📢 TL;DR: #Echidna 🦔integrates scRNA-seq & WGS to decouple gene dosage effects on phenotypic plasticity, tracks clonal evolution, and identifies resistance drivers in tumors.
Read more here: www.biorxiv.org/content/10.1...
Let’s unravel tumor evolution, one clone at a time! 7/7
18.12.2024 13:30 — 👍 2 🔁 0 💬 1 📌 0
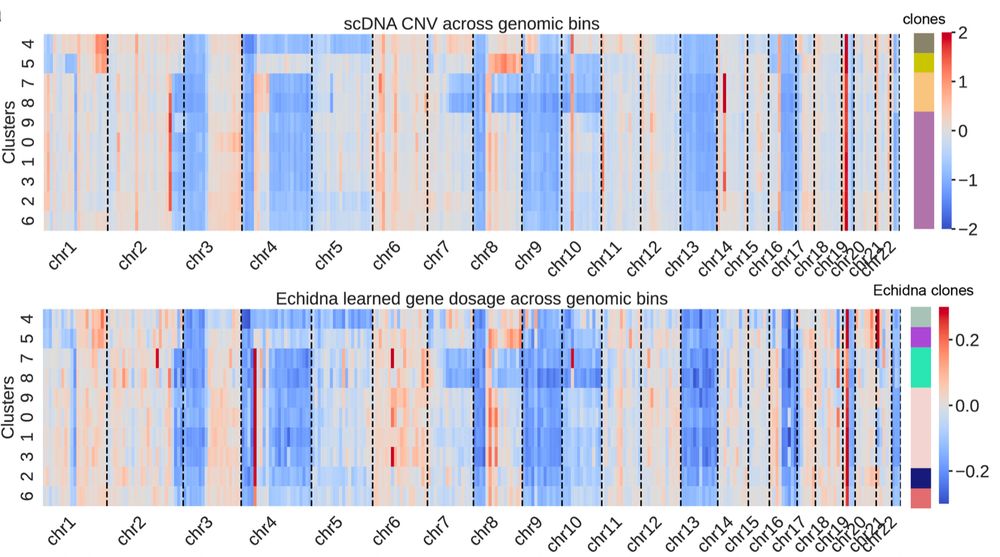
🌟 Why is this impactful? #Echidna doesn’t require joint scDNA/RNA sequencing (rare/expensive). Instead, it pairs scRNA-seq & WGS—readily obtainable from clinical samples. It’s scalable & can reveal mechanisms of cancer progression & treatment response! 🧬 6/
18.12.2024 13:30 — 👍 1 🔁 0 💬 1 📌 0
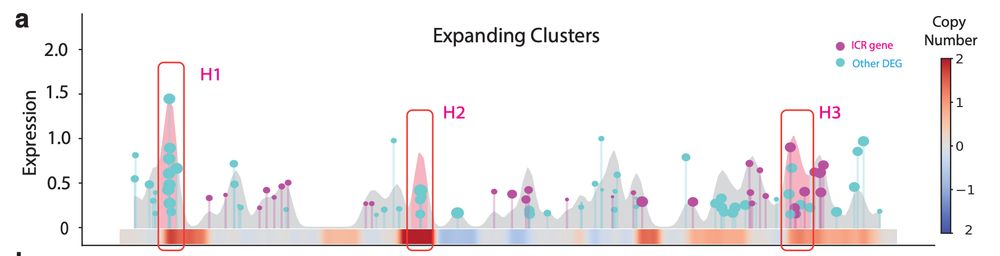
A Case Study in Melanoma: 🧬 Clones resistant to anti-PD-1 therapy showed clusters of phenotype-defining genes in hotspots of amplification! Including:
S100 family and MHC-II genes. #Echidna also disentangles intrinsic (CNA-driven) vs extrinsic expression immune signaling!
18.12.2024 13:30 — 👍 1 🔁 0 💬 1 📌 0
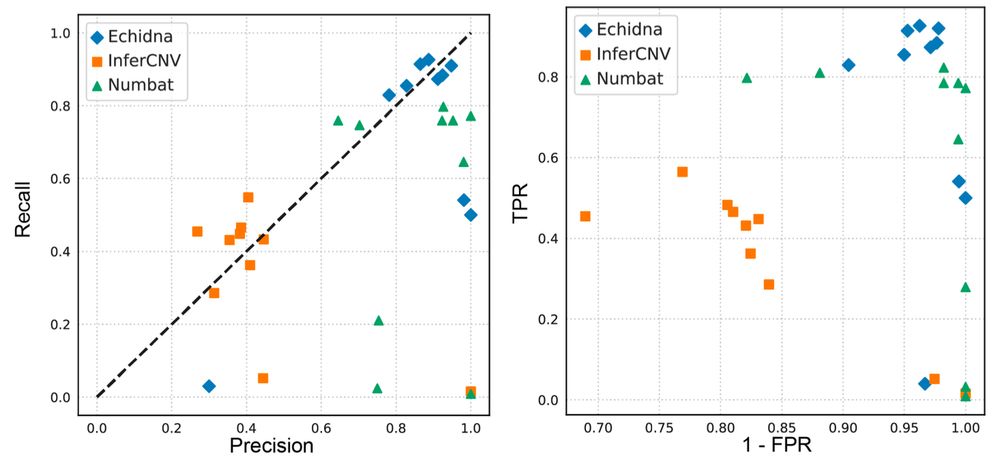
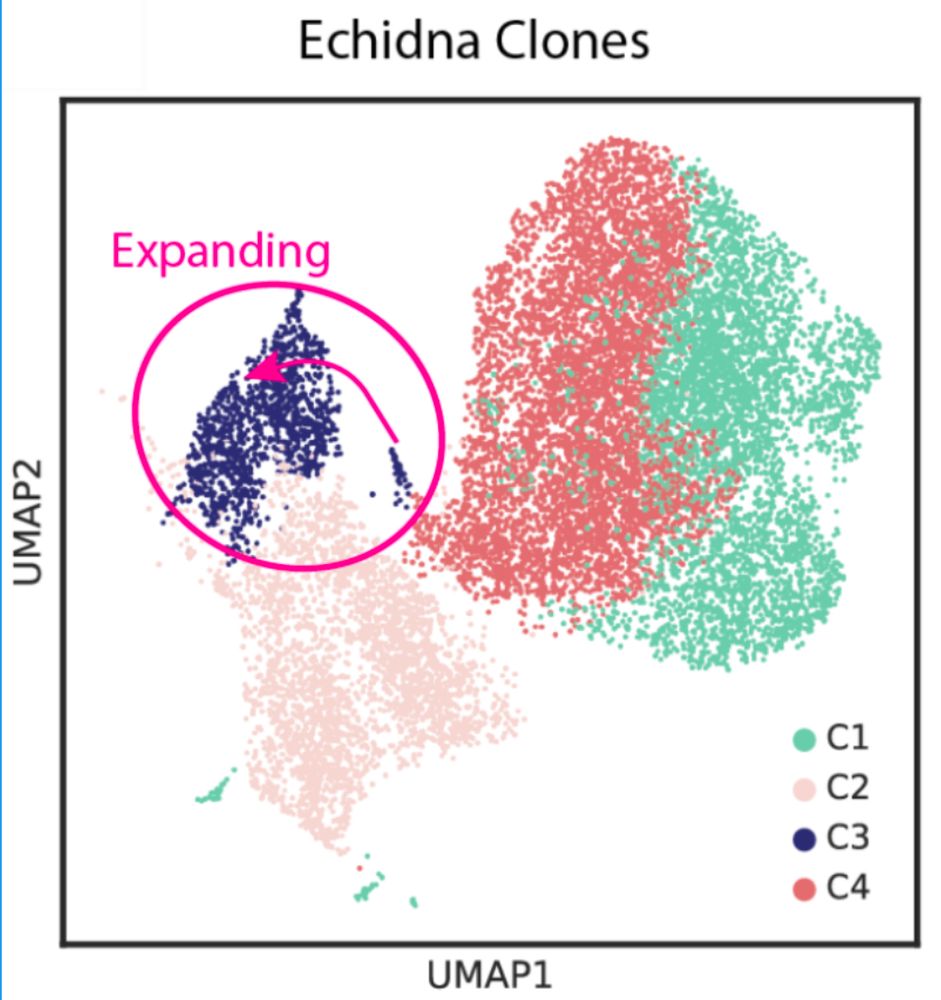
What did we find? Applied to tumor samples: 1️⃣ Clonal structure reconstructed with higher accuracy vs InferCNV/Numbat. 2️⃣ Temporal dynamics captured: tracks clonal evolution pre/post therapy. 3️⃣ Drivers of resistance identified using GDX in genomic hotspots. 4/
18.12.2024 13:30 — 👍 1 🔁 0 💬 1 📌 0
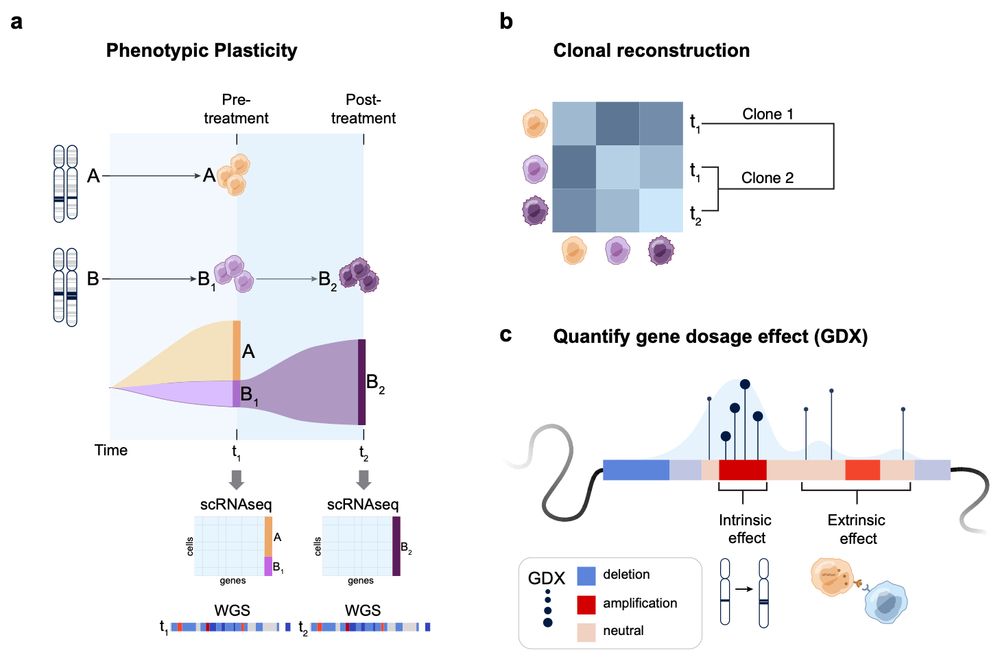
How it works: #Echidna models single-cell RNA & bulk WGS data using a Bayesian hierarchical framework. Key features:
Deconvolves CNA profiles
Tracks clones & their phenotypic states over time
Introduces GDX: a metric quantifying gene dosage effects! 3/
18.12.2024 13:30 — 👍 5 🔁 1 💬 1 📌 0
📌 Why Echidna? Phenotypic plasticity (cell adaptability) is critical in cancer progression & resistance. But how much of this plasticity is driven by gene dosage vs external cues?
Echidna uncouples these effects, bridging genome & transcriptome across timepoints!
18.12.2024 13:30 — 👍 3 🔁 1 💬 1 📌 0
🧵 Excited to share #Echidna, a Bayesian framework for quantifying the impact of gene dosage on phenotypic plasticity: tinyurl.com/296kf7hf!
With @elhamazizi.bsky.social and @mingxz.bsky.social, we integrate scRNA-seq & WGS to uncover how CNAs drive tumor evolution and transcriptional variability.
18.12.2024 13:30 — 👍 15 🔁 6 💬 2 📌 2
Senior Research Scientist at CZ Biohub New York, AI/ML Platform. Building causal AI models of immune cells.
Group leader @schapirolab.bsky.social at the Heidelberg University Hospital focusing on #HighlyMultiplexedImaging and #spatialomics #spatialbiology technologies and analysis. President of the European Society for Spatial Biology e.V. (ESSB)
Ph.D. Student @ Nerurkar Lab, Columbia BME
Developmental biology | biomechanics | stem cell research
PhD Student @ColumbiaBME and @nygenome, B.A. + M.Sci @Cambridge_Uni. Interested in computational methods for single-cell and spatial omics.
Math PhD | Interested in ML
Molecular cell biology and stats / ML, mostly scRNA-seq.
Principal Scientist at Tahoe Tx.
https://www.nxn.se/archive
PhD Candidate @ColumbiaBME 🥼🐣🛠🧬🔬
#embryo2022
computational biologist and statistician interested in how genes encode shape, form, and dynamics during development and regeneration; assistant professor @Columbia University https://www.morpho-lab.com/
CS PhD @ UCLA
https://web.cs.ucla.edu/~wob/
Researcher @bayraktar_lab @teichlab @steglelab.bsky.social @sangerinstitute.bsky.social | Using models & AI to study cells, cell circuits & brains 🧠 | #SingleCell+spatial | 🌍+🇺🇦
PhD candidate@Columbia University.
Generative and Geometric deep learning | Theory of Generative Models| Cancer systems biology
Research at Azizi lab & Califano lab.
Machine learning and biology. Research Scientist at Google DeepMind. adamgayoso.github.io. Views are my own.
My lab at Stanford studies human population genetics and complex traits.
Postdoctoral Fellow at Marioni Lab, Genentech | Aging & bioinformatics
Born in Portugal 🇵🇹 living in California 🐻
Bren Professor of Computational Biology @Caltech.edu. Blog at http://liorpachter.wordpress.com. Posts represent my views, not my employer's. #methodsmatter
MLing biomolecules en route to structural systems biology. Asst Prof of Systems Biology @Columbia. Prev. @Harvard SysBio; @Stanford Genetics, Stats.
A new scientific institution for curiosity-driven biomedical science and technology.




