New pre-print with @wtmatlock.bsky.social!!!
www.biorxiv.org/content/10.6...
What shapes the distribution of plasmids across bacteria? Our paper shows that conjugative plasmids actually have a very narrow distribution compared to mobilizable plasmids. Conjugative systems restrict plasmid transfer!
20.02.2026 08:41 —
👍 31
🔁 15
💬 1
📌 0
Bacteria chromosomes contain Genomic Islands that provide virulence, antibiotic resistance, MGE-defence,... They transfer between cells, but the mechanism of most remains elusive.
Here we explore the conjugative capacity of these mysterious Genomic Islands.
www.biorxiv.org/content/10.6...
14.01.2026 10:14 —
👍 82
🔁 52
💬 4
📌 2
test 3h
venae also produces a clinician-friendly HTML results report summarizing sequencing quality, organism identity, and AMR determinants. See the example report here:
phac-nml.github.io/venae/tests/...
🧵4/4
16.12.2025 15:38 —
👍 0
🔁 0
💬 0
📌 0

Workflow map of the venae pipeline https://github.com/phac-nml/venae
We built venae, a pipeline written in #Nextflow, which removes host reads, assess quality, identifies species, performs assembly, detects #AMR genes, and runs species-specific typing
🧵3/4
github.com/phac-nml/venae
16.12.2025 15:38 —
👍 0
🔁 0
💬 1
📌 0
Our turnaround time was around 4 h after blood cultures flagged positive - much faster than conventional methods! We were fortunate to obtain a lot of clinical samples (n=307🩸) which allowed us to design a really robust workflow
🧵2/4
16.12.2025 15:38 —
👍 0
🔁 0
💬 1
📌 0

Releases · tseemann/shovill
⚡♠️ Assemble bacterial isolate genomes from Illumina paired-end reads - tseemann/shovill
💾 Shovill 1.4.1 has been released!
The best way to de novo assemble microbial genomes from Illumina FASTQ.
Major fixes to the SKESA module, plasmid mode for the Spades module, and more error checking.
#bioinformatiocs #genomics #microbiology
github.com/tseemann/sho...
13.12.2025 21:15 —
👍 69
🔁 25
💬 0
📌 2
To summarise our recent pre-print: Autocycler, the automated consensus assembler, when used with Nanopore long-read only Enterobacterales assemblies, produces more complete chromosomes and plasmids, with an accuracy comparable to hybrid assemblies.
29.09.2025 06:57 —
👍 26
🔁 16
💬 1
📌 0
Check out our ChroQueTas tool (github.com/nmquijada/Ch...) and get ready to screen AMR in your fungal genomes by using FungAMR info!
18.08.2025 20:03 —
👍 4
🔁 4
💬 0
📌 0
This makes it incredibly difficult to track NDM plasmids across the country for surveillance purposes! Long-read sequencing is recommended for resolving NDM plasmid structure, cluster membership, and potential transposon-mediated spread in future studies
8/8
06.08.2025 13:21 —
👍 0
🔁 0
💬 0
📌 0
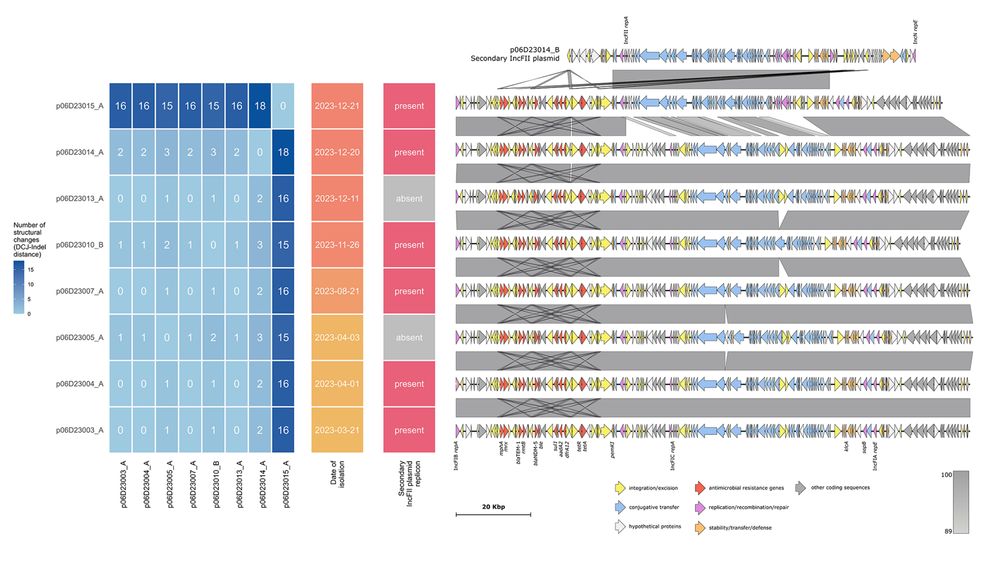
Heatmap and genetic structure of NDM plasmids from a single site site
We searched for trends at specific sites and found examples of how a plasmid can be stable and maintain its structure for months, and also how a single event (recombination with a co-resident plasmid) can dramatically change plasmid structure and consequently classification by MOB-suite/pling
7/8
06.08.2025 13:21 —
👍 0
🔁 0
💬 1
📌 0

Plasmid map depicting OXA and NDM genes
We found 11.4 % of isolates encoded both NDM and OXA-48 in the same cell. Nine plasmids encoded both NDM and OXA-48-type genes on the same backbone, which haven't been seen much in the lit yet. Five were in Kleb pneumo ST16 on an IncX3 backbone
6/8
06.08.2025 13:21 —
👍 0
🔁 0
💬 1
📌 0

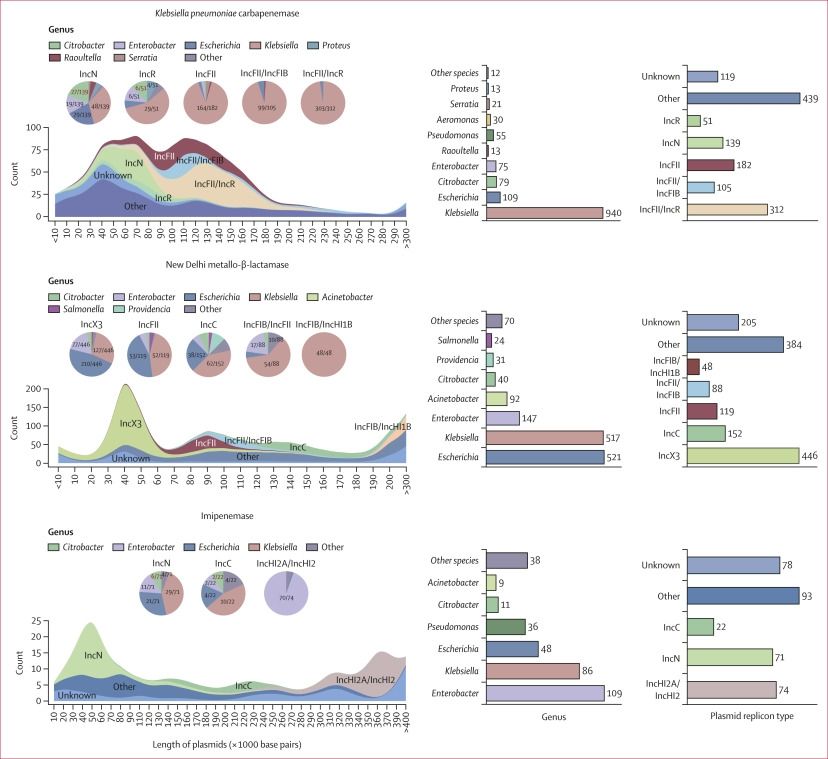
Upset plot of plasmid replicons
Most of the NDM plasmids in our dataset were IncF-type (not a popular type in other studies) and had multiple #AMR genes and mobile elements. This made it impossible to predict plasmid clusters for isolates with short-read only data
5/8
06.08.2025 13:21 —
👍 0
🔁 0
💬 1
📌 0
We used MOB-suite (github.com/phac-nml/mob...) and pling (github.com/iqbal-lab-or...) to group plasmids, and while we didn’t do a thorough comparison, their trends were the same: many small clusters = lots of diversity
4/8
06.08.2025 13:21 —
👍 0
🔁 0
💬 1
📌 0
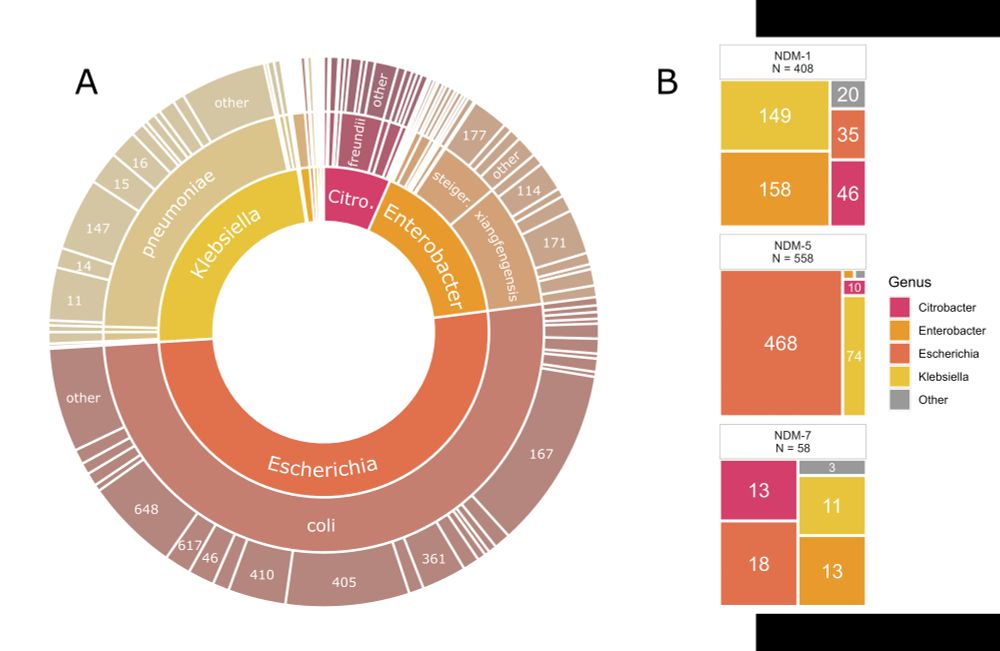
Figure depicting species and genes found in the isolate dataset
We had a big dataset (1032 isolates, 226 complete NDM plasmids) over 14 years of #AMR surveillance data from the Canadian Nosocomial Infection Surveillance Program (CNISP)
2/8
06.08.2025 13:21 —
👍 0
🔁 0
💬 1
📌 0
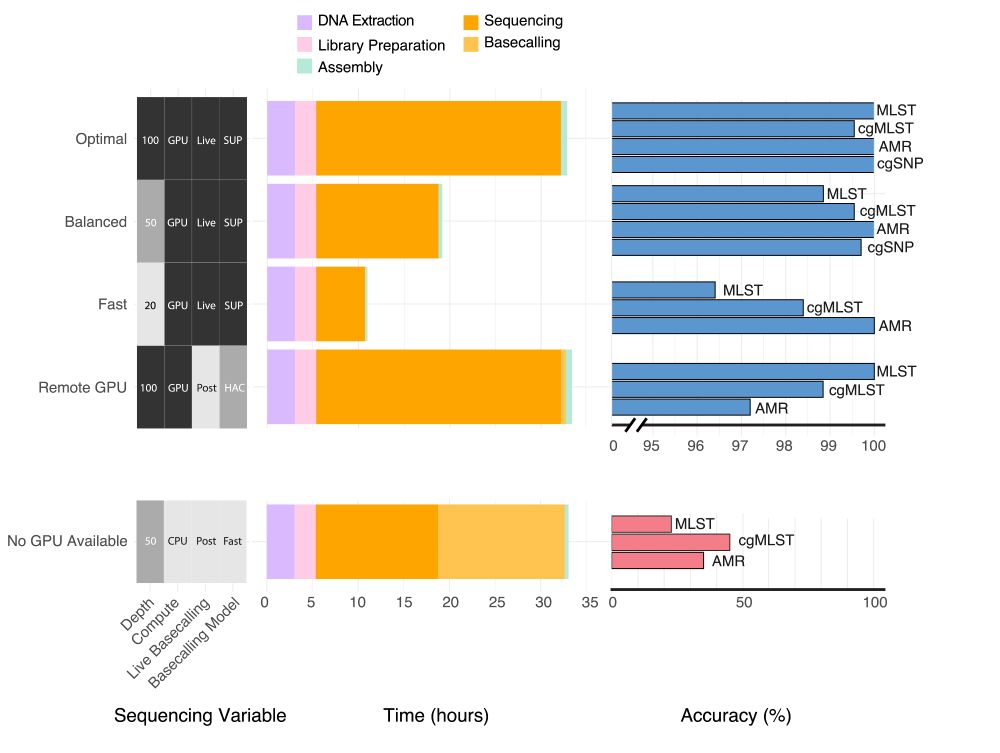
Pleased to say that our preprint benchmarking Nanopore data for MLST, cgMLST, cgSNP & AMR typing from bacterial isolates is out! TL;DR you can get almost perfect results from 50x depth using live SUP basecalling with a GPU in under 20 hours #microsky#IDsky 🦠🧬🖥️ /1
www.medrxiv.org/content/10.1...
30.07.2025 02:10 —
👍 46
🔁 29
💬 3
📌 2
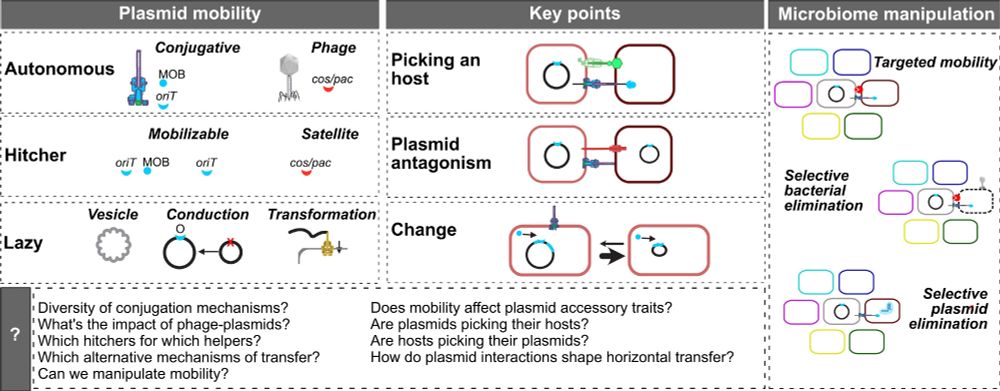
graphical abstract of the article the extended mobility of plasmids
Here's our new broad review on the extended mobility of plasmids, about all mechanisms driving and limiting their transfer. From conjugation to conduction, phage-plasmids to hitchers, molecular to evolutionary dynamics, ecology to biotech. The state of affairs. 1/9 academic.oup.com/nar/article/...
23.07.2025 07:35 —
👍 184
🔁 93
💬 4
📌 9
20231220 3h
An example of the clinician-friendly output report we are designing can be found here: lerminin.github.io/abphm_2025_p...
2/3
22.05.2025 14:47 —
👍 0
🔁 0
💬 1
📌 0

Pitch for ABPHM25 poster
Enjoying #APBHM25 - lots of super talks and excited to try out all the new tools!
I'm virtually presenting poster # 115 on our lab & bioinf workflow to rapidly identify bacteria & fungi organisms and AMR determinants from positive blood cultures, check it out here: github.com/lerminin/abp...
1/3
22.05.2025 14:47 —
👍 3
🔁 0
💬 2
📌 0
Happy to see this out - please try my code and break it! Also come see me at #ABPHM25 in the showcase today, where I’m presenting AMRGen, an R package for combining and exploring AMR geno pheno data. It was used to help make many of these rules interpretamr.github.io/AMRgen/
22.05.2025 11:57 —
👍 14
🔁 3
💬 1
📌 0