
An unhealthy diet disrupts feeding behavior and the gut microbiota, but
whether early-life dietary effects persist, or can be restored later in life,
remains unclear. We investigated whether microbiota-targeted interventions
(FOS + GOS or Bifidobacterium longum APC1472) could restore early-life highfat/
high-sugar (HFHS) diet-induced feeding alterations in adult female and
male mice.
New APC paper in Nature Comms
First author Cristina Cuesta-Marti and colleagues examine how eating unhealthy foods early in life leaves lasting brain and feeding changes but bacteria can help restore healthy eating.
www.nature.com/articles/s41...
#Microbiome #GutBrainAxis
Tagging collaborators ⬇️
24.02.2026 11:06 —
👍 9
🔁 7
💬 1
📌 0
How does industrialization and lifestyle impact our interactions with gut microbiomes? And what is the consequence of these perturbations on our physiology?
Discover findings from our new study addressing these questions:
www.biorxiv.org/content/10.1...
#microbiome #globalhealth #GMbC
12.11.2025 15:45 —
👍 8
🔁 5
💬 1
📌 1
🚀 TaxSEA v1.2 is now live in #Bioconductor release 3.22!
TaxSEA brings enrichment testing to #metagenomics data. Just like GSEA does for transcriptomics making #microbiome & #microbiota results biologically interpretable, fast, and reproducible. We've added new DBs & features in 1.2 👇 🧵
03.11.2025 00:27 —
👍 7
🔁 4
💬 1
📌 0
Orchestrating Microbiome Analysis with Bioconductor https://www.biorxiv.org/content/10.1101/2025.10.29.685036v1
30.10.2025 20:54 —
👍 4
🔁 2
💬 0
📌 1

New paper out: Subspecies of the human gut microbiota carry implicit information for in-depth microbiome research.
29.08.2025 10:06 —
👍 15
🔁 3
💬 2
📌 1

Prehistoric Global Migration of Vanishing Gut Microbes With Humans
The gut microbiome is crucial for health and greatly affected by lifestyle. Many microbes common in non-industrialized populations are disappearing or extinct in industrialized populations. Understand...
Excited to share a new preprint w/ the Sonnenberg lab, led by Matt Carter, @zzzhiru.bsky.social & @mattolm.bsky.social. We analyzed the microbiomes of two non-industrialized populations from opposite sides of the globe to try to reconstruct the recent evolutionary history of our gut microbiota.
16.08.2025 18:25 —
👍 66
🔁 21
💬 2
📌 0
New preprint from lab: www.biorxiv.org/content/10.1... describing the Metalog database of manually annotated contextual data for >110k metagenomics samples around the globe metalog.embl.de
See the thread below from @biocs.bsky.social for more info!
15.08.2025 08:11 —
👍 12
🔁 2
💬 0
📌 0
Thank you so much!
13.08.2025 21:32 —
👍 0
🔁 0
💬 0
📌 0
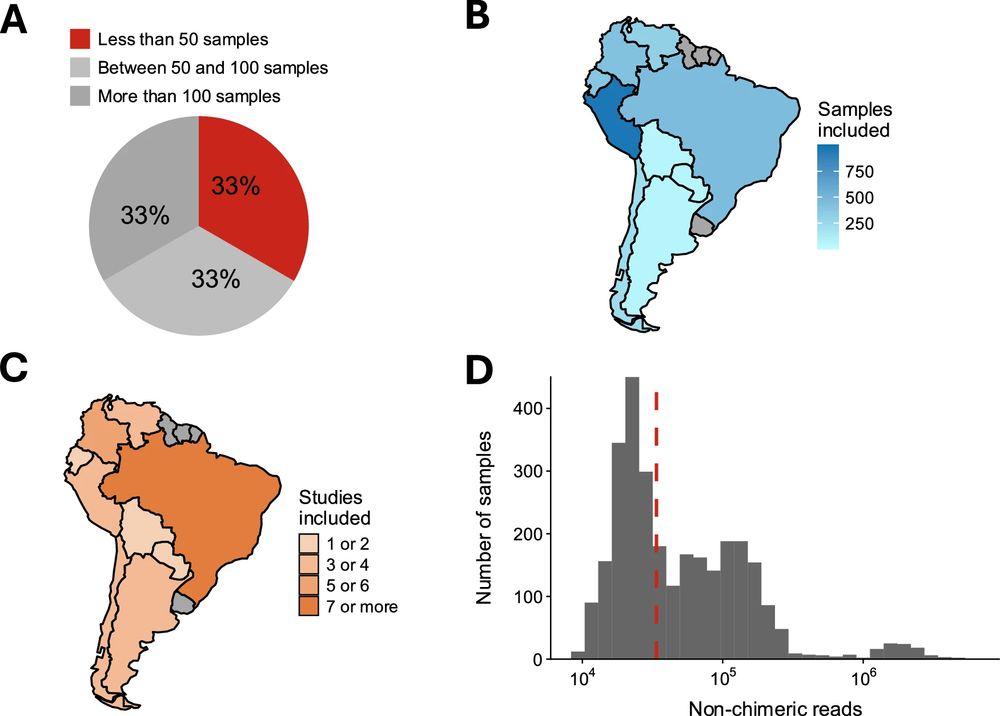
The South American MicroBiome Archive (saMBA): enriching the microbiome field by studying neglected populations. #Microbiome #NegletecPopulations @natcomms.nature.com 🧬
www.nature.com/articles/s41...
13.08.2025 09:15 —
👍 11
🔁 3
💬 1
📌 0
Thanks John for sharing the work. Also for the constant mentoring and support! It's deeply appreciated 🙌🔥
11.08.2025 17:34 —
👍 2
🔁 0
💬 0
📌 0
For more information about our work, see the thread I shared when the preprint was released: bsky.app/profile/bval...
11.08.2025 13:35 —
👍 4
🔁 1
💬 0
📌 0

Common Limitations of Gut Microbiome Meta-Analyses Undermine their Credibility
Although microbiome meta-analyses are fairly common in the literature, there is a lack of critical discussion on their limitations. This post is my (unsolicited) contribution to that topic.
I’ve been thinking about meta-analysis of microbiome data for a on-going collaboration when I noticed that, although they are fairly common in the literature, there is a lack of critical discussion on their limitations. This blog is my (unsolicited) contribution to the matter.
shorturl.at/mXSar
21.07.2025 14:24 —
👍 3
🔁 1
💬 0
📌 0

When Good Science Goes Wrong: How Batch Effects Can Obscure Biology
...and how to successfully avoid it
A blog post about batch effects in bioinformatics and how to address them
--
By me 🤓
shorturl.at/cmF8z
07.07.2025 12:18 —
👍 3
🔁 1
💬 0
📌 0
It's #WorldMicrobiomeDay! 🦠
We are proud to be a part of this exciting research area through contributions by our amazing authors.
To celebrate this occasion, here are some of our favourite microbiome papers.
Do you have a favourite of yours? let us know in the replies.
#MicroSky #MicrobiomeSky
27.06.2025 08:11 —
👍 25
🔁 9
💬 2
📌 1
Apparently it is still possible in 2025 to publish a paper in Nature that *filters* for strong signals before applying un-corrected significance tests. Seeing mistakes like this pass peer review is demoralizing.
17.06.2025 07:25 —
👍 39
🔁 8
💬 2
📌 0

My First Research Project Conducted in the Open
TL;DR I’ll start a biobliometrics analysis on the Microbiome-Gut-Brain axis field.
I started a blog: "The Middle Author's Syndrome" and I've just wrote my first post: bvalderrama.substack.com/p/my-first-r....
It's about a new side research project I just started. We will see if and how these experiments (the blog and the project) develop over time 😂.
11.06.2025 14:23 —
👍 1
🔁 0
💬 0
📌 0

Compendium Manager: a tool for coordination of workflow management instances for bulk data processing in Python
Compendium Manager is a command-line tool written in Python to automate the provisioning, launch, and evaluation of bioinformatics pipelines. Although workflow management tools such as Snakemake and N...
We wrote up the process we've developed for processing microbiome data in bulk! Workflow management tools are miraculous for processing a project with lots of samples, but when you have lots of *projects* too, as we do when pulling data from NCBI databases, it gets hard to juggle. #microbiomesky
19.05.2025 19:40 —
👍 8
🔁 5
💬 1
📌 1

April 11, 2025
Happy Friday and see you next week at MVIF! Gut microbiome The South American MicroBiome Archive (saMBA): Enriching the healthy microbiome concept by evaluating uniqueness and biodiversity of negle…
New #MicrobiomeDigest is OUT microbiomedigest.com/2025/04/10/a...
• The South American MicroBiome Archive / @bvalderrama.bsky.social
• coverM / @aroneys.bsky.social
• #OpenSourceDiaries / Anita Ihuman
• invitation for the next week @microbiomevif.bsky.social
& more.
Happy Friday!
11.04.2025 07:25 —
👍 5
🔁 7
💬 0
📌 0
Finally, I’d like to thank again everyone involved in this research: Paulina Calderón-Romero, Thomaz, @biothomaz.bsky.social, and of course to my supervisors: Aonghus Lavelle, Ger Clarke, and John @jfcryan.bsky.social. Also, thanks to the centre APC @apcmicrobiomeirel.bsky.social
09.04.2025 12:50 —
👍 3
🔁 1
💬 0
📌 0

GitHub - Benjamin-Valderrama/saMBA-pipeline: Workflow to build archives of 16s faecal microbiome samples from neglected populations.
Workflow to build archives of 16s faecal microbiome samples from neglected populations. - Benjamin-Valderrama/saMBA-pipeline
We show how semiautomated project screening can better represent regional biodiversities 🌎. The workflow is available for researchers building similar archives for other regions or body sites. The data generated is open, and new features are coming, so keep an eye on GitHub: shorturl.at/i9P1G (8/8)
09.04.2025 12:50 —
👍 3
🔁 0
💬 1
📌 0

We improved estimates of regional biodiversity and identified countries with the greatest potential to uncover more biodiversity in future sampling efforts—an analysis that’s the first of its kind and a critical guide for regions with limited research resources. (7/8).
09.04.2025 12:50 —
👍 3
🔁 0
💬 1
📌 0

After analysing >30 projects—most not included in any compendium before—we show how saMBA expanded our understanding of the human gut microbiome with samples from nearly every country in the region 🌎. We also found that nearly a third were likely discarded due to past sample size restrictions (6/8).
09.04.2025 12:50 —
👍 2
🔁 0
💬 1
📌 0
Why South America? Simple: it’s one of the regions with the fewest microbiome samples but some of the highest gut microbiome diversity among its inhabitants ☝️ (shorturl.at/qT1I6). Plus, I’m from Chile, so there's an emotional link 😂 (5/8)
09.04.2025 12:50 —
👍 2
🔁 0
💬 1
📌 0





















