
GitHub - lab-cosmo/upet: Universal interatomic potentials for advanced materials modeling
Universal interatomic potentials for advanced materials modeling - lab-cosmo/upet
Now, this is just a milestone on that path, but it's already something worth sharing, so thanks to arXiv you can read about it arxiv.org/html/2603.02..., and thanks to uPET github.com/lab-cosmo/up... and metatomic, you can try already a universal MLIP trained on it.
03.03.2026 11:15 —
👍 2
🔁 2
💬 1
📌 0

📢 We have been working on a new universal atomistic dataset that combines the principles of MAD with a meta-GGA level of theory, so we can all simulate water that does not freeze at 500K 🧊 , and have all our bases covered, with reference data for every isotope with a half-life above 24 hours ☢️
03.03.2026 11:15 —
👍 11
🔁 3
💬 1
📌 0
So let us show you just how *universal* #PET-MAD-1.5 can be. This is a movie of a parallel tempering simulation, with replicas from 300K to 3000K, of what we call a "Mendeleev cluster" - one atom each of every element from 1 to 102.
03.03.2026 11:45 —
👍 10
🔁 5
💬 2
📌 1
ShiftML-3 predictor
No Install. No Setup. Just Chemical Shift Predictions.
At shiftml.materialscloud.io we host the latest ShiftML3 in the web and you can predict chemical shifts of organic crystals for free in the web!
17.02.2026 17:21 —
👍 3
🔁 2
💬 0
📌 0

Zooming in on a large-scale dataset to showcase the new adaptive resolution features in chemiscope


📢 chemiscope.org 1.0.0rc1 just dropped on pypi! We are making (a few) breaking changes to the interfaces, fixing a ton of bugs and introducing some exciting features (you can finally load datasets with > 100k points!). We'd be grateful if you test, break and report 🐛 github.com/lab-cosmo/ch...
05.01.2026 14:42 —
👍 4
🔁 2
💬 0
📌 0
Very nice! Is this able to use WebGPU when running the model?
Also, I guess this is limited to 4GiB of memory? Although it might not matter too much, since it would become very slow for systems that require that kind of memory anyway
28.11.2025 11:26 —
👍 1
🔁 0
💬 1
📌 0
GitHub - lab-cosmo/shiftml: A python package for the prediction of chemical shieldings of organic solids and beyond.
A python package for the prediction of chemical shieldings of organic solids and beyond. - lab-cosmo/shiftml
We're introducing ShiftML3, a new ShiftML model for chemical shielding predictions in organic solids.
* ShiftML3 predicts full chemical shielding tensors
* DFT accuracy for 1H, 13C, and 15N
* ASE integration
* GPU integration
Code: github.com/lab-cosmo/Sh...
Install from Pypi: pip install shiftml
25.08.2025 08:53 —
👍 2
🔁 3
💬 1
📌 0
If you are using or you are considering using CP2K, check this out!
25.08.2025 10:15 —
👍 3
🔁 1
💬 0
📌 0
The dark side of the forces: assessing non-conservative force...
The use of machine learning to estimate the energy of a group of atoms, and the forces that drive them to more stable configurations, have revolutionized the fields of computational chemistry and...
Very proud to send Filippo Bigi to Vancouver to give an oral presentation at @icmlconf.bsky.social about our investigation of the use of "dark-side forces" in atomistic simulations. The final version is here openreview.net/forum?id=OEl... and it's worth a read even if you already read the #preprint
20.06.2025 15:53 —
👍 10
🔁 5
💬 1
📌 2
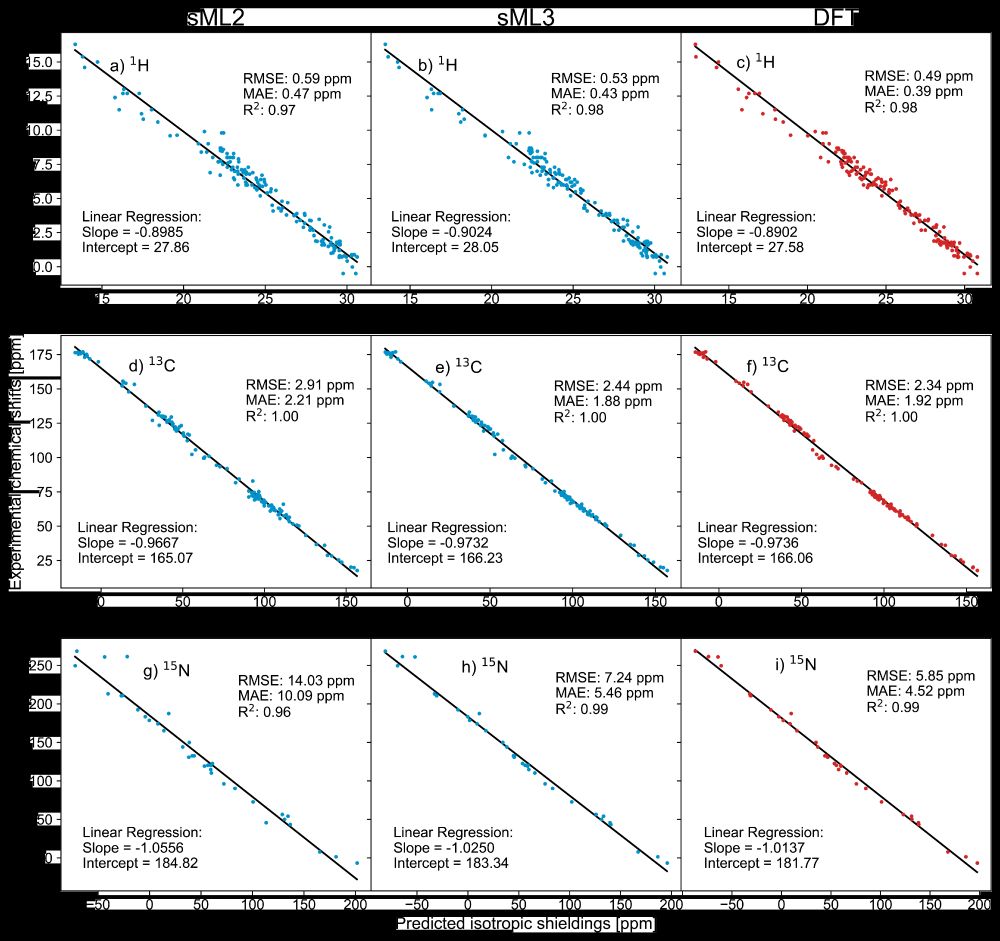
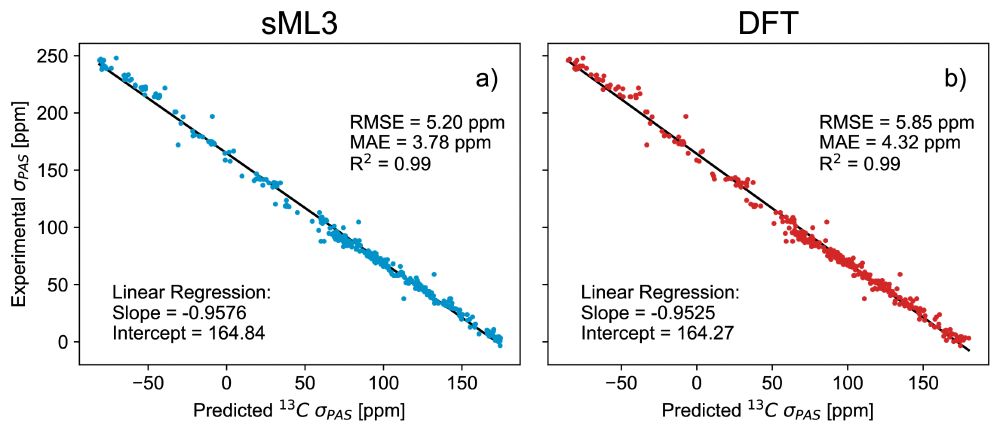
🎉 DFT-accurate, with built-in uncertainty quantification, providing chemical shielding anisotropy - ShiftML3.0 has it all! Building on a successful @nccr-marvel.bsky.social-funded collaboration with LRM🧲⚛️, it just landed on the arXiv arxiv.org/html/2506.13... and on pypi pypi.org/project/shif...
17.06.2025 13:18 —
👍 18
🔁 7
💬 1
📌 0
The link above is missing a password, pleas use epfl.zoom.us/j/6836877674... instead!
13.06.2025 07:42 —
👍 0
🔁 0
💬 0
📌 0
Long-stride trajectories with a universal FlashMD model - The Atomistic CookbookContentsMenuExpandLight modeDark modeAuto light/dark, in light modeAuto light/dark, in dark mode
We wouldn't be @labcosmo.bsky.social if we didn't want you to go break it, so head to atomistic-cookbook.org/examples/fla... for a crash-course 🧑🍳📖 recipe, but not before reading the warnings arxiv.org/html/2505.19.... Have fun! #compchem #machinelearning #md @nccr-marvel.bsky.social @erc.europa.eu
27.05.2025 07:02 —
👍 1
🔁 1
💬 0
📌 0
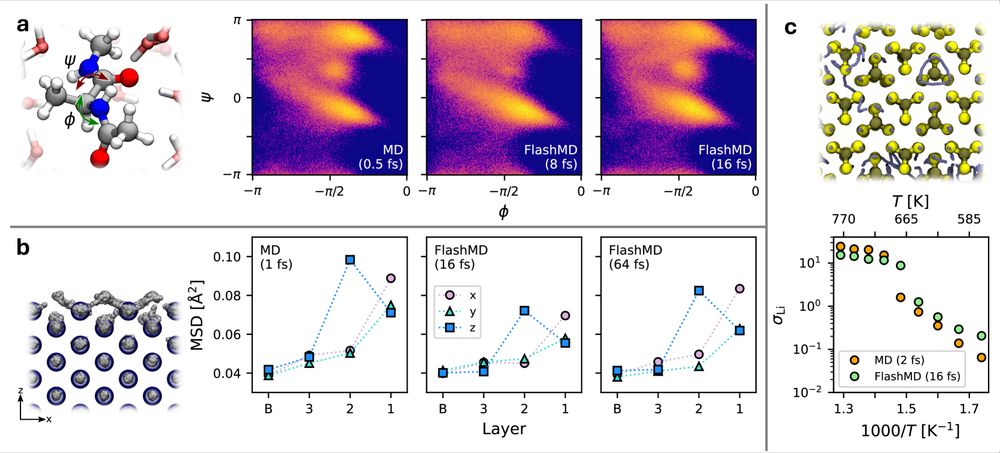
If you do, the rewards can be very impressive: you can run solvated alanine dipeptide and observe superionic behavior in LiPS with 16fs time step, and watch the Al(110) surface pre-melt in strides of 64fs. And all with the same universal model, no fine-tuning needed!
27.05.2025 07:02 —
👍 1
🔁 1
💬 1
📌 0

Darth Vader dressed as Santa presents a non-conservative forcefield
Much as for direct force prediction [ arxiv.org/html/2412.11... ] you better know what you are doing: you've no guarantee of energy conservation, or of equipartition, so you should know your thermostats VERY well. Caveat emptor.
27.05.2025 07:02 —
👍 0
🔁 1
💬 1
📌 0
PET-MAD, a universal interatomic potential for advanced materials modeling
Filippo's idea was to use the heavy-duty PET-MAD model [ arxiv.org/html/2503.14... ] to generate a bunch of trajectories of wildly different compounds and use a PET-like architecture to learn (q',p') from (q,p), and and to think A LOT about the many things that could possibly go wrong.
27.05.2025 07:02 —
👍 0
🔁 1
💬 1
📌 0
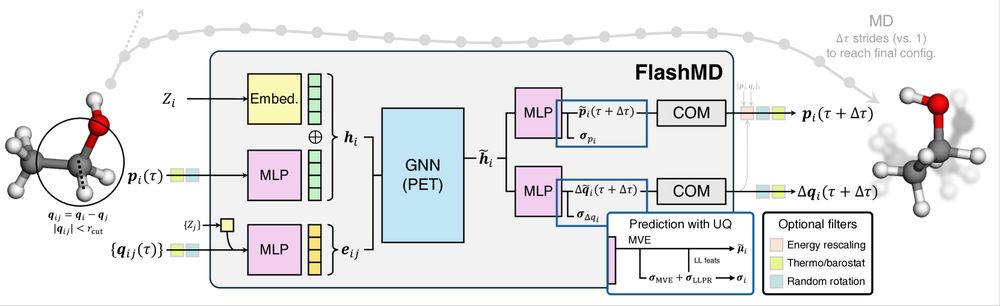
Scheme of the GNN architecture of the FlashMD method.
📢 Running molecular dynamics with time steps up to 64fs for any atomistic system, from Al(110) to Ala2? Thanks to 🧑🚀 Filippo Bigi and Sanggyu Chong, with some help from Agustinus Kristiadis, this is not as crazy as it sounds. Let us briefly introduce FlashMD⚡ arxiv.org/html/2505.19...
27.05.2025 07:02 —
👍 37
🔁 12
💬 1
📌 1
If you don't have time for the 20-pages appendix, the TL;DR is that approximating tensors is harder than the vector case, but can be made as simple as possible using angular momentum theory. The practical implementation we propose is not as rigorous, but works well and is very fast in practice.
09.05.2025 06:50 —
👍 1
🔁 1
💬 1
📌 0
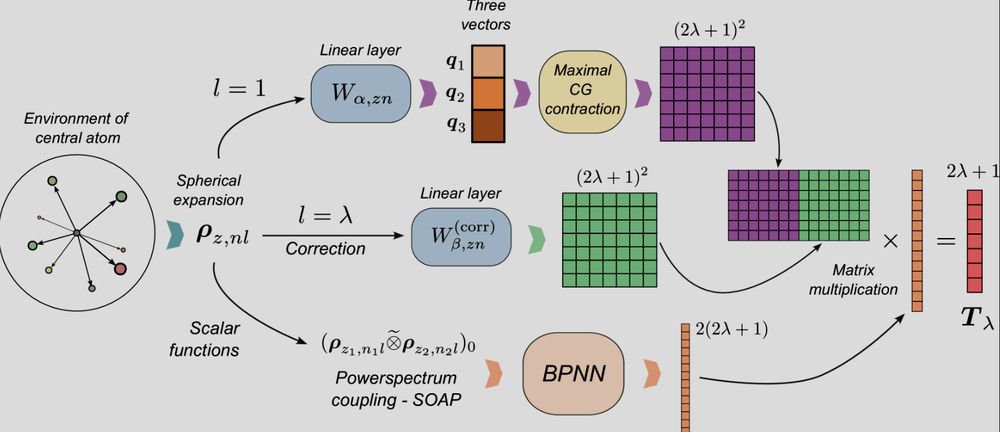
Schematic representation of the lambda-MCoV architecture for predicting tensorial quantities using scalar approximators
First #preprint from recent 🧑🚀 Michelangelo Domina (+ @ppegolo.bsky.social and Filippo) provides theoretical foundations and practical architectures to build scalar-function-based approximations of tensors, in the spirit of the "scalars are universals" paper #compchem arxiv.org/html/2505.05...
09.05.2025 06:50 —
👍 4
🔁 2
💬 1
📌 0
Reversible multiple time scale molecular dynamics
The Trotter factorization of the Liouville propagator is used to generate new reversible molecular dynamics integrators. This strategy is applied to derive reve
You can use these safely in MD, by multiple time stepping (a '90s classic: doi.org/10.1063/1.46...) so you get reliable conservative trajectories, for any material, twice as fast! Let us know if it works for you, and even more importantly, if it doesn't! 👉 atomistic-cookbook.org/examples/pet...
07.05.2025 05:24 —
👍 2
🔁 1
💬 0
📌 0
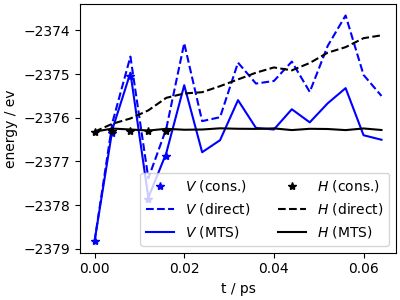
Plot of the potential and total energy trends along a MD trajectory, showing drift for non-conservative forces, and how that is fixed using multiple time stepping.
The PET-MAD universal forcefield mingled with the dark side, and got twice as fast 🚀. Read on, or head to the 🧑🍳📖 atomistic-cookbook.org/examples/pet..., if you are curious of what this is all about. #atomistic-cookbook #compchem #machinelearning #mlip🧵
07.05.2025 05:24 —
👍 10
🔁 3
💬 1
📌 0
Header for the atomistic-cookbook.org recipe for the PET-MAD universal potential
You can read all about it here, arxiv.org/abs/2503.14118, see it in action as a #cookbook recipe 🧑🍳 atomistic-cookbook.org/examples/pet... or rush to install it following the instructions here github.com/lab-cosmo/pe...
19.03.2025 07:23 —
👍 2
🔁 1
💬 1
📌 0
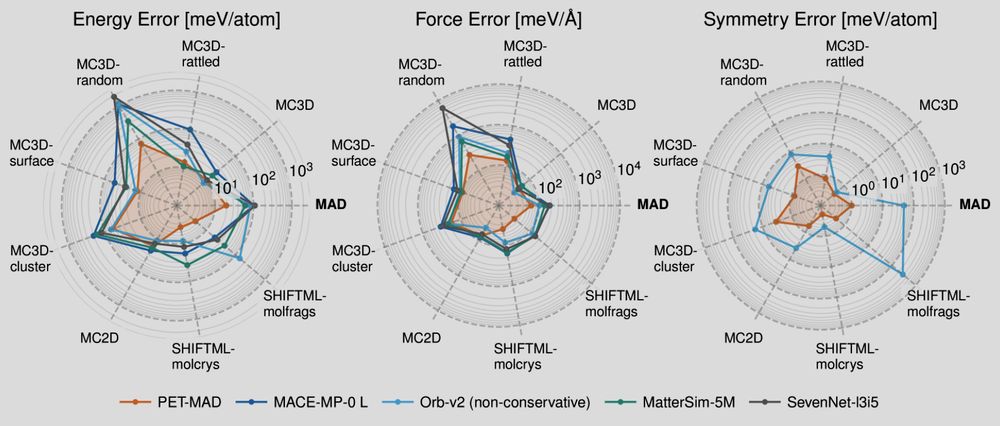
Polar plot showing the errors of several machine-learning potential of different test sets. Smaller is better here!
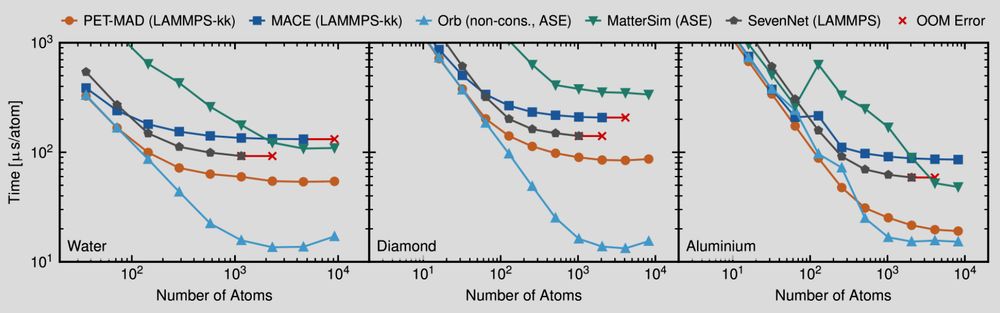
Plots showing the evaluation time per atom for several machine-learning potentials as a function of the number of atoms in a simulation. Smaller is better
📢 PET-MAD has just landed! 📢 What if I told you that you can match & improve the accuracy of other "universal" #machinelearning potentials training on fewer than 100k atomic structures? And be *faster* with an unconstrained architecture that is conservative with tiny symmetry breaking? Sounds like 🧑🚀
19.03.2025 07:23 —
👍 28
🔁 9
💬 1
📌 3
Should be super-easy to adapt to whatever tensor you're trying to learn (and to extend to more sophisticated models than symmetry-adapted regression). Let us know what you cook with it 😋!
13.03.2025 17:30 —
👍 1
🔁 1
💬 0
📌 0
In particular, this recipe relies on metatensor.github.io/featomic to compute the features, and the docs.metatensor.org/latest/torch... backend of #metatensor to export a self-contained ASE-compatible calculator. Easy to use, fast, and accurate.
13.03.2025 17:30 —
👍 1
🔁 1
💬 1
📌 0
Polarizability of a small molecule, represented as an ellipsoid
🧑🍳 A new #cookbook recipe to train, export and use a simple but effective linear model for tensorial properties - specifically molecular polarizabilities
atomistic-cookbook.org/examples/pol...
Thanks @ppegolo.bsky.social for the recipe, and the whole 🧑🚀 team for the underlying infrastructure.
13.03.2025 17:30 —
👍 10
🔁 3
💬 1
📌 0





















