Deeply grateful to the @simonsfoundation.org for launching SCENE and thrilled to join this 10-year journey into ecological neuroscience—unraveling how sensory and motor systems interact. Excited to collaborate with an incredible team of theorists and experimentalists working across species!
25.04.2025 16:17 —
👍 11
🔁 1
💬 0
📌 0
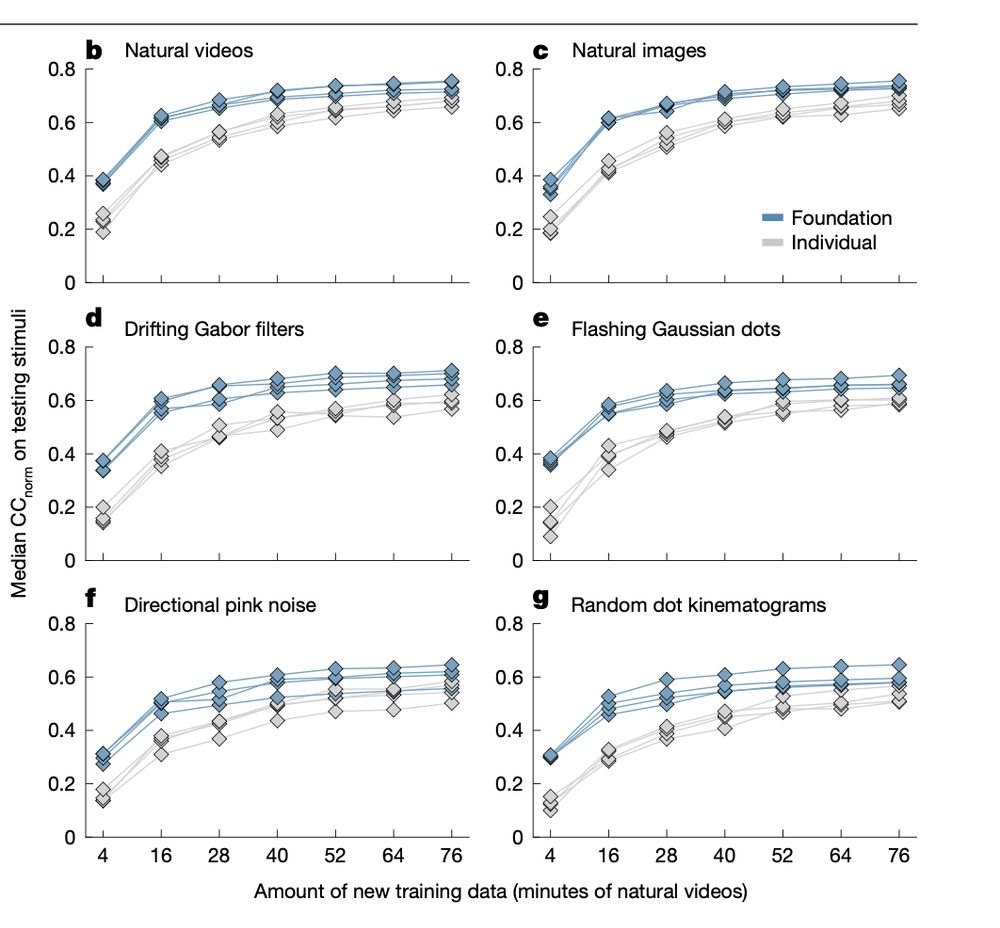
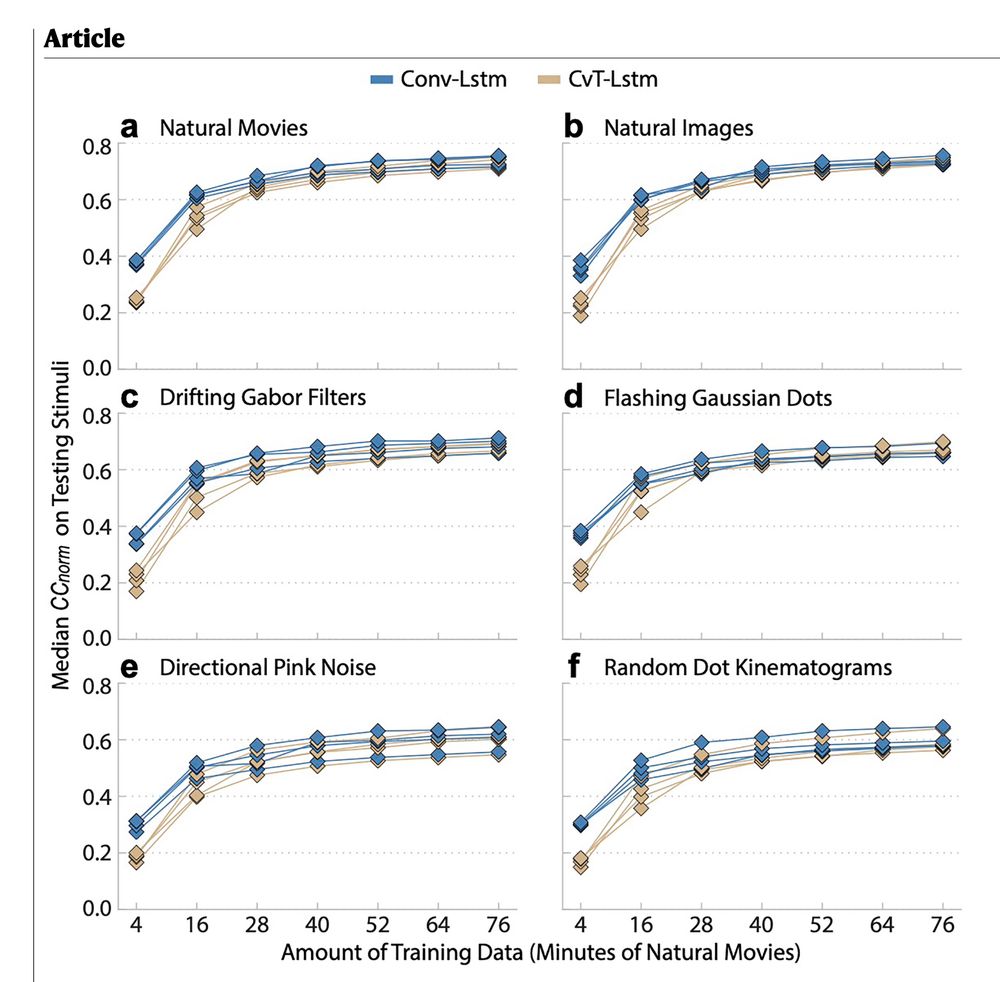
Building on Lurz et al., our new Wang et al. studies movie‑data performance vs. training size and compares scaling for Conv‑LSTM vs. CvT(convolutional vision
transformer)‑LSTM. Details: www.nature.com/articles/s41...
19.04.2025 13:21 —
👍 3
🔁 0
💬 0
📌 0
A super exciting paper by @aecker.bsky.social and Marissa Weis, part of the #MICrONS package, deriving a set of principles to characterize the morphological diversity of excitatory neurons across cortical layers.
www.nature.com/immersive/d4...
15.04.2025 11:37 —
👍 10
🔁 0
💬 0
📌 0

We didn't know the optimal patterns driving mouse V1 neurons until the deep learning model by Walker et al. (2019). FYI: Unlike mice, Gabors actually describe macaque V1 neurons quite well (Fu et al., Cell Reports).
13.04.2025 19:11 —
👍 1
🔁 0
💬 1
📌 0
MICrONS represents a huge step forward for the field. Big-data and AI will drive the next wave of discoveries in neuroscience
13.04.2025 18:48 —
👍 17
🔁 2
💬 1
📌 0
3/3 The core strength of our approach—robust prediction of neural responses to novel visual stimulus domains. Dyer's autoregressive approach generates latent embeddings for neural decoding—an entirely different architectural paradigm with different scientific objectives.
13.04.2025 13:57 —
👍 0
🔁 0
💬 1
📌 0
2/3 However this is not the main point, these models serve fundamentally different purposes. Ours explicitly predicts neural responses to visual stimuli (an encoding model), creating functional digital twins.
13.04.2025 13:57 —
👍 0
🔁 0
💬 1
📌 0
1/3 Just for clarification our foundation model was introduced on March 21st, 2023—predating Dyer et al. by over six months.
www.biorxiv.org/content/bior...
13.04.2025 13:57 —
👍 1
🔁 0
💬 1
📌 0
2
13.04.2025 13:54 —
👍 0
🔁 0
💬 0
📌 0
8/8 Deep learning simulation enables systematic representational-level characterization, though detailed circuit-cell-type-level mechanistic comprehension remains beyond current capabilities in the cortex.
13.04.2025 13:54 —
👍 1
🔁 0
💬 2
📌 0
7/8 and characterization of the feature landscape of mouse visual cortex (Tong et al., bioRxiv 2023)—just a few examples of their applications. Most importantly, they yield in silico predictions which are subsequently verified through experimental testing.
13.04.2025 13:54 —
👍 0
🔁 0
💬 1
📌 0
6/8 Predictive models also enabled to systematically characterize single neuron invariance properties (Ding et al., bioRxiv 2023), center-surround interactions (Fu et al., bioRxiv 2023), color-opponency mechanisms (Hofling et al., Elife 2024),
13.04.2025 13:54 —
👍 0
🔁 0
💬 1
📌 0
5/8 Our models also revealed that mouse V1 neurons shift their selectivity toward UV when pupil dilation or running begins, despite maintaining stable spatial stimulus structure—discovered in the digital twin and validated experimentally in closed-loop studies (Franke et al., Nature 2022).
13.04.2025 13:54 —
👍 2
🔁 0
💬 1
📌 0
4/8 For example, these simulations revealed that mouse V1 neurons exhibit complex spatial features deviating from the common notion that Gabor-like stimuli are optimal (Walker, Sinz et al., Nature Neuroscience 2019).
13.04.2025 13:54 —
👍 0
🔁 0
💬 2
📌 0
3/8 When ANNs accurately simulate neural function, they facilitate 'mechanistic interpretability' (to borrow the AI term)—enabling rigorous representational-level analysis of neuronal tuning.
13.04.2025 13:54 —
👍 0
🔁 0
💬 2
📌 0
2/8 Moreover, both task- and data-driven neural predictive models are powerful tools to gain neuroscientific insights as we and others have demonstrated repeatedly.
13.04.2025 13:54 —
👍 0
🔁 0
💬 2
📌 0
1/8: This quote from our abstract refers to task-driven modeling approaches (e.g., Yamins, DiCarlo, et al.) which define computational objectives and reveal hidden representations closely matching brain activity—widely recognized for deepening insights into brain computations.
13.04.2025 13:54 —
👍 0
🔁 0
💬 2
📌 0
3/3 The core strength of our approach—robust prediction of neural responses to novel visual stimulus domains. Dyer's autoregressive approach generates latent embeddings for neural decoding—an entirely different architectural paradigm with different scientific objectives.
13.04.2025 13:47 —
👍 1
🔁 0
💬 1
📌 0
2/3 However this is not the main point, these models serve fundamentally different purposes. Ours explicitly predicts neural responses to visual stimuli (an encoding model), creating functional digital twins.
13.04.2025 13:47 —
👍 0
🔁 0
💬 1
📌 0
1/3 Just for clarification our foundation model was introduced on March 21st, 2023—predating Dyer et al. by over six months.
www.biorxiv.org/content/bior...
13.04.2025 13:47 —
👍 1
🔁 0
💬 1
📌 0
Huge thanks to @IARPAnews for funding this groundbreaking effort through the @BRAINinitiative, and to our amazing team at
@stanforduniversity.bsky.social @stanfordmedicine.bsky.social @BCM @Allen @Princeton @unigoettingen.bsky.social
#MICrONS #NeuroAI #Connectomics #FoundationModels #AI
10.04.2025 23:46 —
👍 5
🔁 0
💬 0
📌 0
Foundation models offer a powerful way to systematically decode the neural code of natural intelligence, bridging the gap between brain structure and function.
10.04.2025 23:46 —
👍 2
🔁 0
💬 1
📌 0
Instead, they preferentially connect based on shared functional tuning, choosing partners with similar feature selectivity (“what”) rather than merely receptive field overlap (“where”).
10.04.2025 23:46 —
👍 1
🔁 0
💬 1
📌 0
Crucially, this robust generalization allowed us to create precise functional digital twins of individual mouse brains, combining functional predictions with known anatomical wiring.
10.04.2025 23:46 —
👍 1
🔁 0
💬 1
📌 0
Our foundation model generalized robustly to new neurons, new animals, and even previously unseen stimulus domains. It also accurately predicted entirely new modalities, such as anatomically defined cell types.
10.04.2025 23:46 —
👍 1
🔁 0
💬 1
📌 0