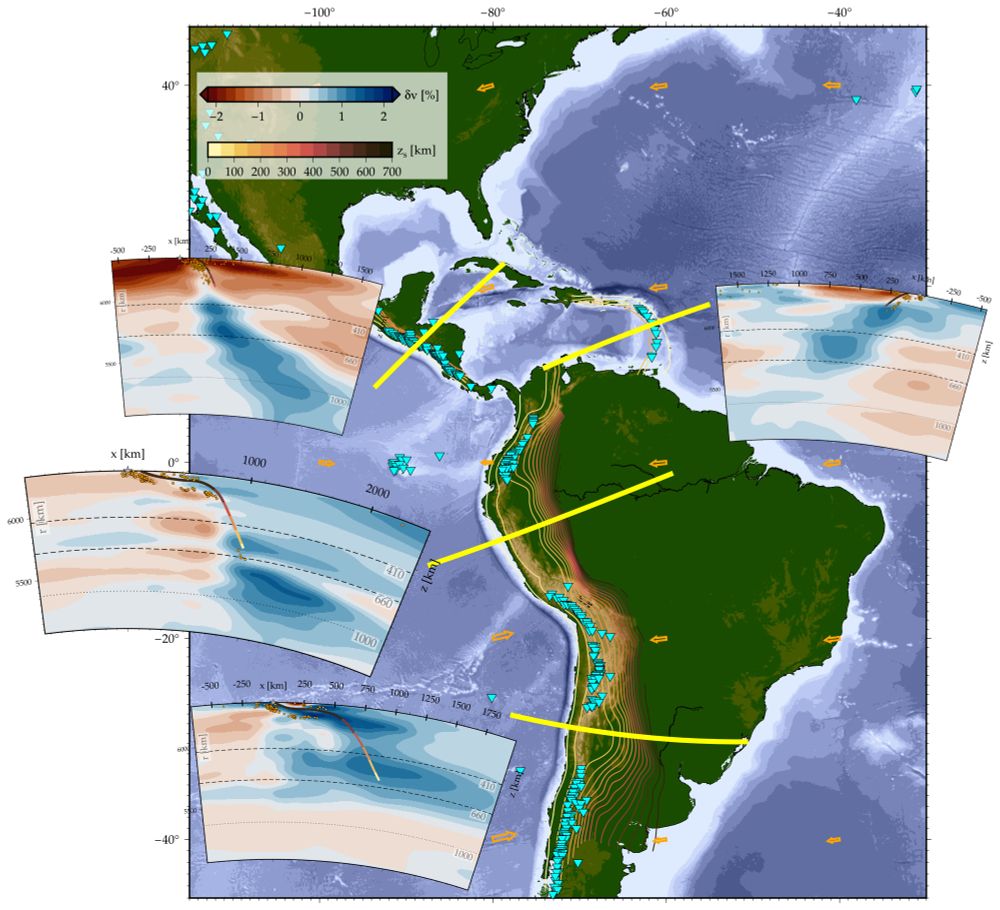
Subduction zones along the west coast of the Americas. In the podcast, Faccenna described how the slab subducting below the Andes has entered the lower mantle where its lateral movement is restricted by the higher viscosity there. The subduction zones beneath northern South America and Central America show evidence of penetration into the lower mantle, reaching depths of 1,300 km or more. By contrast, the younger Caribbean subduction zone does not.
Becker, T and Faccenna, C. (2025), Tectonic Geodynamics, Princeton University Press

Left: map showing seafloor ages in the western Pacific. The back-arc basins shown appear mainly as the red and orange regions, i.e., having the youngest ages of 0 and 30 million years. Center: more detailed seafloor age map of the Izu-Bonin-Marianas region. Right: P-wave tomography sections along the green lines labeled A, B, and C, revealing the varying shape of the subduction zone going from north to south.
The western Pacific subduction zone is characterized by back-arc extension in the overriding plate that began within the last 30 million years (left panel). Tomography and seismicity data show that along the Izu–Bonin region (section B1–B2), a double subduction system is present, resulting from subduction of both the Pacific Plate (to the east) and the Philippine Plate (to the west).
Becker, T and Faccenna, C. (2025), Tectonic Geodynamics, Princeton University Press

The Izu–Mariana subduction system is unique in that the trench is advancing toward the overriding plate (right panel). Bottom left: tomographic section along the red line on the map at right shows the two subducting slabs. Top left: numerical model of single subduction and double subduction. The double subduction simulation replicates quite a few features appearing on the tomographic section, such as the apparent flattening out of the subducting slabs between the depths of 400 and 600 km. In a single-slab system, the trench migrates backward, whereas in a double-slab system, the rear slab drives trench advance. The simulation also reproduces the tomographic section (bottom left, section along the red line on the map at right), which clearly shows the presence of two subducting slabs.
Faccenna, C. et al. (2018), Tectonophysics, 746, 229
🧪⚒️Just released an episode on the dynamics of subduction zones with Claudio Faccenna. Not only do trenches roll back and move laterally, they also advance and flip polarity. But when they penetrate the viscous lower mantle they get locked in place. Enjoy listening!
17.09.2025 16:24 —
👍 6
🔁 3
💬 0
📌 0
A novel NGS-compatible Enzymatic Strategy Enables Carryover Contamination Removal and Enhances Sequencing Performance. https://www.biorxiv.org/content/10.1101/2025.08.01.668201v1
03.08.2025 03:33 —
👍 1
🔁 1
💬 0
📌 0
Read the full story & explore the data here:
www.biorxiv.org/content/10.1...
👏 This is an amazing work from Amanda DeLiberto at #NEB @nebiolabs.bsky.social
#NGS #genomics #molecularbiology #preprint #contamination #sequencing
03.08.2025 18:07 —
👍 4
🔁 0
💬 0
📌 0
This dual-function strategy is ideal for:
🔹 Clinical & diagnostic sequencing
🔹 Low-input or precious samples
🔹 Metagenomics
🔹 Any workflow where contamination is a problem
03.08.2025 18:07 —
👍 0
🔁 0
💬 1
📌 0
Implementation is simple:
➕ Add Fpg & 7-deaza-dGTP directly to your PCR mix
⏱️ Brief pre-amplification incubation
No added steps, no workflow disruption
03.08.2025 18:07 —
👍 0
🔁 0
💬 1
📌 0
Why is this different?
7-deaza-dGTP is accepted by high-fidelity polymerases (unlike dUTP)
Fpg cleaves 7-deaza-dG-containing contaminants
Bonus: 7-deaza-dGTP reduces GC bias and improves sequencing coverage!
03.08.2025 18:07 —
👍 0
🔁 0
💬 1
📌 0
Our method changes that.
We introduce a new strategy using:
✅ 7-deaza-dGTP
✅ Fpg glycosylase
Together, they provide similar carryover contamination removal as dUTP/UDG and work seamlessly in standard NGS pipelines.
03.08.2025 18:07 —
👍 0
🔁 0
💬 1
📌 0
Contamination between NGS runs can compromise sensitive applications—from clinical diagnostics to ancient DNA.
Until now, enzymatic solutions like dUTP/UDG had one major flaw: they're are not compatible with high-fidelity NGS workflows.
03.08.2025 18:07 —
👍 0
🔁 0
💬 1
📌 0

Dream of avoiding carryover contamination in your NGS experiment?
Check out our latest preprint:
🧬 “A Novel NGS-Compatible Enzymatic Strategy Enables Carryover Contamination Removal and Enhances Sequencing Performance” www.biorxiv.org/content/10.1...
03.08.2025 18:07 —
👍 5
🔁 2
💬 1
📌 0
(Sorry for the late reply - I was hiking the Alpine section of the GR5 trail over the past six days)
27.07.2025 13:31 —
👍 1
🔁 0
💬 0
📌 0
Phages containing dU can be sequenced using polymerases that tolerate dU, such as Taq or Q5U. But thermostable polymerases are not strictly required, as only the first round of amplification needs to successfully copy the dU-containing templates to allow their propagation in downstream PCR cycles.
27.07.2025 13:31 —
👍 0
🔁 0
💬 1
📌 0
dU containing phages have been described (for ex : PMID: 34033748, PMID: 35934259) while Mikael Skurnik's lab isolated YerA41, a Yersinia ruckeri Bacteriophage that cannot be sequenced due to bulky modifications (PMID: 32517038).
27.07.2025 13:31 —
👍 0
🔁 0
💬 2
📌 0
And yes- standard high-throughput sequencing misses bulky DNA modifications and those recognized as damage, like deoxyuridine (dU), because most standard protocols use high-fidelity polymerases that typically contain a dU pocket and stall at dU.
27.07.2025 13:31 —
👍 0
🔁 0
💬 1
📌 0
This estimate comes from metaGPA for which high-throughput sequencing of DNA resistant to cytosine deamination is compared to total viral metagenomic DNA. Of course, only DNA that can be amplified (e.g., with Q5 polymerase) is detectable.
27.07.2025 13:31 —
👍 0
🔁 0
💬 1
📌 0

We estimate that 2–3% of contigs from viral metagenomes originate from genomes in which all canonical cytosines (C) have been replaced by modified bases (elifesciences.org/articles/70021). And that’s just for cytosine. (data from sewage and coastal metagenomes)
27.07.2025 13:31 —
👍 1
🔁 0
💬 1
📌 0
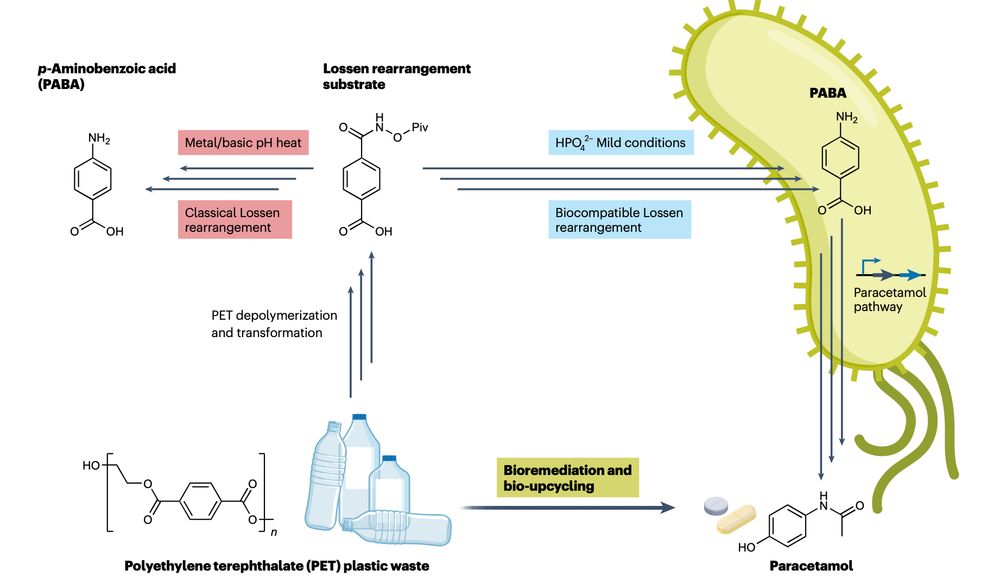
This is wild!
Engineering E. coli bacteria to turn plastic waste into paracetamol (Tylenol)
www.nature.com/articles/s41...
www.nature.com/articles/s41...
23.06.2025 18:54 —
👍 617
🔁 179
💬 17
📌 26