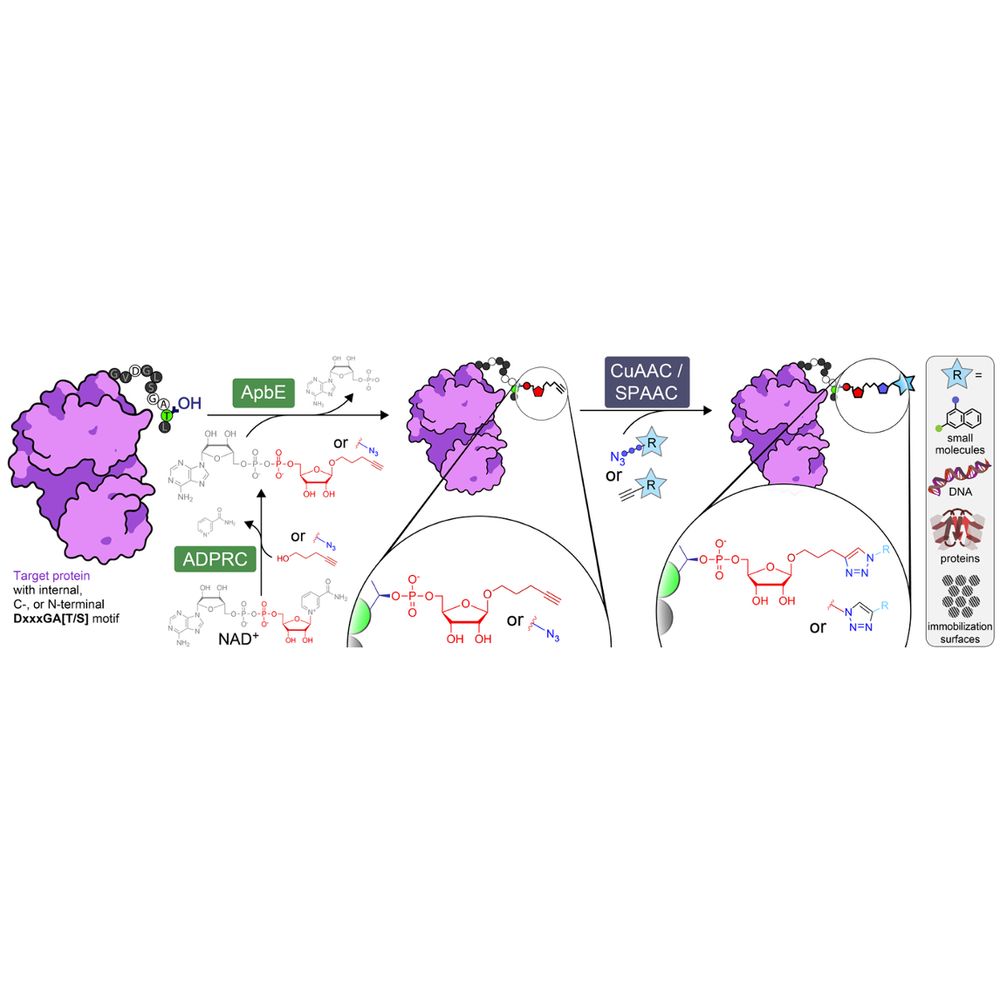
Overview of ADD-tagging including ADP-ribosyl cyclase (ADPRC)-catalysed dinucleotide substrate generation and two step chemoenzymatic labelling of target proteins with the flavin transferase ApbE, followed by click chemistry-based functional group attachment.
🚨 preprint 2️⃣ this month: our (purely experimental🧪) venture into #ChemBio
We prouldy present: ADD-tagging of proteins (or "ADDing") —a super convenient enzymatic technique to install click chemistry handles on proteins.
Led by superstar @wahyuwidodo.bsky.social
www.biorxiv.org/content/10.1...
A 🧵👇🏽
20.05.2025 12:29 —
👍 53
🔁 15
💬 4
📌 2

Roelfes group - Open Positions
PhD position on Hybrid biocatalytic cascades aided by Machine learning (1.0 FTE)
📢📢📢 PhD Opportunity!
Join our research group at RUG! We're seeking a motivated PhD student for a project on biocatalytic cascade processes combining natural and designer enzymes, supported by machine learning.
Click below to learn more — and feel free to share!
sites.google.com/rug.nl/roelf...
12.05.2025 09:10 —
👍 11
🔁 13
💬 0
📌 1
Thank you @francdefel.bsky.social, Andy and Gerard for the great collaboration!
@roelfesgroup.bsky.social @stratinghinst.bsky.social
23.04.2025 09:17 —
👍 2
🔁 0
💬 0
📌 0
Last but not least, the simple and straightforward genetic incorporation of aY facilitated the application of the evolved Friedel-Crafts alkylase in whole-cells!
23.04.2025 09:17 —
👍 1
🔁 0
💬 1
📌 0
X-ray crystal structures of the parent and evolved mutant (2.2 Å and 1.2 Å!) showed a significant change in the rotameric state of the aY catalytic residue, and a narrowing of the active site cavity.
23.04.2025 09:17 —
👍 1
🔁 0
💬 1
📌 0
Directed evolution gave rise to a quadruple mutant that showed increased activity and excellent enantioselectivity (up to 95% ee).
23.04.2025 09:17 —
👍 1
🔁 0
💬 1
📌 0
Through genetic incorporation of aY into LmrR, we create an artificial Friedel-Crafts alkylase that is enantiocomplementary to a previous design featuring p-aminophenylalanine as catalytic residue.
23.04.2025 09:17 —
👍 1
🔁 0
💬 1
📌 0
We demonstrate the first example of using noncanonical 3-aminotyrosine (aY) as a catalytic residue for iminium activation in a designer enzyme
23.04.2025 09:17 —
👍 1
🔁 0
💬 1
📌 0
Very proud to share that one of my main PhD projects has been published in @chemicalscience.rsc.org, and that it has been selected as a #ChemSciPicks of the week paper!
Keep reading below for more details, or check out the full paper for free online: doi.org/10.1039/D5SC...
Thread ⬇️
23.04.2025 09:17 —
👍 19
🔁 2
💬 1
📌 1
Grateful to be part of this awesome project lead by @brouwerb.bsky.social. Take a look to our preprint about unlocking iminium ion catalysis when using amino-tyrosine as a genetically encoded amino acid! 🤓
#MSCA @roelfesgroup.bsky.social
12.02.2025 08:31 —
👍 4
🔁 3
💬 0
📌 0
Thank you Franco, Andy and Gerard for the great collaboration!
Looking forward to seeing it published soon, keep your eyes peeled!
@roelfesgroup.bsky.social @stratinghinst.bsky.social
11.02.2025 20:05 —
👍 0
🔁 0
💬 0
📌 0
Last but not least, the simple and straightforward genetic incorporation of aY facilitated the application of the evolved Friedel-Crafts alkylase in whole-cells!
11.02.2025 20:05 —
👍 0
🔁 0
💬 1
📌 0
X-ray crystal structures of the parent and evolved mutant (2.2 Å and 1.2 Å!) showed a significant change in the rotameric state of the aY catalytic residue, and a narrowing of the active site cavity.
11.02.2025 20:05 —
👍 0
🔁 0
💬 1
📌 0
Directed evolution gave rise to a quadruple mutant that showed increased activity and excellent enantioselectivity (up to 95% ee).
11.02.2025 20:05 —
👍 0
🔁 0
💬 1
📌 0
Through genetic incorporation of aY into LmrR, we create an artificial Friedel-Crafts alkylase that is enantiocomplementary to a previous design featuring p-aminophenylalanine as catalytic residue.
doi.org/10.26434/che...
11.02.2025 20:05 —
👍 1
🔁 0
💬 1
📌 0
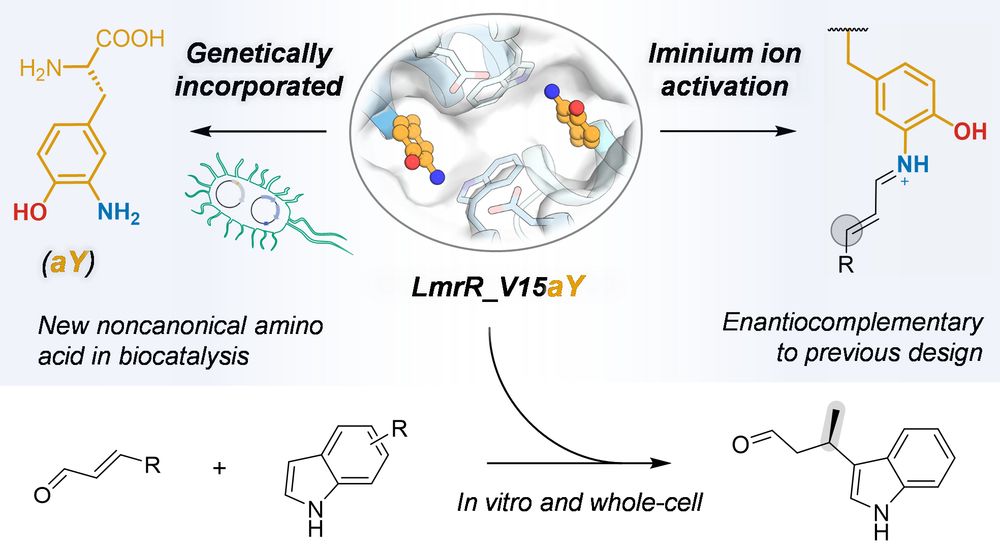
Check out our latest preprint, in which we demonstrate the first example of using noncanonical 3-aminotyrosine (aY) as a catalytic residue for iminium activation in a designer enzyme!
Thread ⬇️
11.02.2025 20:05 —
👍 16
🔁 5
💬 1
📌 2



