Discover and share the CBS Montpellier starter pack, to follow current and formers members of the Centre de Biologie Structurale on Bluesky.
go.bsky.app/G682pvP
@cnrs.fr
@inserm.fr
@umontpellier.bsky.social
Discover and share the CBS Montpellier starter pack, to follow current and formers members of the Centre de Biologie Structurale on Bluesky.
go.bsky.app/G682pvP
@cnrs.fr
@inserm.fr
@umontpellier.bsky.social
This graph basically shows that the MSCA are, as of now, useless unless the EU governance does something about it 🧪
11.02.2026 09:42 — 👍 21 🔁 8 💬 3 📌 0
The killing of Alex Pretti is a heartbreaking tragedy. It should also be a wake-up call to every American, regardless of party, that many of our core values as a nation are increasingly under assault.
25.01.2026 17:39 — 👍 60226 🔁 19558 💬 3137 📌 1541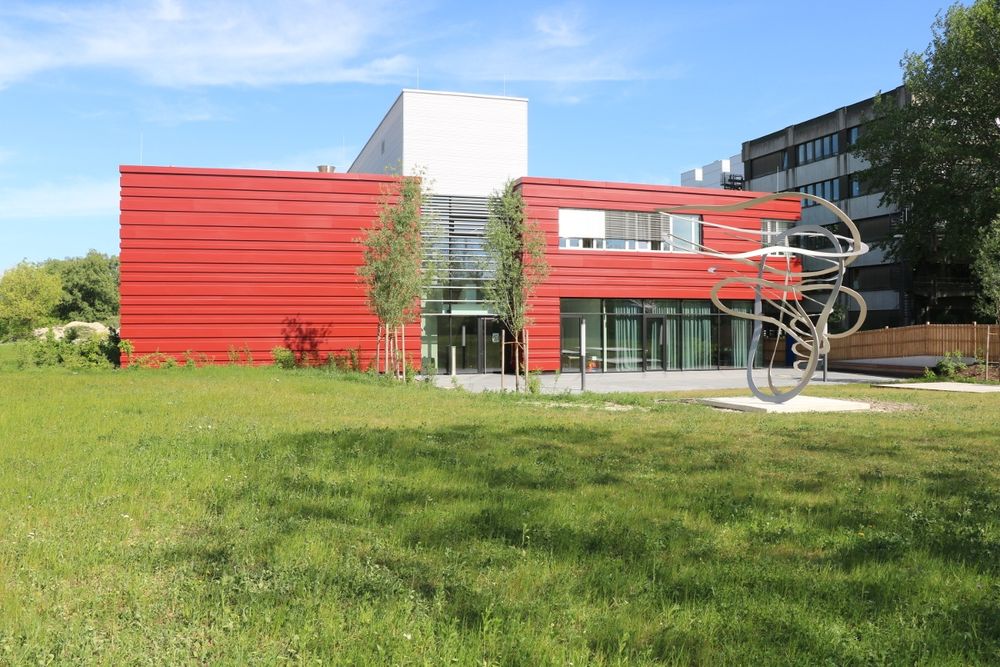
EMBO Practical Course: Structural dynamics, interactions & function of biological macromolecules by NMR
EMBO Practical Course: Structural dynamics, interactions & function of biological macromolecules by NMR @embo.org
📅 July 24–31, 2026
📍 BNMRZ, Technical University of Munich (Garching)
📝 Registration open | Deadline: Apr 17, 2026
🔗 embo2026.bnmrz.org
#NMR #StructuralBiology #EMBO #NMRchat 🧲
Join us in Montpellier in June 2026 for the second edition of the European South Atlantic Biophysical conference
#ESAB2026
The latest article from a great collaboration between the Buchner Lab, @sattler-lab.bsky.social and the Rosenzweig Lab!
07.01.2026 11:26 — 👍 1 🔁 0 💬 0 📌 0
Schematic showing the function of Sgt1 in yeast.
Our latest work is now out in @cp-molcell.bsky.social !
We have looked at Sgt1, an essential Hsp90 co-chaperone in yeast. We determined its interaction with Hs90 and showed that it stabilises the chaperone-client complex and prevents its dissociation by Aha1.
www.cell.com/molecular-ce...

Online Now: The essential co-chaperone Sgt1 regulates client dwell time in the Hsp90 chaperone cycle Online now:
25.12.2025 16:19 — 👍 6 🔁 3 💬 0 📌 0
Congrats to our lab member Philipp, who defended his PhD this week! 🎓🎉
05.12.2025 16:10 — 👍 6 🔁 2 💬 0 📌 0Watch Robert Quast @rbquast.bsky.social from our Team presenting our work on single molecule #FRET on #GPCR in the first FBI Connect webainar !
04.12.2025 19:20 — 👍 3 🔁 3 💬 0 📌 0
We are hiring a 2-year postdoctoral researcher to join our Membrane Biophysics team at the CBS in Montpellier. Join us to reinvent Atomic Force Microscopy to image the most common interface on our planet: the surface of water @membranesbiophy.bsky.social @cbsmontpellier.bsky.social
25.11.2025 13:04 — 👍 11 🔁 11 💬 0 📌 2
graphical abstract showing the combination of techniques used to analyze the dynamics of a de novo enzyme.
Our latest work is out! We combined #NMR with #Crystallography, #MD, and #ProteinMPNN to analyze the conformational dynamics of a de novo enzyme and redesigned it! Great collaboration between
@cathleenzeymer.bsky.social and the @sattler-lab.bsky.social.
www.cell.com/structure/fu...
Don’t miss this opportunity to work with a fantastic mentor, a great team, in a beautiful city of France!
07.11.2025 12:47 — 👍 9 🔁 4 💬 0 📌 0
Misfolded proteins can cause #Alzheimers, #Parkinsons and #Cancer. Johannes Buchner, part of the CHAPEROME team, receives an #ERC Synergy Grant to study how chaperone #proteins keep cells healthy: go.tum.de/904052 👏
#ERCSyG #biotechnology
@erc.europa.eu
📷A. Heddergott
We are hiring a PhD student!!
The project starts as early as January 2026 and is fully funded for three years.
Everyone with interest in NMR-centered structural biochemistry of nucleic acid-protein interactions in virus-infected cells, get in contact!
biochemie.uni-greifswald.de/forschung/fo...

Lancement ce jour du projet d'extension de notre laboratoire. 1100m² de nouveaux espaces et 280m² rehabilités.
Merci pour leur soutien à nos tutelles et nos partenaires.
@inserm.fr @cnrs.fr @umontpellier.bsky.social
@occitanie.bsky.social @villedemontpellier.bsky.social
& l'Etat
Thrilled to share my first corresponding author paper as a project leader in @hillerlab.bsky.social Amazing work from first author @annaleder.bsky.social We discovered multi-chaperone condensates in the ER that revolutionise the current vision of protein folding!
www.nature.com/articles/s41...

BNMRZ Symposium Ultra-Highfield NMR and 4D Structural Biology – From Mechanisms to Therapies, November 6–7, 2025, TUM Institute for Advanced Study, Garching @ias-tum.bsky.social registration opens soon www.bnmrz.org/4dstructbiol #BNMRZ #NMRchat #NMR 🧲
04.08.2025 20:14 — 👍 10 🔁 4 💬 0 📌 1
New paper in collaboration with the Förstemann Lab & Boekhoven Lab is out! Combining #NMR & mutational analysis, we characterize the unique features of the #Ago2 IDR region which drives Ago2/RNA #condensate formation.
doi.org/10.1093/nar/...
@clarahipp.bsky.social
J'entends Aurore Bergé sur @franceinfo.fr expliquer que les gens sont contres la loi Duplomb parce qu'ils sont désinformés.
Désinformés par qui ? 14 sociétés savantes ? La ligue contre le cancer ? Le CNRS écologie environnement ?
Very happy to share our preprint on heavy metal associated (HMA) domain-containing rice proteins bound by the rice blast MAX effector AVR-Pia! 🌾
15.07.2025 05:34 — 👍 13 🔁 6 💬 0 📌 0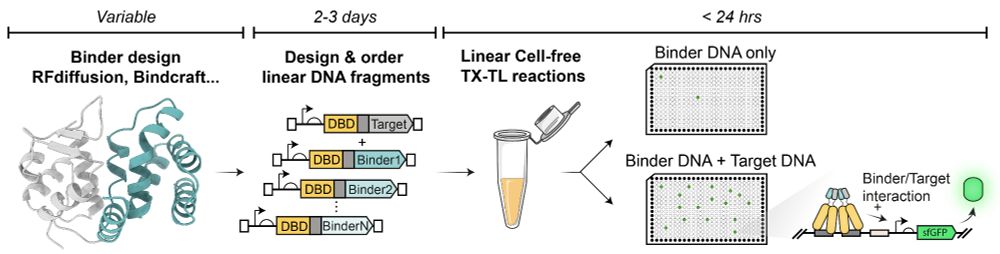
You designed binders for your favourite protein and wish there was a way to experimentally screen them within 24h w/ only a set of pipettes and a plate reader?
Check out our Cell-Free 2-Hybrid approach (CF2H)
Full post: tinyurl.com/48cz5nb6
Preprint: www.biorxiv.org/cgi/content/...



One of our group members @clarahipp.bsky.social had the great opportunity to attend the Nobel Laureate Meeting last week in sunny Lindau ☀️
#lino25

Read our preprint related to an important topic: climate change 👉 mr.copernicus.org/preprints/mr...
As scientists, we rely on data for rational decisions. Climate science is clear: human-made CO₂ emissions are changing our climate fast. But what can we do about it—individually or as a community? 1/n

📢 Paper with @sattler-lab.bsky.social now published in @narjournal.bsky.social ! doi.org/10.1093/nar/... We combine #NMR, #SAXS, and #MD to construct conformational ensembles, showing that A-to-I hyper-editing leads to significant structural dynamics. MD simulations by @piompons.bsky.social
26.06.2025 14:36 — 👍 13 🔁 6 💬 0 📌 0
We are happy to announce the “10th European solid-state NMR summer school” (October 19-24, 2025) that will take place at the Technical University Munich (@TUM.de) in Garching, Germany. #NMRchat #BioNMR
For more detailed information, please follow the link:
www.bio.nat.tum.de/ocb/upcoming...

"a fascinating case study on the limits of AI in biology and the harms of current publishing incentives" rachel.fast.ai/posts/2025-0...
04.06.2025 15:53 — 👍 93 🔁 45 💬 1 📌 5Thrilled to share our new method in-cyclo NMR that lets us watch molecular machines in action — decoding molecular machine kinetics with atomic precision and exceptional time resolution in one go! 🚀 Check it out: doi.org/10.1038/s414... #NMR #StructuralBiology
02.06.2025 09:16 — 👍 23 🔁 5 💬 4 📌 0
A new paper from the CBS !
Structural and Functional Characterization of the 28 kDa Structured Core of BmSA1, the Major Surface Antigen of Babesia Microti
onlinelibrary.wiley.com/doi/10.1002/...
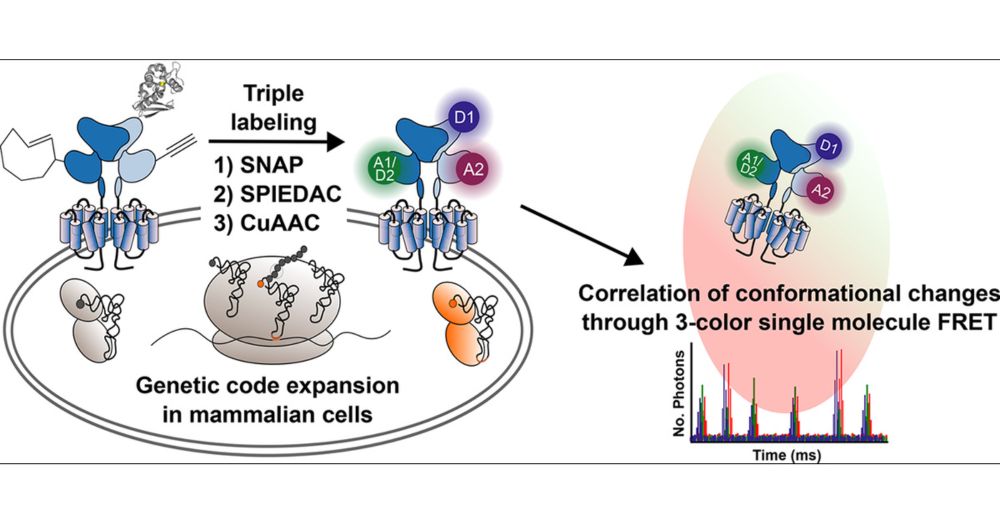
Our new study is out !
We introduce a triple labeling strategy (2 non canonical Amino Acids + 2 click chemistries and 1 SNAP-tag) to triple label a G-protein coupled receptor.
Open access in JACS
pubs.acs.org/doi/10.1021/...