Mapping targetable sites on the human surfaceome for the design of novel binders | PNAS
The human cell surfaceome, integral to cell communication and disease mechanisms, presents a prime target for therapeutic intervention. De novo pro...
A new @pnas.org study caught the eye of our bioinformatics team with its identification of ~4,500 targetable binding sites & binding seeds across human cell surface proteins—giving IPI and other labs a protein design launch point.
26.02.2026 16:05 — 👍 1 🔁 1 💬 0 📌 0
The final version of our new paper is out now - and open access @acs.org Central Science!!
Such a fun collaboration!
pubs.acs.org/doi/10.1021/...
25.01.2026 01:05 — 👍 33 🔁 8 💬 0 📌 1

New preprint 🥳! We made photoclickable HaloTag ligands to precisely control protein labeling on living cells. With it, we can do some cool multicolor stuff. Huge congrats to Franzi and all co-authors! Check it out 👇
www.biorxiv.org/content/10.1...
13.11.2025 16:20 — 👍 157 🔁 40 💬 0 📌 4
Chemical biology 2026
Registration for THE chemical biology conference of 2026 is now open! EMBO ChemBio 2026 in Heidelberg
DeGrado, Arikin, Picotti (Keynotes). @lmkdassama.bsky.social @brianliau.bsky.social @rhodamine110.bsky.social @benlehner.bsky.social @alitavassoli.bsky.social
www.embl.org/about/info/c...
11.11.2025 17:46 — 👍 32 🔁 19 💬 2 📌 0
PNAS
Proceedings of the National Academy of Sciences (PNAS), a peer reviewed journal of the National Academy of Sciences (NAS) - an authoritative source of high-impact, original research that broadly spans...
A new, nerdy paper. We figured out (some) of the rules underlying cell-permeability of probes and designed ligands that light up, grab, and move proteins around. Awesome @hhmijanelia.bsky.social x @uwmadison.bsky.social x @stjuderesearch.bsky.social collaboration! www.pnas.org/doi/10.1073/...
27.10.2025 21:43 — 👍 87 🔁 30 💬 0 📌 1

ProteinDJ logo
I am thrilled to release ProteinDJ: a high-performance and modular protein design pipeline. Our open-source workflow incorporates #RFdiffusion, #ProteinMPNN, #FAMPNN, #AlphaFold2 and #Boltz-2. It is a fast, free, and fun way to design proteins (1/5)
doi.org/10.1101/2025.09.24.678028 #proteindesign
28.09.2025 21:16 — 👍 88 🔁 31 💬 2 📌 6
Paralog-specific intrabodies for PSD-93 and SAP102 expand the molecular toolkit to resolve excitatory synapse organization https://www.biorxiv.org/content/10.1101/2025.08.24.671450v1
26.08.2025 00:46 — 👍 1 🔁 1 💬 0 📌 0
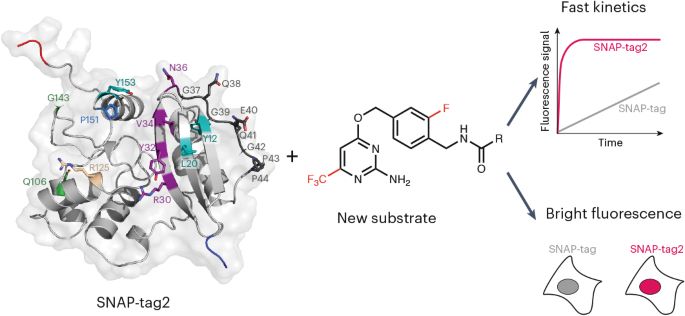
Also check out the associated News & Views by Masayasu Taki and Masayoshi Nakamura highlighting the work from @kjohnsson.bsky.social lab on SNAP-tag2
www.nature.com/articles/s41...
08.07.2025 16:02 — 👍 17 🔁 10 💬 0 📌 0

We are thrilled to welcome Barbara Imperiali at #ChemBioParis2025 in Paris, October 6-9, 2025. Sign up here: chembioparis2025.com – Early bird ends June 10 with reduced fees for @icbschembio.bsky.social - euchems.eu - @eu-openscreen.bsky.social - @scf-chembio.bsky.social - #ChemBioEvent
01.05.2025 11:19 — 👍 7 🔁 4 💬 0 📌 0

Toward single-molecule protein sequencing using nanopores - @unigroningen.bsky.social go.nature.com/3FL0g3k
18.03.2025 13:39 — 👍 33 🔁 12 💬 0 📌 1
Super excited to finally share PANCS-Binders: our group's decade-long quest to accelerate protein binder discovery.
TLDR: PANCS-binders is fast (2 days), cheap (pennies), has extremely high fidelity (low false positive and negatives), and high-throughput.
1/n
07.01.2025 20:50 — 👍 94 🔁 36 💬 4 📌 4
Our group at LMU Munich and @mpibiochem.bsky.social uses DNA nanotechnology to develop next-generation super-resolution microscopy techniques. #DNAPAINT
Associate Professor at UCLA, all things chemical biology, chemoproteomic-, covalent-, cysteine- and redox-related
let's talk about microscopy and/or neuroscience, anything at any time!
PI's at the Laboratory of Membrane Biogenesis, CNRS, Bordeaux. Working on plasmodesmata-mediated cell-to-cell communication in plants
Chemical biologist working on tools for fluorescence microscopy | Assistant Professor ETH DCHAB | Postdoc UCSDPharm | PhD MPI-MR and EPFL
Structural biologist in Aus @WEHI. Cryo-electron microscopy of viruses and protein complexes #cryo-EM. Co-creator of ProteinDJ (He/Him)
European Research Council, set up by the EU, funds top researchers of any nationality, helping them pursue great ideas at the frontiers of knowledge. #HorizonEU
𝐂𝐮𝐬𝐭𝐨𝐦 𝐒𝐢𝐧𝐠𝐥𝐞-𝐃𝐨𝐦𝐚𝐢𝐧 𝐀𝐧𝐭𝐢𝐛𝐨𝐝𝐲 𝐃𝐢𝐬𝐜𝐨𝐯𝐞𝐫𝐲 & 𝐍𝐚𝐧𝐨𝐛𝐨𝐝𝐲-𝐁𝐚𝐬𝐞𝐝 𝐑𝐞𝐬𝐞𝐚𝐫𝐜𝐡 𝐓𝐨𝐨𝐥𝐬
Single-molecule biophysics group at Goethe University Frankfurt/Germany. Super-resolution microscopy. Optical structural cell biology. Computational microscopy. Posts by MH.
http://www.smb.uni-frankfurt.de
#superresolution, #livecellimaging, #imageanalysis
Posting our lab life studying the neuronal cytoskeleton and chromatin with cool microscopes🔬Proud #GenX, #firstgen college, #intj @RITscience @stonybrooku Alum
Verified: https://orcid.org/0000-0001-8481-0403
Chemical Biologist. Tenure track assistant prof @EPFL. Trying to make sense of the world by changing one variable at a time. https://www.epfl.ch/labs/libn/
Computational chemist/structural bioinformatician working on improving molecular simulation at MRC Laboratory of Molecular Biology. jgreener64.github.io
PhD student@MIT
https://sites.google.com/view/yehlincho/home
Publishing the best of biotech science and business. Find us on Twitter, Facebook & Instagram. Part of @natureportfolio.nature.com.
Group of young academic and industrial researchers from Société de chimie Thérapeutique (SCT) devoted to support young medicinal chemists and students who wish to become one!
CNRS senior researcher @unistra, @cnrs, @BrightSens Diagnostics, @AstraNICE. Working on fluorescent probes, organic nanoparticles, biomembranes, biosensing & bioimaging. #StandwithUkraine
Structural biologist and microbiologist at the IECB, CNRS, Bordeaux. Loves her kiddos, homies, science and bacteria!
www.krasteva-lab.com