Has anyone else noticed that the bathrooms at #bps2026 have low-key good ambient electronic music for no reason?
23.02.2026 18:14 — 👍 0 🔁 0 💬 0 📌 0

I'm giving a poster later today @ 1:45. Come learn about how we're using MD and free energy calculations to better understand specific antibody-epitope recognition in concert w/ experiment! #bps2026 #BPS2026
23.02.2026 15:18 — 👍 2 🔁 0 💬 0 📌 0

@filizolalab1 kicking off at #bps2026 with a poster presentation by Kirill, on a collaborative work with the Skiniotis and Robertson labs. Go listen to Kirill!
22.02.2026 16:25 — 👍 3 🔁 1 💬 0 📌 0

BOLD-GPCRs: A Transformer-Powered App for Predicting Ligand Bioactivity and Mutational Effects across Class A GPCRs
G Protein-Coupled Receptors (GPCRs) are important targets for drug discovery owing to their ability to respond to a broad range of stimuli and their involvement in numerous pathologies. Although traditional ligand-based and structure-based approaches have facilitated the development of effective therapeutics for many GPCRs, these approaches often fall short when applied to receptors with limited ligand or structural data. This limitation highlights the critical need for advanced strategies capable of accurately predicting ligand bioactivity across the entire GPCR family, especially for understudied receptor subtypes. In this study, we introduce BOLD-GPCRs (BERT-Optimized Ligand Discovery for GPCRs), a deep learning framework designed to enhance the prediction of ligand bioactivity across class A GPCRs. Accessible via a user-friendly web interface, BOLD-GPCRs employs transfer learning and leverages curated data sets of known class A GPCR ligands, receptor sequences, and signaling-relevant mutations. By integrating dense neural network classifiers with transformer-based protein language models, BOLD-GPCRs captures complex relationships between receptor sequence/function and ligand activity. Our results demonstrate that BOLD-GPCRs achieves robust predictive performance for both ligand bioactivity and mutational effects across a broad range of class A GPCRs, underscoring its potential as a valuable tool for ligand discovery, especially for poorly characterized receptors.
After the most surreal peer review process we are out in the wild with BOLD-GPCRs: A Transformer-Powered App for Predicting Ligand Bioactivity and Mutational Effects across Class A GPCRs | Journal of Chemical Information and Modeling pubs.acs.org/doi/full/10....
14.01.2026 16:48 — 👍 5 🔁 1 💬 0 📌 0
First preprint of the year, led by Junjie Zhu from Haifeng Chen’s lab
Extending Conformational Ensemble Prediction to Multidomain Proteins and Protein Complex
doi.org/10.64898/202...
16.01.2026 17:07 — 👍 32 🔁 8 💬 0 📌 0
"Academics/scientists, stop using AI generated images" challenge, any%
20.11.2025 11:03 — 👍 17 🔁 10 💬 1 📌 0

Desireless performing in 1986
Physical Chemist at the Biochem conference #chemchat
07.12.2025 20:48 — 👍 16 🔁 3 💬 0 📌 0

It’s official: ChatGPT can't draw a GPCR, but you’ll master them in the Filizola Lab
😜 Join Us! (send your CV and the names of at least two references to my institutional email) #Postdoc #scientist #research
17.10.2025 04:50 — 👍 12 🔁 12 💬 0 📌 0
How many crystal structures do you need to trust your docking results? https://www.biorxiv.org/content/10.1101/2025.09.19.677428v1
25.09.2025 02:48 — 👍 7 🔁 3 💬 0 📌 0

Our first protein design paper out in Protein Science
onlinelibrary.wiley.com/doi/10.1002/...
24.09.2025 14:53 — 👍 14 🔁 4 💬 3 📌 0
This seems super cool, excited to give it a read!
24.09.2025 15:01 — 👍 1 🔁 0 💬 1 📌 0
Awesome! I'm excited to give these a read and try this out!
16.09.2025 20:33 — 👍 1 🔁 0 💬 0 📌 0

Abstract: Under the banner of progress, products have been uncritically adopted or
even imposed on users — in past centuries with tobacco and combustion engines, and in
the 21st with social media. For these collective blunders, we now regret our involvement or
apathy as scientists, and society struggles to put the genie back in the bottle. Currently, we
are similarly entangled with artificial intelligence (AI) technology. For example, software updates are rolled out seamlessly and non-consensually, Microsoft Office is bundled with chatbots, and we, our students, and our employers have had no say, as it is not
considered a valid position to reject AI technologies in our teaching and research. This
is why in June 2025, we co-authored an Open Letter calling on our employers to reverse
and rethink their stance on uncritically adopting AI technologies. In this position piece,
we expound on why universities must take their role seriously toa) counter the technology
industry’s marketing, hype, and harm; and to b) safeguard higher education, critical
thinking, expertise, academic freedom, and scientific integrity. We include pointers to
relevant work to further inform our colleagues.

Figure 1. A cartoon set theoretic view on various terms (see Table 1) used when discussing the superset AI
(black outline, hatched background): LLMs are in orange; ANNs are in magenta; generative models are
in blue; and finally, chatbots are in green. Where these intersect, the colours reflect that, e.g. generative adversarial network (GAN) and Boltzmann machine (BM) models are in the purple subset because they are
both generative and ANNs. In the case of proprietary closed source models, e.g. OpenAI’s ChatGPT and
Apple’s Siri, we cannot verify their implementation and so academics can only make educated guesses (cf.
Dingemanse 2025). Undefined terms used above: BERT (Devlin et al. 2019); AlexNet (Krizhevsky et al.
2017); A.L.I.C.E. (Wallace 2009); ELIZA (Weizenbaum 1966); Jabberwacky (Twist 2003); linear discriminant analysis (LDA); quadratic discriminant analysis (QDA).

Table 1. Below some of the typical terminological disarray is untangled. Importantly, none of these terms
are orthogonal nor do they exclusively pick out the types of products we may wish to critique or proscribe.

Protecting the Ecosystem of Human Knowledge: Five Principles
Finally! 🤩 Our position piece: Against the Uncritical Adoption of 'AI' Technologies in Academia:
doi.org/10.5281/zeno...
We unpick the tech industry’s marketing, hype, & harm; and we argue for safeguarding higher education, critical
thinking, expertise, academic freedom, & scientific integrity.
1/n
06.09.2025 08:13 — 👍 3751 🔁 1879 💬 109 📌 386
Congrats Roland :)!
06.09.2025 01:03 — 👍 1 🔁 0 💬 0 📌 0
I actually love this!
30.08.2025 14:31 — 👍 1 🔁 0 💬 0 📌 0
Exciting to see our protein binder design pipeline BindCraft published in its final form in @Nature ! This has been an amazing collaborative effort with Lennart, Christian, @sokrypton.org, Bruno and many other amazing lab members and collaborators.
www.nature.com/articles/s41...
27.08.2025 16:14 — 👍 305 🔁 109 💬 14 📌 11
Thanks! I spoke with Ivan earlier this year about our intersections, I gave a talk on this paper and our pylambdaopt work at the Folding@NYC meeting in Jan. I'm really excited for that as well and generally for your tools. I think they are very powerful and beneficial for the field 💪!
19.08.2025 17:47 — 👍 1 🔁 0 💬 0 📌 0

Expanded ensemble was also able to well capture average residue-wise mutation effects, potentially allowing for prediction of beneficial position-wise mutation sites.
19.08.2025 16:20 — 👍 0 🔁 0 💬 0 📌 0

We found that, although Flex-ddG was more accurate, this accuracy came from a conservative prediction tendency (predicting most mutations to be neutral). Expanded ensemble however, was better able to predict significantly stabilizing or destabilizing mutations.
19.08.2025 16:16 — 👍 0 🔁 0 💬 1 📌 0

Additionally, we re-ran Rosetta-based SSM using the Flex-ddG protocol.
19.08.2025 16:12 — 👍 0 🔁 0 💬 1 📌 0

Using Bayesian inference of the convoluted high-throughput FACS data from the publication of these designs [Chevalier, A.; Silva, D. A.; Rocklin, G. J.; Nature 2017, 550, 74–79 doi.org/10.1038/natu... ] we estimated the experimental binding affinities of all site-saturated mutants in the 3 binders.
19.08.2025 16:10 — 👍 0 🔁 0 💬 1 📌 0

We used the massive amount of statistics from these calculations to quantify sources of uncertainty. Our uncertainties were distributed bimodally with a mean of ~2 kcal/mol. Sources of larger uncertainties include number of alchemical atoms, charge changes, and slow DOFs.
19.08.2025 16:05 — 👍 0 🔁 0 💬 1 📌 0
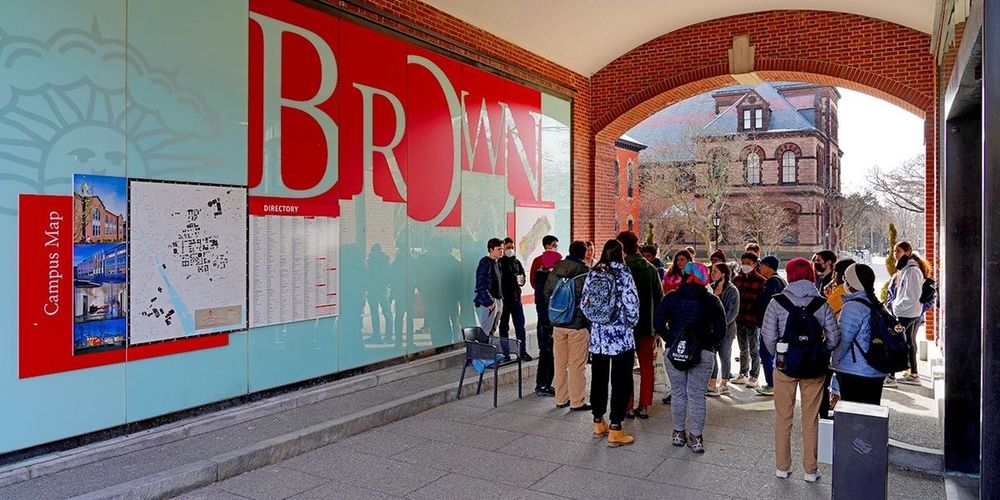

Brown University is ‘functionally inaccessible’ to transgender students after Trump settlement
“Everyone I talked to thought it only applied to sports. But it applies to everything," one transgender student told The Advocate.
The Trump administration effectively created a de facto bathroom ban at Brown University and none of the major media is covering it, and if they do, they character it as about sports. This is how the trans visa ban was enacted and characterized.
www.advocate.com/news/transge...
06.08.2025 03:55 — 👍 2150 🔁 876 💬 30 📌 38
CASP is the main reason the protein structure prediction technology and research field advanced over the last 30 years. And the main reason AI based methods have been accepted and widely applied in biology. So shortsighted of NIH to postpone or even halt funding. John Moult is a scientific hero.
02.07.2025 22:36 — 👍 41 🔁 14 💬 1 📌 0
Very funny in the saddest and scariest way that people who are apparently in love with western civilization don't know that trans is a latin word that has an actually meaning and are willing to dismantle science in the u.s. using ctrl-f over their hatred of trans people. Very cool place 😎
27.06.2025 14:43 — 👍 0 🔁 0 💬 0 📌 0
Postdoc in biophysics/biochemistry working on the interaction between the phosphorylated Tau protein and microtubules. Molecular dynamist. Tamer of IDRs and IDPs since 2022
(They/Them)
https://scholar.google.com/citations?user=4S1QUPgAAAAJ&hl=fr
Harvard-based consortium curating structural biology (CryoEM Crystallography NMR Tomography) software, hosting educational webinars, supporting reproducibility, validation, reprocessing of data, & access to scientific resources sbgrid.org / biogrids.org
optimization, inverse problems, also proteins. ml at escalante. formerly: atomicai, xgenomes, broad, berkeley.
The community-wide CAPRI protein docking and assembly modeling experiment
Computational physicist at https://peptone.io
PhD @GroupParrinello, PostDoc @franknoe.bsky.social
Disordered Proteins, AI for Science, Molecular Dynamics, Enhanced Sampling
🔗 https://scholar.google.com/citations?user=fnJktPAAAAAJ
OpenBind is building the world’s largest open-access dataset of drug–protein interactions to speed up new treatments. Through cutting-edge science and open collaboration, we power AI tools for structure-based drug design – free for all.
We are senior researchers at the Icahn School of Medicine at Mount Sinai forming a union to have a voice in our working conditions and improve research at Sinai as whole.
Automated discovery of AI x Bio papers, blogs, and news.
A growing coalition championing state-backed biomedical research in New York—advancing discovery, education & new therapies for real-world impact. www.nycures.org
We are researchers organizing with @nycures.bsky.social @AMSNewYork @uaw.org to fund NY science by establishing state-funded biomedical research in New York. Join us!
Computational Structural Biologist
Assistant Professor @VanderbiltMPB
wankowiczlab.com
(she/her)
Past: UCSF, Dana-Farber, Broad Institute, UMass Amherst
Ad Astra Fellow, Asst. Prof. of Digital Chemistry, @ucddublin.bsky.social |
Editor, @joss-openjournals.bsky.social |
Personal: ewcss.info |
Research group (@coreacter.org): CoReACTER.org |
ORCID: orcid.org/0000-0003-1554-197X |
All opinions mine
Ph.D Student at http://chembio.triiprograms.org
Interested in the history of life, language, and science.
https://apayne97.github.io/
Organizing the largest workshop concerning Free Energy calculations within the United States.
The original & only WomensArt created/ curated by art historian and author PL Henderson
Real art...real artists
Images © to respective owners
Ph.D. candidate in Biophysics Shukla Group | University of Illinois | Molecular dynamics simulations 🧬 + ML 🤖
#compchem | organic semiconductors | functional biomaterials | polymers | @mdanalysis core dev | made in Sardinia 🏳️🌈 she/her
structural biology, biophysics
guitarist for @KulfiGirls
she/her
🧬🎵🏳️⚧🏳️🌈
Immunologist, professor at the Karolinksa Institutet, member of the Nobel Assembly
Assistant Professor at Stockholm University.
Dedicated Scientist.