A wonderful first for me at the upcoming @biophysicalsoc.bsky.social meeting in SF is having many lab members present! See 6 brilliant graduate students postdocs from the lab present talks and posters on how they are pushing the boundaries of technology and mechanistic membrane protein biology
20.02.2026 18:26 — 👍 14 🔁 4 💬 1 📌 0

Two #CRISPR talks on Feb3!
1️⃣ Stephan Riesenberg (Group Leader, Max Planck Leipzig): CRISPR-mediated generation of genetic variants for functional analysis.
2️⃣ Yuriy Baglaenko @baglaenkolab.bsky.social: Causal variants with CRISPR editing in primary human cells www.varianteffect.org/seminar-seri...
28.01.2026 17:50 — 👍 3 🔁 2 💬 1 📌 1

Specificity, length and luck drive gene rankings in association studies - Nature
Genetic association tests prioritize candidate genes based on different criteria.
How do GWAS and rare variant burden tests rank gene signals?
In new work @nature.com with @hakha.bsky.social, @jkpritch.bsky.social, and our wonderful coauthors we find that the key factors are what we call Specificity, Length, and Luck!
🧬🧪🧵
www.nature.com/articles/s41...
07.11.2025 00:05 — 👍 171 🔁 74 💬 5 📌 11

🎙️ Next up Dec 2 in VESS!
Thea Schulze (Lindorff-Larsen Lab): Predicting mutated protein abundance @tkschulze.bsky.social
Taylor Mighell (Lehner Lab): Massive mutagenesis to understand GPCRs @taylor-mighell.bsky.social
🔗 More info at varianteffect.org/seminar-series
@varianteffect.bsky.social
04.11.2025 18:31 — 👍 5 🔁 4 💬 0 📌 0
MSS26
Mutational Scanning Symposium 2026, 25-27 March, Melbourne
🌏 🧬 Join the global functional genomics community in Melbourne this March for MSS26! The 9th Annual Mutational Scanning Symposium #VariantEffect26 runs March 25–27, 2026 at the Aikenhead Centre for Medical Discovery.
🗓️ Early bird registration & abstract submissions close Nov 2, 2025.
www.mss2026.org
10.10.2025 14:40 — 👍 3 🔁 1 💬 0 📌 0

Coming up next in the VESS (Nov 4):
Population genetics × variant effects:
🧬 Nikhil Milind (Stanford) on gene dosage and complex traits @nikhilmilind.dev
🧬 Leslie Smith (U Florida) on equitable ML in cancer genomics
www.varianteffect.org/seminar-seri...
@varianteffect.bsky.social
10.10.2025 14:43 — 👍 5 🔁 1 💬 0 📌 1

Postdoc Description.docx
Title: Postdoctoral Associate Summary statement: The postdoctoral research associate is responsible for developing novel computational methodology for high-throughput sequence genomics tasks, as well ...
Have you recently completed (or finishing soon) a PhD in CS or a related discipline? Do you want to do research advancing the theory & practice of algorithmic genomics & build tools that people love to use? I'll be looking to hire a postdoc! Official ad coming soon:
docs.google.com/document/d/1...
08.10.2025 16:03 — 👍 14 🔁 17 💬 0 📌 2

VESS is happening tomorrow (Oct 7). See you there!
First speaker: Shelby Hemker (Dr. Jacob Kitzman Lab, University of Michigan)
Second speaker: Karl Romanowicz (Dr. Calin Plesa Lab, University of Oregon) @kroman.bsky.social
Link: www.varianteffect.org/seminar-seri...
06.10.2025 17:05 — 👍 1 🔁 0 💬 0 📌 0
The first (of hopefully many) reports to come from our collaboration with @hjp.bsky.social
We present a new type of cell fitness assay that allows you to both quantify and explain differences across human donors in cell proliferation and sensitivity to environmental toxicants.
30.09.2025 19:17 — 👍 17 🔁 7 💬 1 📌 0
Super excited to have this out. Thanks very much to the reviewers who helped improve this manuscript. Congrats to @jingyour.bsky.social!
bsky.app/profile/bioi...
22.09.2025 17:35 — 👍 20 🔁 5 💬 1 📌 0

We are excited to share GPN-Star, a cost-effective, biologically grounded genomic language modeling framework that achieves state-of-the-art performance across a wide range of variant effect prediction tasks relevant to human genetics.
www.biorxiv.org/content/10.1...
(1/n)
22.09.2025 05:29 — 👍 175 🔁 90 💬 4 📌 5

What we talk about when we talk about risk
How embryo selection exploits our flawed intuitions about risk
I wrote about how genetic risk works in the context of embryo selection and how people often think about it all wrong. A short 🧵:
10.08.2025 19:07 — 👍 77 🔁 32 💬 3 📌 1


@jengreitz.bsky.social l & my lab want to co-hire a computational biologist/biostatistician with project management expertise to help map the regulatory code of the human genome and discover genetic mechanisms of disease.
Details below
careersearch.stanford.edu/jobs/computa...
Plz RT
19.08.2025 00:29 — 👍 55 🔁 60 💬 1 📌 1

Why yes! You can watch previous Variant Effects Seminar Series talks on our YouTube channel!
ℹ️ www.varianteffect.org/previous-sem... 📺 www.youtube.com/playlist?lis...
#Genomics #Seminar #EarlyCareerResearchers #ScientificSeminar #PrecisionMedicine
19.08.2025 01:34 — 👍 8 🔁 6 💬 0 📌 0
Very excited to have this out! Here, we take inspiration from model selection MR to decouple direct and indirect effects in DMS experiments. Check out Jingyou's explainer and paper below:
bsky.app/profile/jing...
15.08.2025 16:22 — 👍 20 🔁 5 💬 0 📌 0
Bittersweet to be leaving @docedge.bsky.social after a wonderful postdoc, but excited to share that I'm joining @uoregon.bsky.social next month as an Assistant Professor in the Department of Data Science.
06.08.2025 17:18 — 👍 146 🔁 28 💬 21 📌 4

We applied Cosmos to 3 DMS datasets:
Kir2.1: abundance → surface expression
PDZ3: abundance → CRIPT binding
KRAS: abundance → RAF1_RBD binding
Cosmos clearly separates direct binding residues from those with indirect effects.
[6/n]
04.08.2025 15:10 — 👍 0 🔁 0 💬 1 📌 0
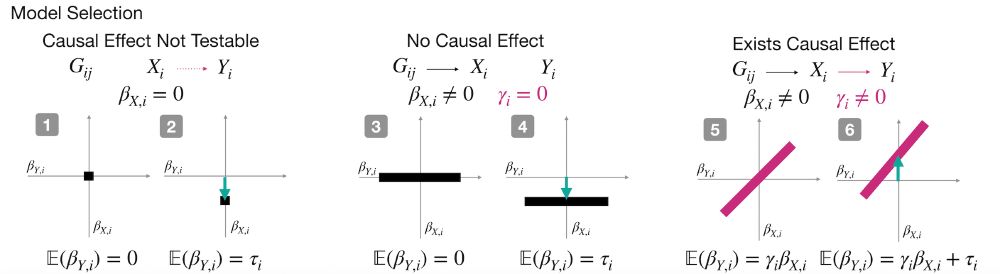
How does it work?
Cosmos aggregates mutation effects by position, learns interpretable causal graphs via Bayesian model selection, and outputs residue-level direct vs indirect effects.
[5/n]
04.08.2025 15:10 — 👍 0 🔁 0 💬 1 📌 0
Cosmos answers three key questions:
1️⃣ Is there a causal link between the phenotypes?
2️⃣ How strong is it?
3️⃣ What would the downstream phenotype look like if we “normalized” the upstream one (i.e., counterfactual inference)?
[4/n]
04.08.2025 15:09 — 👍 1 🔁 0 💬 1 📌 0

Cosmos is a Bayesian framework that performs residue-level causal inference to decouple how mutations influence upstream vs downstream functions.
No need for detailed biophysical models—just stats and data.
[3/n]
04.08.2025 15:09 — 👍 0 🔁 0 💬 1 📌 0
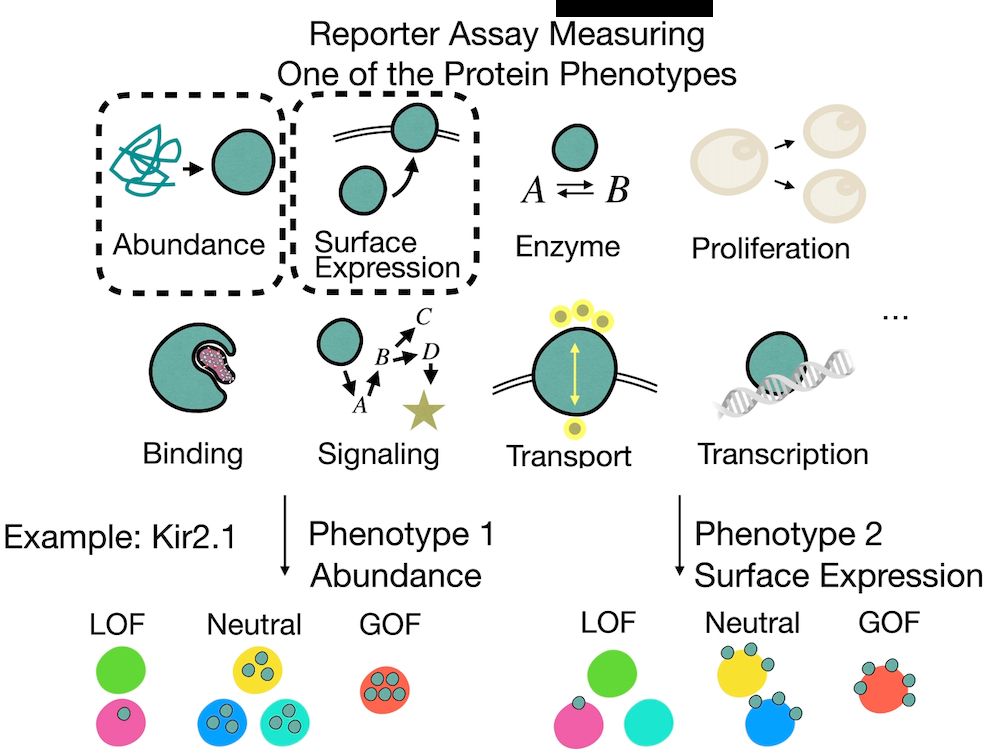
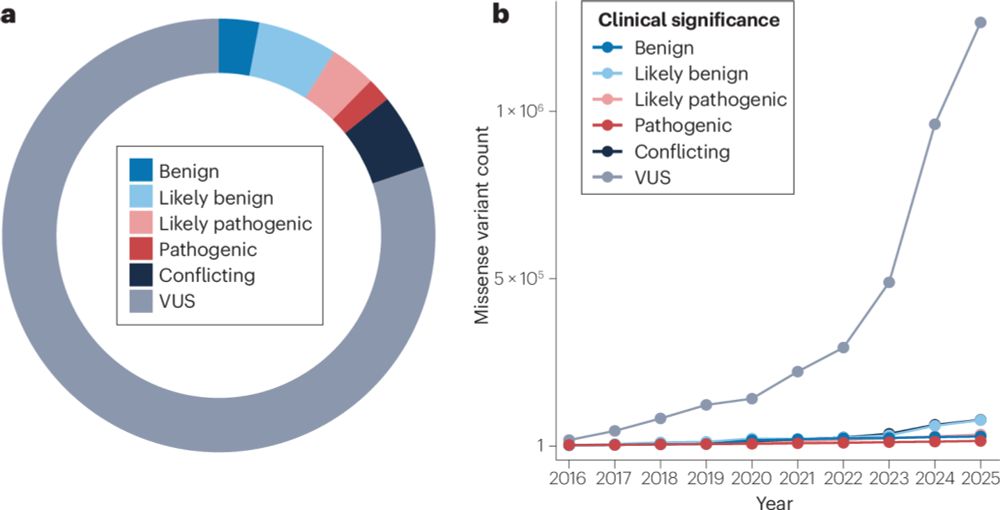
Multi-phenotype DMS experiments are revealing how mutations impact different protein functions.
But these phenotypes are often causally linked—e.g., when measuring activity, we may also capture effects propagated from abundance.
So how do we tell what’s direct vs indirect?
[2/n]
04.08.2025 15:09 — 👍 0 🔁 0 💬 1 📌 0
Home - ProbGen 2026
Your Site Description
The 2026 Probabilistic Modeling in Genomics (ProbGen) meeting will be held at UC Berkeley, March 25-28, 2026. We have an amazing list of keynote speakers and session chairs:
probgen2026.github.io
Please help spread the news.
06.06.2025 17:52 — 👍 69 🔁 36 💬 2 📌 0

Perfect first day: receiving a set of new pipettes! Can’t wait to do more cool experiments with the lab for the next few years. @willowcoyote.bsky.social
31.07.2025 22:57 — 👍 3 🔁 0 💬 0 📌 1
Super proud of my first student, @jingyour.bsky.social! Well done! Looking forward to the amazing work you will do in the future 🥲 bsky.app/profile/jing...
21.07.2025 23:15 — 👍 31 🔁 7 💬 0 📌 0

Thrilled to share that I just successfully defended my PhD! Thanks to my committee, collaborators, and everyone who’d supported me throughout my seven years at UCLA. A special thank you to my PI Harold for his incredible mentorship! @hjp.bsky.social
21.07.2025 21:35 — 👍 287 🔁 14 💬 16 📌 2
Very excited to have this work out by @jeromics.bsky.social ! Please check it out.
I think my favorite story from the supplement is how impactful normalization can be in this context.
bsky.app/profile/jero...
02.07.2025 15:56 — 👍 8 🔁 5 💬 0 📌 0
Harvard Neuro PhD student in Beth Stevens’ lab | Studying interactions between genes, environments, and neurological dysfunction | NSF & HHMI Gilliam fellow 🏳️🌈
PhD Candidate in the Pritchard Lab at Stanford University. Interested in statistical and population genetics.
https://nikhilmilind.dev/
genetics, genomics, and gossip • NHGRI #NIHMOSAIC K99/R00 fellow at Michigan, formerly at MIT/UNC
https://adelaidetovar.com
Postdoc at Siepel Lab, CSHL
Statistical genetics, phylogenetics, and optimization
Postdoc in Zoghbi Lab at Baylor College of Medicine | Interested in gene regulation, neurogenetics, variant interpretation, and disease mechanism | Opinions are my own
Postdoc at @crg.eu Barcelona,
previously PhD at Cambridge University
Genomics | Mutational Scanning | Molecular Evolution 🧬🌱
www.pyrisentinel.eu ⛰️🌊 www.puntseq.com
Same content, different website! Human evolutionary genomics, functional genomics and archaic hominins. Group leader in Human Genomics at SVI in Melbourne/Naarm, Australia. Durian evangelist. Sometimes I go to Estonia.
PhD candidate - Machine Learning and Genomics @CRG.eu with @jonnyfrazer.bsky.social and @MafaldaFigDias
Doing science @UCSF in the Coyote-Maestas and Manglik Labs. Former Jackrel Lab @WUSTL www.matthewkhoward.com
@CompMedUcla PhD Student | Somewhere in between EHR + Genetics 🧬
Assistant Professor @ UCLA Pathology, Computational Medicine, Biostatistics, and Genetics
PhDing at the Sanger Institute, i'm evolving every day
🌼 isabellezane.bio
Assistant professor of Psychiatry, Human Genetics, and Computational Medicine at UCLA
Probabilistic machine learning to address questions in evolution and health #EvolutionaryMedicine. PI at the Centre for Genomic Regulation, co-leading a group with Mafalda Dias. Previously Harvard.
🇮🇹, Computational biophysicist, NNF Postdoc in #OPIG at University of Oxford | previously PhD and PostDoc at University of Copenhagen (KLL group)
Sudmantlab.org - Assistant professor Berkeley
Stanford CS PhD student working on ML/AI for genomics with @anshulkundaje.bsky.social
austintwang.com
Scientist, #MachineLearning and #AI for Moleculear Sciences. Scuba Diver. Loves @cecclementi.bsky.social