
Systemic inflammation impairs myelopoiesis and interferon type I responses in humans
www.nature.com/articles/s41...
@natimmunol.bsky.social
@hemagene.bsky.social

Systemic inflammation impairs myelopoiesis and interferon type I responses in humans
www.nature.com/articles/s41...
@natimmunol.bsky.social

There’s an arms race going on between a disease-causing fungus and its host, and it’s not the one portrayed in HBO’s postapocalyptic series The Last of Us. #ScienceMagArchives scim.ag/42QS3Un
18.04.2025 17:44 — 👍 60 🔁 13 💬 0 📌 1
The Science Visuals team has been honored with three
@commarts.com Awards of Excellence.
These prestigious awards recognize creative excellence and outstanding execution in visual storytelling. (THREAD 🧵) scim.ag/4jDJ6TY

A great ressource from the Khoriaty lab at University of Michigan to understand deeper the molecular regulation of erythropoiesis, this week in @natcomms.nature.com
www.nature.com/articles/s41...
Merci Ismael 😉
05.04.2025 15:12 — 👍 0 🔁 0 💬 0 📌 0
Scalable spatial transcriptomics through computational array reconstruction - @broadinstitute.org go.nature.com/3QYfSDd
03.04.2025 15:04 — 👍 36 🔁 9 💬 1 📌 0Our work nicely complements recently released papers from @thenormanlab www.cell.com/cell-systems... and @satijalab www.nature.com/articles/s41... that describe the value of multimodal perturb-seq using CRISPRi in cell lines to decipher gene regulatory networks ! 🧬 🧫
03.04.2025 21:25 — 👍 0 🔁 0 💬 0 📌 0We are also grateful to the @broadinstitute.org and the @GRO_Broad for their support, and to our editor and reviewers at @science.org! (end)
03.04.2025 21:20 — 👍 1 🔁 0 💬 0 📌 0I want to sincerely thank all our amazing co-authors and, in particular, my wonderful co-first author Jorge Martin-Rufino (@jmartinrufino), as well as my mentors @bloodgenes.bsky.social and @eric_lander. (12/n)
03.04.2025 21:20 — 👍 1 🔁 0 💬 1 📌 0Our findings reveal that heritable variation is predominantly enriched in specific gene regulatory networks controlled by master TFs, offering novel insights into the genetic architecture of complex traits. Protocols and much more can be found in our manuscript! (11/n)
03.04.2025 21:20 — 👍 2 🔁 0 💬 1 📌 0
Most strikingly, we discovered that TF-sensitive regions comprise <0.3% of the genome but are enriched ~100-fold for SNPs associated with certain blood-cell phenotypes, which is remarkably higher than that of any other chromatin elements active throughout hematopoiesis. (10/n)
03.04.2025 21:20 — 👍 1 🔁 0 💬 1 📌 0
…such as a GATA1-dependent regulatory network comprising the CPEB4 gene and an enhancer harboring variants with multiple associations to blood cell traits, exemplifying how Perturb-multiome can help systematically pinpoint associations between phenotypes and GWAS variants. (9/n)
03.04.2025 21:20 — 👍 1 🔁 0 💬 1 📌 0
For example, we faithfully identified various well-characterized gene regulatory mechanisms in hematopoiesis, such as the BCL11A-mediated fetal-to-adult hemoglobin switch. Importantly, it also uncovered thousands of previously uncharacterized regulatory interactions… (8/n)
03.04.2025 21:20 — 👍 1 🔁 0 💬 1 📌 0
As such, Perturb-multiome allowed us to reconstruct transcription factor-dependent gene regulatory networks throughout hematopoietic differentiation, capturing the complexity of the differentiation continuum and enabling a detailed understanding of regulatory mechanisms. (7/n)
03.04.2025 21:20 — 👍 1 🔁 0 💬 1 📌 0
Perturb-multiome is highly effective in disease-relevant primary cells, such as hematopoietic stem and progenitor cells (which give rise to all blood cells) by overcoming challenges in perturbation efficiency and identification. (6/n)
03.04.2025 21:20 — 👍 1 🔁 0 💬 1 📌 0
With Perturb-multiome we can now generate high-resolution analyses of gene regulatory networks in single cells, which enables genome-wide variant-to-function inferences. (5/n)
03.04.2025 21:20 — 👍 1 🔁 0 💬 1 📌 0
To achieve this, we developed Perturb-multiome, which recovers from each cell: (1) the identity of the genetic perturbation (sgRNA), (2) the effect on chromatin accessibility at cis-regulatory elements (scATAC-seq), and (3) the effect on expression of genes (scRNA-seq). (4/n)
03.04.2025 21:20 — 👍 1 🔁 0 💬 1 📌 0
Rather than recreating each variant, which is hard to scale, given the enrichment of noncoding causal variants in regulatory regions bound by transcription factors (TFs), we reasoned that systematic perturbation of TF networks in single cells could provide new insights (3/n)
03.04.2025 21:20 — 👍 1 🔁 0 💬 1 📌 0While GWAS have been successful in identifying a large number of genetic variants associated with complex traits, biological insights gained have been limited. This is largely due to the lack of systematic methods for linking these variants to their functional impacts. (2/n)
03.04.2025 21:20 — 👍 2 🔁 0 💬 1 📌 0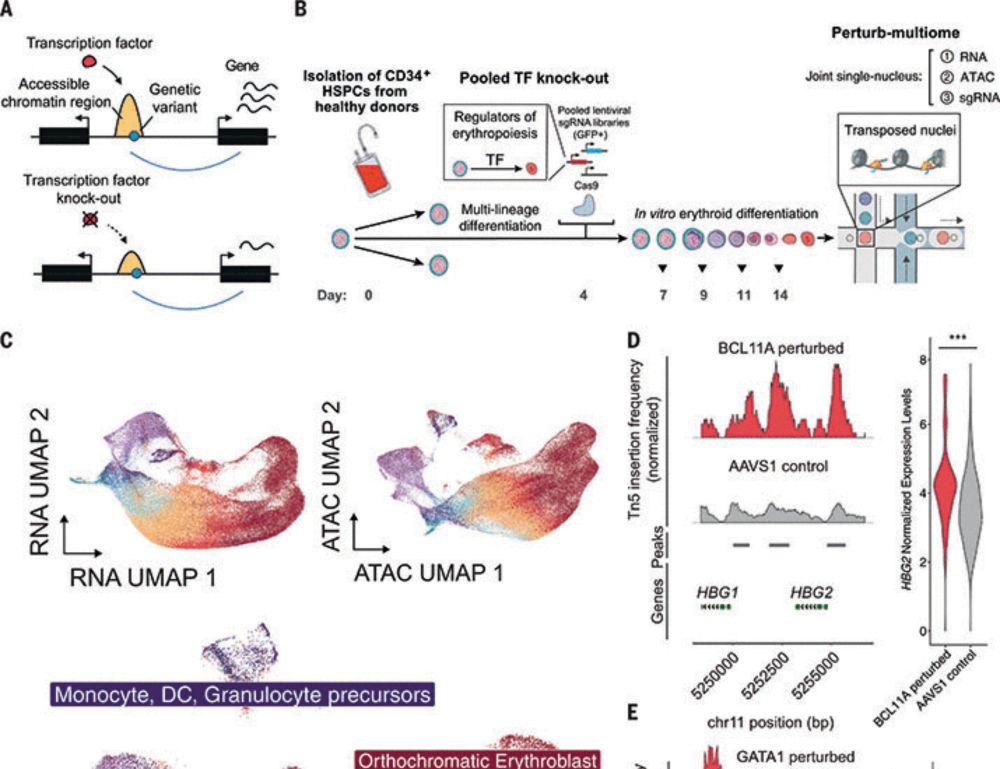
Out today in @science.org!
What if you could chart cells' regulatory programs at unprecedented resolution?
In my work with Jorge Martin-Rufino from the @bloodgenes.bsky.social lab, we dissect the genome’s control circuits and find where key genetic variation hides
bit.ly/3YhBMoO

Congratulations to @danafarber.bsky.social’s Lara Wahlster, MD, PhD, and Nina Weichert-Leahey, MD, for receiving the American Society for Clinical Investigation Emerging-Generation Awards. Read about Dr. Wahlster at bit.ly/4haxdTW and Dr. Weichert-Leahey at bit.ly/4h6BYxL.
06.03.2025 19:26 — 👍 4 🔁 2 💬 0 📌 0