
1/ I’m excited to share recent work on inferring cell-cell interactions using attention by @justjhong.bsky.social and me, supervised by @elhamazizi.bsky.social . Open the thread 🧵 for a brief overview of our method. bioRxiv link: www.biorxiv.org/content/10.1....
25.09.2025 02:06 — 👍 14 🔁 6 💬 1 📌 1

For our third Computational Spatial Biology seminar, we will be speaking with @wencksternjo.bsky.social about VirTues, a foundation model framework for analyzing complex tissue architectures.
Tune in this Wed, May 7 @ 11AM ET! Full schedule and last week's recording: csbseminar.github.io
05.05.2025 14:03 — 👍 1 🔁 0 💬 0 📌 0

For our next Computational Spatial Biology seminar, we will talking with @siyuhe.bsky.social about CORAL, a deep graph generative model learning integrated representations of multimodal spatial omics data.
Tune in next Wednesday, April 30 @ 12PM ET!
Full schedule: csbseminar.github.io
24.04.2025 16:29 — 👍 5 🔁 0 💬 0 📌 1

To kick off the Computational Spatial Biology seminar series with Nathan Levy, we will be discussing LUNA with Yist Yu from EPFL. LUNA is a generative AI model that reassembles complex tissue structures from gene expressions of cells.
Tune in next Wednesday, April 9 @ 11AM ET!
02.04.2025 16:06 — 👍 1 🔁 0 💬 0 📌 0

Cover photo of the Computational Spatial Biology Seminar website
Announcing the Computational Spatial Biology seminar!
With Nathan Levy (Yosef Lab), we’re bringing together researchers at the intersection of AI & spatial biology to share their work.
Kicking off April 9th with Yist Yu presenting LUNA!
Explore the lineup here:
csbseminar.github.io
26.03.2025 19:48 — 👍 12 🔁 4 💬 1 📌 1
🧵 Excited to share #Echidna, a Bayesian framework for quantifying the impact of gene dosage on phenotypic plasticity: tinyurl.com/296kf7hf!
With @elhamazizi.bsky.social and @mingxz.bsky.social, we integrate scRNA-seq & WGS to uncover how CNAs drive tumor evolution and transcriptional variability.
18.12.2024 13:30 — 👍 15 🔁 6 💬 2 📌 2
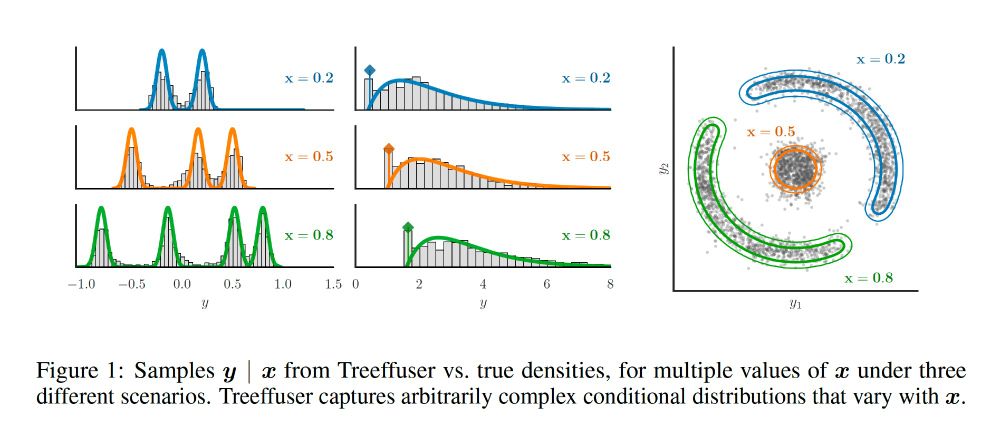
Samples y | x from Treeffuser vs. true densities, for multiple values of x under three different scenarios. Treeffuser captures arbitrarily complex conditional distributions that vary with x.
I am very excited to share our new Neurips 2024 paper + package, Treeffuser! 🌳 We combine gradient-boosted trees with diffusion models for fast, flexible probabilistic predictions and well-calibrated uncertainty.
paper: arxiv.org/abs/2406.07658
repo: github.com/blei-lab/tre...
🧵(1/8)
02.12.2024 21:48 — 👍 154 🔁 23 💬 4 📌 4
I've followed just the right set of accounts that my feed is just pictures of space and cats doing weird stuff. I'll add more, but for now... it's perfect
25.11.2024 14:51 — 👍 2440 🔁 58 💬 110 📌 3
Your Jeopardy! pal. Author of THE COMPLETE KENNECTIONS (http://bit.ly/4qUcbhK) and a bunch of other stuff.
Assistant Professor, BME, Columbia University
Immunologist at the Ido Amit, Nir Yosef and Robert Zeiser labs. Formerly PhD Student in Burkhard Bechers lab. Interested in innovative single cell technologies and computational biology.
trying to learn bio and computation at Berkeley
CS PhD @ UCLA
https://web.cs.ucla.edu/~wob/
Biomedical Informatics PhD candidate @ Harvard
ELLIS PhD student @HelmholtzMunich, Student Researcher @Apple. Interested in ML, Single-Cell Genomics, and People.
PhD candidate@Columbia University.
Generative and Geometric deep learning | Theory of Generative Models| Cancer systems biology
Research at Azizi lab & Califano lab.
PhD candidate in BME @Columbia | computational #cancer research at http://azizilab.com
T32 GEIC Postdoctoral research scientist co-mentored by @elhamazizi and @joselmcfaline @Cancer_dynamics, Ph.D. @columbiaBME, NSF Graduate Research Fellow (2019-2022)
Ray and Stephanie Lane Professor of Computational Biology at CMU School of Computer Science. https://www.cs.cmu.edu/~jianma/
Machine Learning Professor
https://cims.nyu.edu/~andrewgw
Postdoctoral fellow at Harvard Data Science Initiative | Former computer science PhD at Columbia University | ML + NLP + social sciences
https://keyonvafa.com
Machine learning PhD student @ Blei Lab in Columbia University
Working in mechanistic interpretability, nlp, causal inference, and probabilistic modeling!
Previously at Meta for ~3 years on the Bayesian Modeling & Generative AI teams.
🔗 www.sweta.dev
Machine learning lab at Columbia University. Probabilistic modeling and approximate inference, embeddings, Bayesian deep learning, and recommendation systems.
🔗 https://www.cs.columbia.edu/~blei/
🔗 https://github.com/blei-lab
Machine Learning PhD Student
@ Blei Lab & Columbia University.
Working on probabilistic ML | uncertainty quantification | LLM interpretability.
Excited about everything ML, AI and engineering!
Computer Science PhD candidate at Columbia University. Interested in applying deep learning to functional genomics. NSF GRFP Fellow
Statistician,Mother,Prof,Grandmother(she/her)🇫🇷🇵🇹🇺🇸🧪.
🖋️with @wkhuber.bsky.social:Modern Statistics for Modern Biology (https://www.huber.embl.de/msmb/) #stats,#rstats.OSS,arXiv, microbial ecology,cytof,multi-omics,Bioconductor:phyloseq, DADA2





