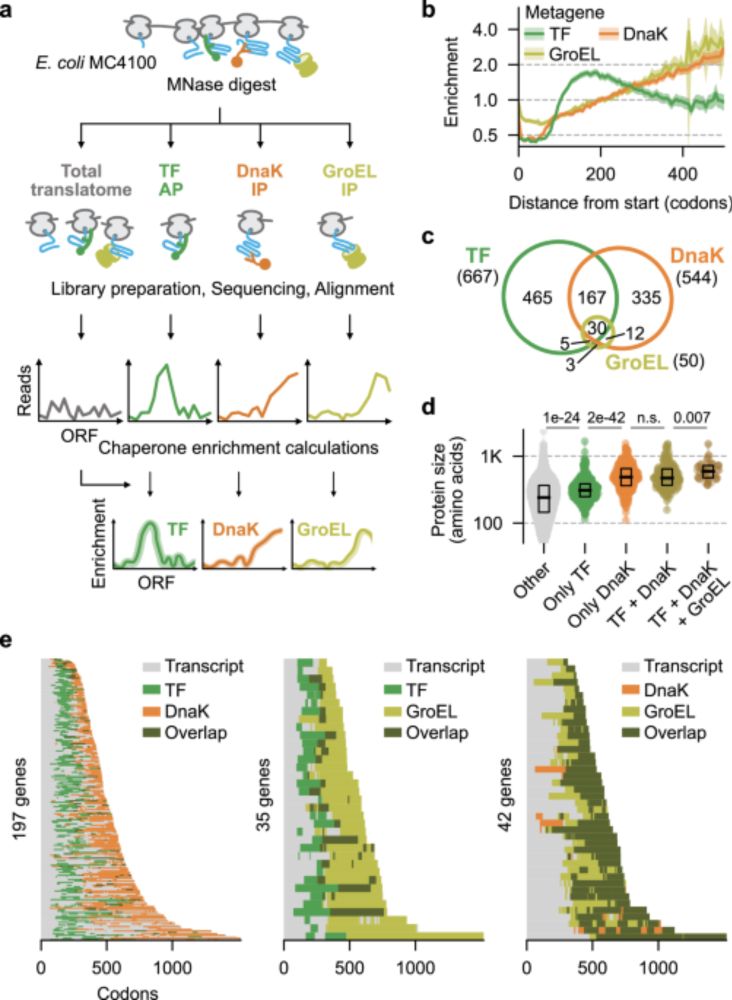
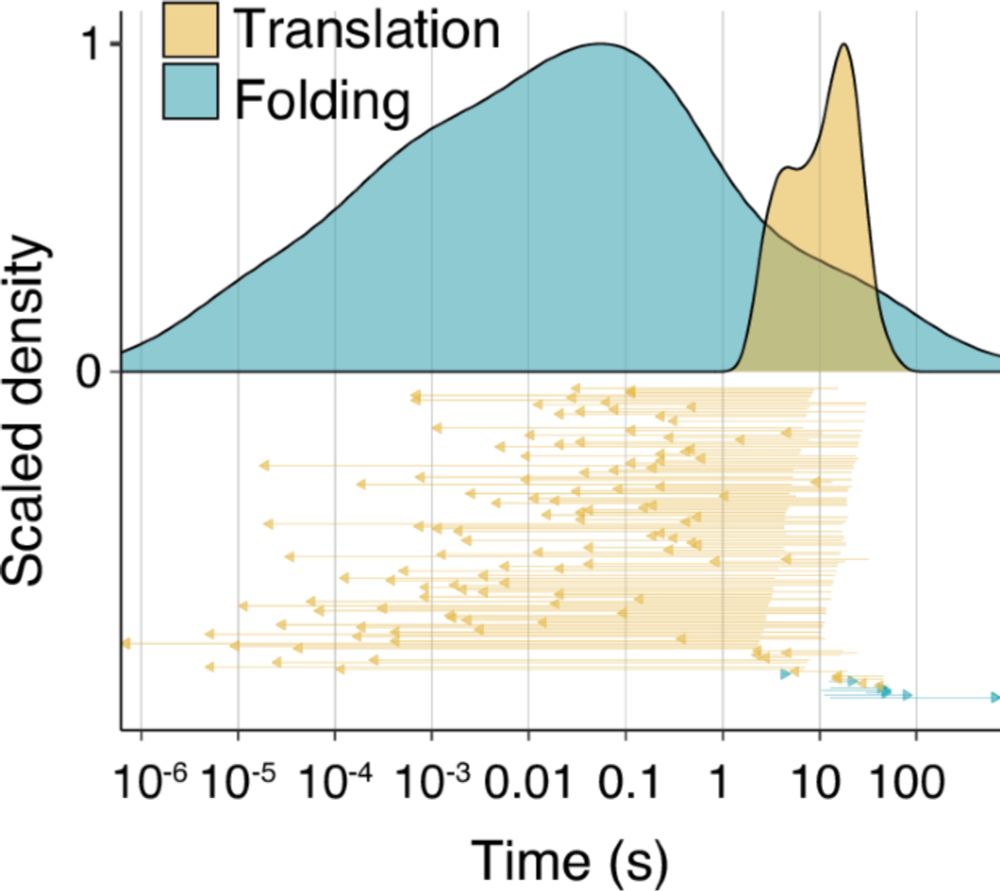
A new @MolecularCell paper shows that bacterial sRNAs can bind within coding sequences and enhance protein activity without affecting mRNA levels, pointing towards a new mechanism of co-translational folding regulation!
Could this be used as tool to induce translation pausing? 👀
#translation
06.05.2025 09:53 —
👍 1
🔁 0
💬 0
📌 0
Had a fantastic week at @embl.org to learn the ins and outs of ribosome profiling directly from some of the best in the field! Grateful to the amazing organisers, trainers, and fellow participants for such an enriching experience.
#ProteinTranslation #EMBORiboProf
31.03.2025 15:16 —
👍 1
🔁 0
💬 0
📌 0
Very excited to share our new preprint together with @daniedaaboul.bsky.social, where we studied the gene regulatory code that hippocampal granule cells (GCs) use during synapse formation (1/n)
31.03.2025 05:57 —
👍 15
🔁 8
💬 2
📌 0