
A wonderful collection of contemporary topics and perspectives on secretion and Golgi biology edited by Akihiko Nakano and Jaakko Saraste.
The Golgi Network, Volume II @SN and link.springer.com/book/10.1007...
@natmachinery.bsky.social
Prof. Univ. of Bergen & Helse Bergen | Basic & translational science | Molecular Biology | Cancer | Endocrinology | Rare diseases | Cytoskeleton | Ribosome | Protein biogenesis | PTMs | N-terminal acetylation https://www.uib.no/en/rg/protein

A wonderful collection of contemporary topics and perspectives on secretion and Golgi biology edited by Akihiko Nakano and Jaakko Saraste.
The Golgi Network, Volume II @SN and link.springer.com/book/10.1007...

My review “Co-translational control of protein stability and quality in plants” is now online at @jxbotany.bsky.social, in which I describe how co-translational processing and ribosome-associated quality control together establish protein stability and fate early in synthesis. tinyurl.com/5dhhz9h6
24.02.2026 09:45 — 👍 41 🔁 24 💬 0 📌 0Indeed! Feel free to suggest here or to ISPT. Also, many speakers will be selected from Abstracts so this route is open.
23.02.2026 10:40 — 👍 0 🔁 0 💬 0 📌 0
8 days left to register | 28 invited speakers | Keynotes from F. Ulrich Hartl, Roland Beckmann and Michael Rapé | #chaperones #degradation #ubiquitin #cryoEM #acetylation #lipidation #ribosomes #proteins
proteintermini.org/meeting/

Actin N-terminal maturation can be blocked by a butynamide stereoprobe which acts as an ACTMAP inhibitor. Preprint by Xiong et al. Benjamin Cravatt and Bruno Melillo. @ispt-proteinterm.bsky.social
www.biorxiv.org/content/10.6...

Join us in Palermo!
FEBS Workshop Protein Termini 2026: the power of protein termini across bacteria, plants, and animals—from ribosome biology to proteostasis and applications. Deadline: 3 March 2026
proteintermini.org/meeting
#ProteinTermini #Proteostasis #FEBS #EMBO @iubmb.bsky.social
Join us at the @crick.ac.uk for the 2026 meeting of the UK proteostasis community!
We especially encourage students and postdocs to attend and share their work. All talks (except the keynotes) will be selected from abstracts.
Surprisingly: "Conditional stability of HY5 through the ATE N-degron pathway regulates environmental responses in Arabidopsis thaliana".
The shining bounds of N-degron pathway influence expands! @charlene-kunaka.bsky.social www.biorxiv.org/content/10.6...

Protein event of the year - sign up today! See you at beautiful Palazzo dei Normanni in Palermo for the FEBS 2026 Protein Termini Workshop.
#ProteinTermini #Proteostasis #ProteinModifications #StructuralBiology #PalazzoDeiNormanni #Palermo2026
proteintermini.org/meeting/
Remember to register for this year's Protein Termini FEBS Advanced course. Keynote by Roland Beckmann @beckmannlab.bsky.social "It takes more than two to tango: structural basis and coordination of co-translational N-terminal nascent chain modification" www.conferencecentral.org/webpage/view...
07.02.2026 15:49 — 👍 3 🔁 2 💬 0 📌 0Protein degradation in Turnip - an adapted Arg/N-degron pathway. Preprint from Brian Mooney et al. Emmanuelle Graciet lab. www.biorxiv.org/content/10.6...
30.01.2026 14:09 — 👍 6 🔁 5 💬 0 📌 0
Ribosome-NAC collaboration: A regulatory platform for cotranslational chaperones, enzymes, and targeting factors. New review out in @cp-molcell.bsky.social from Elke Deuerling's lab. www.sciencedirect.com/science/arti...
28.01.2026 11:18 — 👍 6 🔁 3 💬 0 📌 0
Complexity at the ribosomal polypeptide tunnel exit. The major N-terminal acetyltransferase NatA may form different complexes on the ribosome to facilitate protein maturation. By Marius Klein & Klemens Wild in Irmgard Sinning's lab BZH & Nina McTiernan in my lab. www.nature.com/articles/s41...
26.01.2026 11:24 — 👍 16 🔁 8 💬 0 📌 1
This is how co-translational N-terminal myristoylation occurs on the ribosome. NMT1 acts together with NAC, but after the release of MetAP. Cryo-EM study by Timo Denk et al. @beckmannlab.bsky.social @giglionelab.bsky.social www.nature.com/articles/s41...
26.01.2026 11:00 — 👍 17 🔁 6 💬 0 📌 0
🎉 As we celebrate #TIBS50, we're highlighting #CitedClassics and revisiting the top cited article from each of the last 50 years!
Check out this one from 1976! Read it here 👉 lnkd.in/egdN-AXt
1. Apply for one of the positions below (lab or theory) by Jan 18th
2. Learn interdisc skills and discover cool new things about how mitochondria move and socialise
3. Explore some of Norway's beautiful nature (both the below, Rundemanen and Gullfjellet, <10km from work)
Intricate regulation of Src kinases via N-terminal modifications and specific degradation pathways. A novel degradation pathway involving the CUL4A-DDB1-DCAF10 E3 ligase found by Kremer et al. Tanja Bange lab @andrea-musacchio.bsky.social @ispt-proteinterm.bsky.social www.nature.com/articles/s41...
05.01.2026 13:34 — 👍 8 🔁 4 💬 0 📌 0
Last X-Mas, the ribosome gave you methionine,
but the very next day, MetAP took it away.
This year, to save histones from tears,
NatD gives you an acetyl group. ⭐️
Explore our latest paper with the Deuerling lab @uni-konstanz.de and Shu-ou Chan lab @caltech.edu!
www.science.org/doi/10.1126/...

Online Now: HYPK promotes N-terminal protein acetylation through rapid ribosome exchange of NatA Online now:
10.12.2025 16:19 — 👍 9 🔁 4 💬 0 📌 2
📢 Our December issue is now online! Access the full issue 👉 www.cell.com/trends/bioch...
🎨 The cover brings warm, summertime vibes ☀️ and highlights a Feature Review on #AromaticAminoAcid biosynthesis in 🌱 from Jorge El-Azaz and @hiroshi-maeda.bsky.social. Cover art credit to Tae Park.
(1/n)

We’ve made some new tools to manipulate N-recognins. Check them out in our preprint.
www.biorxiv.org/content/10.1...
Just registered for FEBS advanced workshop Protein termini 2026 in Palermo #proteintermini2026 (?) Looking forward to a great meeting. @ispt-proteinterm.bsky.social
www.conferencecentral.org/webpage/view...
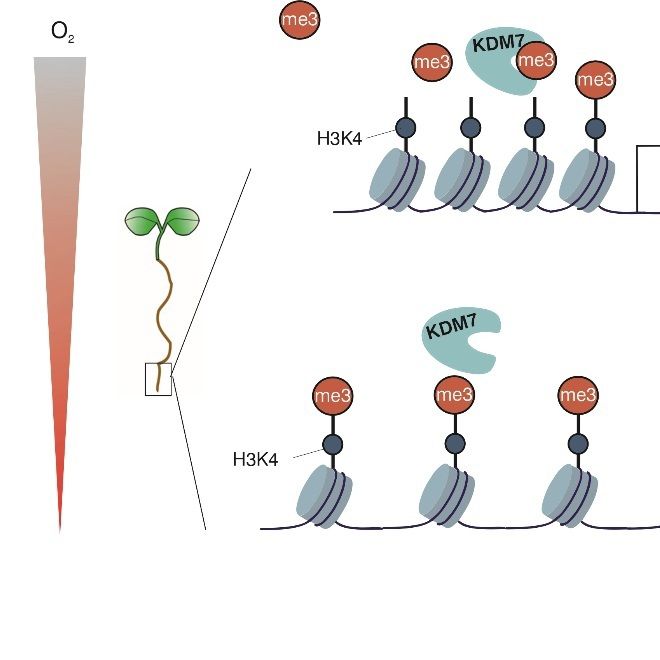
Our study led by the extraordinary multitasking Daai Zhang shows here (tinyurl.com/3uhbkh54) that an additional O2 sensing mechanism based on histone methylation helps roots to prepare for potentially lethal hypoxic stress (such as in waterlogging)
27.11.2025 10:45 — 👍 21 🔁 8 💬 1 📌 2
📝 Online now - the Review "Emerging functions for nonhistone protein acetylation in budding yeast" from Michael Downey and colleagues.
#PostTranslationalModifications #DNADamageResponse #Proteostasis #Autophagy #MetabolicRegulation
Read it here: authors.elsevier.com/a/1m4nM3S6Gf...

Want to assess N-terminal acetyltransferases without radioactive labelling or mass spec? Look here! @ispt-proteinterm.bsky.social @cellsignal.com
27.10.2025 20:25 — 👍 4 🔁 3 💬 1 📌 0
See you at Palazzo dei Normanni in Palermo for the 2026 Protein Termini Conference | Co- and post-translational modifications | Proteolytic processing and degradation pathways | Microbes, plants, animals #ProteinTermini2026 #Proteostasis #ProteinModifications #PalazzoDeiNormanni #Palermo2026
26.10.2025 18:59 — 👍 7 🔁 7 💬 2 📌 1
We are advertising a PhD project investigating a cellular mechanism that ensures protein quality control during mRNA translation in plants. Please get in touch on here or via email if you are interested and I can provide further details!
Lab website: sites.google.com/site/danielg...

Uptake of anti-cancer drugs depends on N-terminally acetylated anion channels | @ispt-proteinterm.bsky.social www.nature.com/articles/s42...
07.10.2025 11:03 — 👍 5 🔁 2 💬 0 📌 0
📢 Our October issue is now online. Access the full issue 👉 www.cell.com/trends/bioch....
🎨 The cover highlights a Review from @chiosislabmskcc.bsky.social and co on how PTMs reprogram chaperones into epichaperomes.