
I wrote about the recent autism-microbiome paper, why I think it's the most important microbiome paper this year, and what it says about the field
open.substack.com/pub/blekhman...

I wrote about the recent autism-microbiome paper, why I think it's the most important microbiome paper this year, and what it says about the field
open.substack.com/pub/blekhman...

Introducing Foli-seq in @natbiotech.nature.com today. Foli-seq can be used to measure gut-derived RNA from fecal matter (signals of immune, secretory & epithelial cells). We used this to map gut inflammation, drug response, and host-microbe interactions. 💩🧬💊🦠
www.nature.com/articles/s41...

Another banger from the @harriswang.bsky.social Lab this month!
“Fecal exfoliome sequencing captures immune dynamics of the healthy and inflamed gut” www.nature.com/articles/s41...
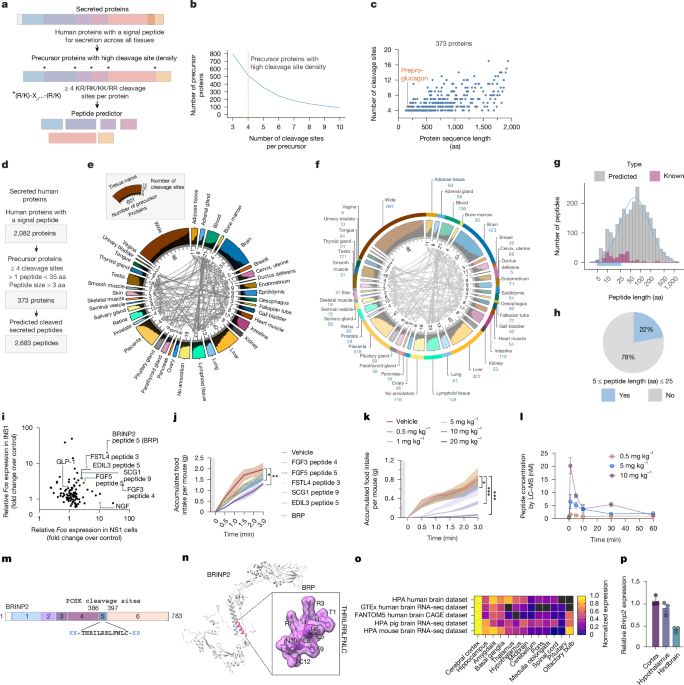
Exciting discovery by Svensson lab and collaborators @stanfordmedicine.bsky.social, computational discovery of 2500 new bioactive peptides, including 12-mer named BRP that reduces food intake leading to weight loss w/o nausea in mice!
www.nature.com/articles/s41...




Contributions from: Jeffrey Moffitt, @drmingyaoli.bsky.social, Qing Nie, Kazumasa Kanemaru, Sarah Teichmann, Daniel Dar, @ltkagohara.bsky.social, Shalev Itzkovitz, Roser Vento-Tormo, @fertiglab.bsky.social, @fabiantheis.bsky.social
23.02.2025 17:39 — 👍 4 🔁 2 💬 1 📌 0Mitosis?
14.12.2024 04:34 — 👍 1 🔁 0 💬 0 📌 0
Tabula Sapiens reveals transcription factor expression, senescence effects, and sex-specific features in cell types from 28 human organs and tissues https://www.biorxiv.org/content/10.1101/2024.12.03.626516v1
04.12.2024 13:31 — 👍 2 🔁 2 💬 0 📌 0New research from the Xavier lab at Broad reveals the gut's adaptability and immune-driven control in specific regions. A spatial map of the mouse intestines highlights its resilience to perturbations like inflammation, with implications for understanding diet, food allergies, and IBD in humans 🧬🦠
21.11.2024 15:12 — 👍 1 🔁 2 💬 1 📌 0Thank you Micha! 🙏
20.11.2024 13:38 — 👍 0 🔁 0 💬 0 📌 0We believe our approach marks a paradigm shift in microbiome science, integrating spatial context to uncover how the host interacts with the microbiome at relevant scales. Reach out if you want to try it yourselves
19.11.2024 22:59 — 👍 1 🔁 0 💬 0 📌 0
Finally, we explore a vignette of host-microbe interaction in a tumor-bearing tissue section, revealing microbial invasion into the host, possibly because misplaced mucus-producing cells leave the tissue unprotected, enabling generalized host-microbe interaction.
19.11.2024 22:59 — 👍 1 🔁 0 💬 1 📌 0
What excites me most: the achieved micron resolution lets us measure colony sizes for individual taxa, potentially linking them to their in vivo growth rates.
The cherry on top? We can identify colocalizing taxa and capture not just rRNA but also microbial genes!

Regarding the host, we retrieve both coding and non-coding signatures along the crypt-villus axis.
We observe high and diverse expression at villus tips, high % unspliced molecules near crypts, and high non-coding RNA percentages closer to the gut wall

Combining in situ polyadenylation with the StereoSeq platform, we reconstruct detailed maps of both cell types and microbial taxa at micron resolution, unlocking unprecedented insights into host-microbe interaction
19.11.2024 22:59 — 👍 0 🔁 0 💬 1 📌 0
Using Visium with in situ polyadenylation we explore microbial niches across the transverse and longitudinal axes of the GI
19.11.2024 22:59 — 👍 0 🔁 0 💬 1 📌 0
We first tested whether in situ polyadenylation enhances recovery of microbiome-derived RNA at low spatial resolution.
Applying our method across murine gut sections, we observed up to a 100-fold enrichment in microbial RNA while preserving host gene expression.

Building on previous work from @dwmckellar.bsky.social
, we combine a single enzymatic step of in situ polyadenylation with commercial spatial transcriptomics platforms—Visium and StereoSeq—to map host gene expression and microbiome in mouse gut tissue sections
Previous methods to map the interface, like imaging-based HiPR-FISH from our lab and sequencing-based with modified arrays like those developed by the Vickovic and Giacomello labs, capture target microbes but limit unbiased discovery or produce low-resolution maps.
Here we address these challenges.

Spatial structure shapes the host-microbiome interactome.
Our new method to map the host-microbiome interface at high-resolution, now on bioRxiv!
Co-led with Lena Takayasu & @dwmckellar.bsky.social collab between the De Vlaminck and Brito labs
doi.org/10.1101/2024...
A thread⬇️