New preprint! We unexpectedly discovered that some Caenorhabditis species delete parts of their somatic genome early in development, which fragments their chromosomes and eliminates key germline genes. Multiple lines of evidence suggest this bizarre process was present in the ancestors of C. elegans
28.10.2025 12:11 — 👍 49 🔁 17 💬 0 📌 0

Amjad Khalaf follows the symbiosis genomics session at #biodiversity25 presenting about #microsporidia tetraploid genomes. Most of them are recent autotetraploids. An interesting model of reproduction is proposed with two different paths: sexual and asexual reproduction.
28.10.2025 11:49 — 👍 3 🔁 3 💬 0 📌 0
Oh hello! He's only gone and re-written the Microsporidian reproductive tree and published it in @plosbiology.org!
Heroic effort, Amjad. Congrats to all the authors!
You can read about Amjad and his doctoral research in the blog I wrote all about him: sangerinstitute.blog/2025/07/29/s...
22.10.2025 16:48 — 👍 2 🔁 2 💬 0 📌 0

Top: Prevalence of Microsporidia in DToL insect genomes. Microsporidian genomes recovered from insect hosts, split by taxonomic order. F: female, M: male, U: unspecified sex. Bottom: Simplified proposed generalized lifecycle for Microsporidia. This proposed model posits that each nucleus is a diploid, and that microsporidian reproduction mirrors reproduction in Fungi with stages similar to karyogamy, plasmogamy, and a stable “heterokaryon” (known as a diplokaryon in Microsporidia). Importantly, both the diplokaryotic and monokaryotic phases are parasitic, and species may spend most of their lifecycle in one or the other phase, giving rise to “diploid” and “tetraploid” lineages.
#Microsporidia are #parasites of growing importance. @amjadkhalaf.bsky.social @mblaxter.bsky.social @marakat.bsky.social &co present 40 new genomes from #DarwinTreeofLife sequencing of their arthropod hosts, revealing insights into #tetraploidy & #reproduction @plosbiology.org 🧪 plos.io/48HsAQJ
22.10.2025 13:07 — 👍 7 🔁 2 💬 0 📌 1
Really, really happy that our work on microsporidian genomes is now out in @plosbiology.org! A huge thank you to my coauthors, my supervisors @mblaxter.bsky.social & @marakat.bsky.social, the editors @roliroberts.bsky.social & Joseph Heitman, and the reviewers ❤️ plos.io/48HsAQJ
22.10.2025 10:47 — 👍 28 🔁 13 💬 0 📌 2

Image shows Amjad Khalaf sitting on a bench outside, working on his laptop.
Meet Amjad Khalaf – who has spent his PhD panning for parasite DNA, allowing him to create a goldmine of information for the research community🔎
Amjad shares what first brought him to the Sanger Institute, his research and what he is dreaming about next:
sangerinstitute.blog/2025/07/29/s...
29.07.2025 14:49 — 👍 5 🔁 3 💬 0 📌 0
it's really an honour to be interviewed by the wonderful @carmcdhume.bsky.social -- a huge thank you! ❤️🫂
03.08.2025 20:37 — 👍 2 🔁 0 💬 0 📌 0
it was so so lovely to do this together! 🇭🇷❤️🇵🇸
03.08.2025 19:16 — 👍 1 🔁 1 💬 0 📌 0

If you’re curious about Microsporidia, polyploidy, or assembling cobionts from non-target organisms — please check out our preprint! A HUGE thank you to my wonderful coauthors, and my supervisors @mblaxter.bsky.social and @marakat.bsky.social
16.05.2025 10:58 — 👍 1 🔁 0 💬 0 📌 0

The tetraploid genomes that we see are consistent with autopolyploidy, not allopolyploidy. While we’re unable to place the timing of polyploidisation events in Microsporidia, we propose that tetraploidy is a shared & ancient event in the group, with diploid lineages representing “reduced” forms.
16.05.2025 10:58 — 👍 2 🔁 0 💬 1 📌 0

Using the Hi-C data, we find that a tetraploid microsporidian genome is organised into two diploid compartments, likely the two nuclei in a diplokaryon. We also see evidence of recombination both within and between the two compartments - providing evidence for a sexual cycle in Microsporidia.
16.05.2025 10:58 — 👍 0 🔁 0 💬 1 📌 0
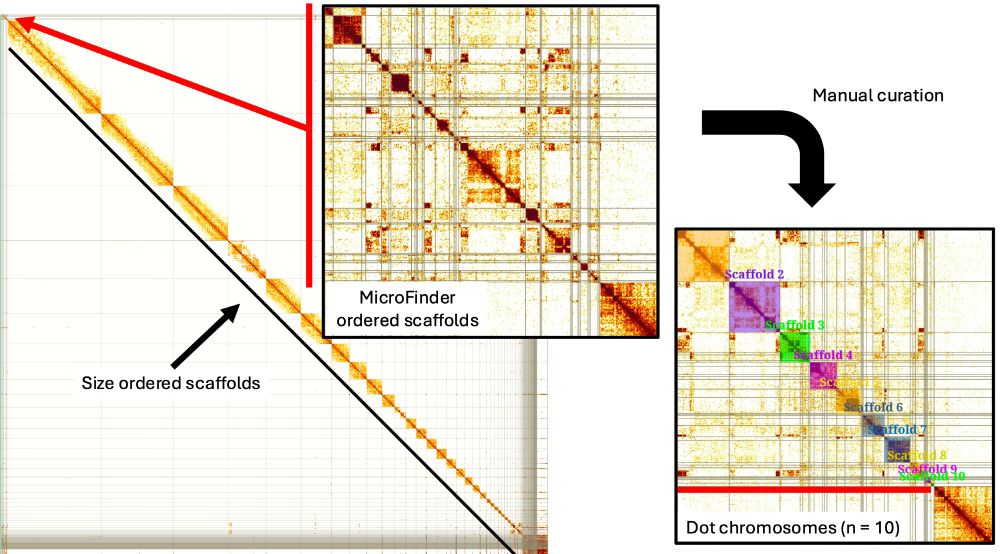
Struggling to find the tiny microchromosomes in draft bird genome assemblies?
Our new #preprint introducing MicroFinder can help: www.biorxiv.org/content/10.1...
🧵 below
(1/n)
@sangerinstitute.bsky.social
16.05.2025 10:07 — 👍 40 🔁 22 💬 3 📌 1
Postdoc in computational genomics studying RNA viruses at Cambridge. She/Her https://orcid.org/0000-0002-8400-6922
We foster path-breaking scientific discovery, environmental conservation, and preservation of the special character of the Bay Area.
Genome annotation geek at University of Greifswald, always interested in friendly scientific collaboration. Developer of BRAKER, GALBA, MakeHub and other Gaius-Augustus tools.
Cambridge (UK) based science arts and crafts
created by @fionatel.bsky.social and @petrathepostdoc.bsky.social
https://theintrovertebrates.carrd.co/
Research PI @leibnizlib.bsky.social Bonn Germany #biodiversity #genomics #museomics #sexchromosomes #cichlids
PhD student at the Sanger Institute / University of Cambridge
Prev. Associate Computational Biologist at the Broad Institute
UWaterloo CS
anant-maheshwari.owlstown.net
Head of the Biodiversity Genomics lab
All enthusiastic about butterflies, holocentric chromosomes, speciation, evolution and conservation!
Asst. Prof. at University of Geneva
Evolutionary cell biology & multicellular developmental diversity of protists
www.dudinlab.com
#Ichthyosporea, #Multicellularity, #Evolution, #Embryo, #Development, #Protist, #UExM, #Cytoskeleton, #Expansion #Microscopy
I summon things into existence by typing. 🐪 OΛ
🌉 bridged from ⁂ https://mstdn.science/@jgrg, follow @ap.brid.gy to interact
Zoologist at Girton College, University of Cambridge. Author of 'The Zoologist's Guide to the Galaxy' and 'Why Animals Talk' https://bit.ly/482925U. Animal communication researcher, https://www.bioacousticsresearchgroup.org/
Senior Editor at @plosbiology.org; Scientist, Humanist, Optimist and Recovering Academic. All opinions my own.
Eco-evo biologist - post-doc @kuleuvenuniversity.bsky.social - Research interests: population & landscape #genomics 🧬, (co-) #evolution, #bats 🦇, #viruses 🦠, #Leishmania & #fish 🐟
a doodle every day!
🎨 art director on Monopoly GO!
🌱 silly pink cat
🌟 x.com/stefscribbles
Dr in Evolutionary Genetics. Genomics of complex eukaryotes in symbiosis, biodiversity, speciation 🧬👩🏻💻 She/Her🌈
Professor, Molecular & Cellular Biology, Harvard University
Biosequence analysis using profile hidden Markov Models.
HMMER is managed by EMBL’s European Bioinformatics Institute (EMBL-EBI).
Professor of Marine Microbiology at GEOMAR. A particular interest in marine sponge symbioses and in ocean health & disease matters.
Postdoc in Lars Velten's lab @crg.eu, working on gene regulation in haematopoiesis using ML techniques
https://jacobhepkema.github.io








