https://doi.org/10.1038/s41589-026-02145-w
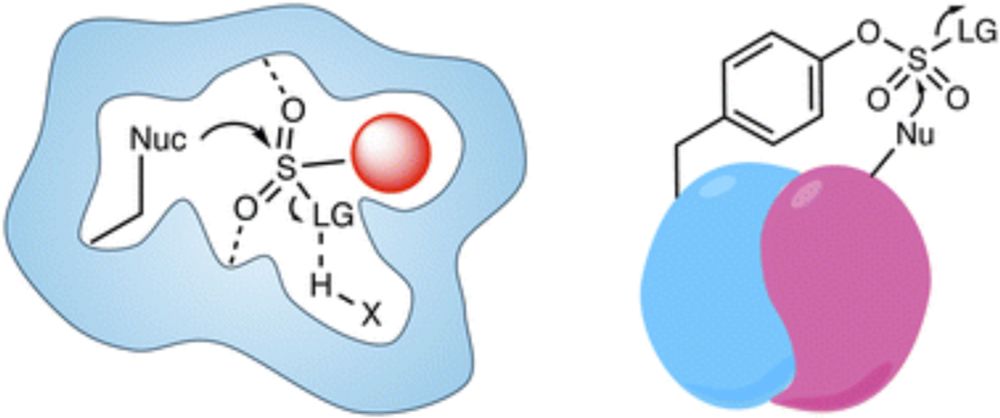
Excited to share our new Perspective in @natchembio.nature.com with @Yang_J_lab.
How do we move from isolated cysteine modifications to a systems-level understanding of redox signaling? We outline a unified chemical framework for decoding the cysteine redoxome.
Read here: t.co/6XY1FCdtN6
02.03.2026 14:24 —
👍 6
🔁 4
💬 0
📌 0

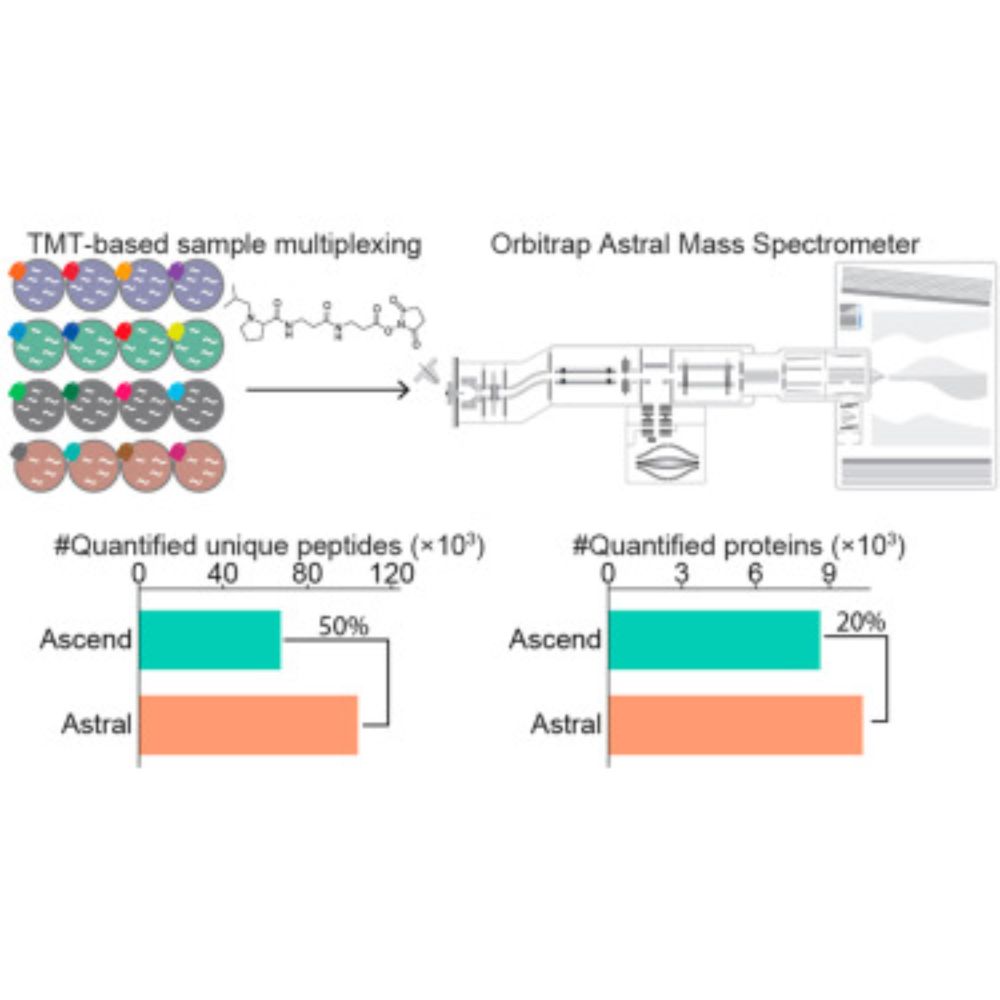
pmultiqc: An open-source, lightweight, and metadata-oriented QC reporting library for MS proteomics www.sciencedirect.co...
---
#proteomics #prot-paper
18.02.2026 20:20 —
👍 3
🔁 1
💬 0
📌 0

📯👨🚀 Calling all #bioimage analysis aficionados. #EZInput is out! We wanted to bring #ImageJ’s capacity to record settings to the #Python ecosystem 🐍. It is a declarative library for generating UIs that just work. Done with @guijacquemet.bsky.social lab
arxiv.org/abs/2601.08859 #ScientificComputing
15.01.2026 11:36 —
👍 44
🔁 14
💬 2
📌 1
ProteoBench: the community-curated platform for comparing proteomics data analysis workflows https://www.biorxiv.org/content/10.64898/2025.12.09.692895v1
11.12.2025 19:46 —
👍 0
🔁 1
💬 0
📌 0
#Chemoproteomics
13.12.2025 02:33 —
👍 0
🔁 0
💬 0
📌 0

AlphaFold Protein Structure Database 2025: a redesigned interface and updated structural coverage
Abstract. The AlphaFold Protein Structure Database (AFDB; https://alphafold.ebi.ac.uk), developed by EMBL–EBI and Google DeepMind, provides open access to
From Sameer Velankar & colleagues in @narjournal.bsky.social #NARDatabaseIssue | #AlphaFold #Protein #Structure #Database 2025: a redesigned interface and updated structural coverage | #Bioinformatics #Proteomics #OpenScience #AFDB 🧪🔓 CC/ @ebi.embl.org
⬇️
academic.oup.com/nar/advance-...
24.11.2025 00:56 —
👍 30
🔁 14
💬 0
📌 0

A framework for AI-ready data
Define AI readiness and its underlying principles through our new research.
Data is the foundation of AI. Poor quality data drives up costs and can lead to hidden problems for AI, while biased data negatively affects the performance of AI models. In our research, we seek to define AI readiness and its underlying principles. Read more.
12.11.2025 11:47 —
👍 5
🔁 4
💬 0
📌 2

Chemical biology 2026
Registration for THE chemical biology conference of 2026 is now open! EMBO ChemBio 2026 in Heidelberg
DeGrado, Arikin, Picotti (Keynotes). @lmkdassama.bsky.social @brianliau.bsky.social @rhodamine110.bsky.social @benlehner.bsky.social @alitavassoli.bsky.social
www.embl.org/about/info/c...
11.11.2025 17:46 —
👍 32
🔁 19
💬 2
📌 0
This project started 5 years ago. It led us to add isotope-labeling support to #FragPipe/#IonQuant. Since then, the tools have grown so much and are now widely used in #Chemoproteomics.
Huge thanks to everyone, and special thanks to @stephanhacker2.bsky.social and @pzanon.bsky.social
30.10.2025 14:15 —
👍 15
🔁 5
💬 0
📌 0

Deep phenotyping of health–disease continuum in the Human Phenotype Project - Nature Medicine
Deep phenotyping of an ancestrally diverse group of 13,000 individuals in the Human Phenotype Project highlights diversity and variations in lifestyle factors, clinical features and molecular…
A new Nature Medicine paper introduces the Human Phenotype Project, a large-scale cohort integrating multi-omics, imaging, and lifestyle data. The project reveals molecular signatures of disease and uses AI to create predictive health models.
🧬💻 #MedSky
15.07.2025 16:33 —
👍 3
🔁 2
💬 0
📌 0
quantms - Quantitative Mass Spectrometry Analysis
A bioinformatics best-practice analysis pipeline for Quantitative Mass Spectrometry (MS)
🚀 We're launching quantms.org — new hub for all #quantms related projects. Explore its components #pmultiqc & #ibaqpy, and check all reanalysis we're sharing with the community: quantms.org/datasets
Meet the Team and Labs behind the project
👉 quantms.org/about
#OpenSource #Proteomics #ASMS2025
02.06.2025 15:39 —
👍 18
🔁 8
💬 0
📌 0

Screenshot of the marimo-quarto documentation website showing a browser window with the URL marimo-team.github.io/quarto-marimo/. The page displays "marimo + quarto" as the main heading, with a left sidebar navigation containing tutorial sections including Intro, Dataflow, UI, Markdown, Plots, Layout, and Fileformat. The main content area features explanatory text about integrating Quarto with marimo, followed by a blue-highlighted interactive example box titled "Hello from marimo!" that demonstrates a reactive Python cell. Below this is visible Python code using mo.md() function, and a "What?" section that begins explaining marimo and the qmd format. The page has a clean, documentation-style layout with syntax highlighting for the code examples.
After disrupting the notebook world, #marimo is expanding to markdown with a new #Quarto extension (WIP)!
marimo-team.github.io/quarto-marimo/
What's marimo? Reactive #jupyter -like notebooks that automatically run cells when code changes. marimo islands is making this tech portable everywhere 🏝️
30.05.2025 02:10 —
👍 5
🔁 4
💬 1
📌 0

The pan-cancer proteome atlas, a mass spectrometry-based landscape for discovering tumor biology, biomarkers, and therapeutic targets www.cell.com/cancer-...
---
#proteomics #prot-paper
29.05.2025 19:40 —
👍 5
🔁 2
💬 0
📌 0
#ChemBio #Chemoproteomics
26.05.2025 22:08 —
👍 3
🔁 0
💬 0
📌 0

ChemBio in the PUB 2025
Chemical Biology in the PUB
Chembio in the Pub! The tastier, chattier version of the popular Boston conference. Jun 12th is pretty close to the GRC too so whether you're a local or visiting it would be great to hang out www.eventbrite.com/e/chembio-in... #chembio #chemicalbiology #bostonbiotech
20.05.2025 15:39 —
👍 0
🔁 2
💬 1
📌 0

The Open Molecules 2025 (OMol25) Dataset, Evaluations, and Models
Machine learning (ML) models hold the promise of transforming atomic simulations by delivering quantum chemical accuracy at a fraction of the computational cost. Realization of this potential would en...
It's finally done (enough for a preprint)! Today, in collaboration with so many folks at Meta (shout-out Daniel Levine and Muhammed Shuaibi, who put in superhuman levels of work), Berkeley, Stanford, NYU, and more, I'm proud to announce the Open Molecules 2025 (OMol25) dataset!
#CompChem #ML 🧪 ⚗️
14.05.2025 16:06 —
👍 39
🔁 10
💬 1
📌 0

Announcement for the latest version of RStudio (2025.05.0) and Posit Workbench, highlighting new features likely related to console output and package management, as illustrated by the console screenshots showing warnings, informative messages, and package installation details
We’re excited to announce the release of Posit Workbench and RStudio 2025.05.0!
The latest release includes Quarto 1.6 support, genAI integrations, better errors/warnings in the console, and other improvements.
Read more in the blog post: posit.co/blog/rstudio...
#RStats #Python
08.05.2025 19:13 —
👍 28
🔁 5
💬 0
📌 5
![A thumbnail with a dark background and the VS Code logo that reads “What’s new in Visual Studio Code April Update [1.100] -Performance improvements in chat -Auto-attach instructions to chat -Reuse prompts for common tasks -Smarter chat responses with GitHub, extensions, and notebook tools -Identify staged changes in editor -Improved multi-window support -Quick create for Python Environments -Image and Streamable HTTP support for MCP servers”](https://cdn.bsky.app/img/feed_thumbnail/plain/did:plc:2xibtqlq4oplg5hkbqw56xgu/bafkreih3cfvsxjemy3ni2symoe77ag7tbbarx77wn4e6uzwzae6eztk73y)
A thumbnail with a dark background and the VS Code logo that reads “What’s new in Visual Studio Code April Update [1.100] -Performance improvements in chat -Auto-attach instructions to chat -Reuse prompts for common tasks -Smarter chat responses with GitHub, extensions, and notebook tools -Identify staged changes in editor -Improved multi-window support -Quick create for Python Environments -Image and Streamable HTTP support for MCP servers”
💯 v1.100 of VS Code is here! And we’ve got some great updates for you, like:
- Smarter chat responses with new tools
- Improved multi-window support
- Image and Streamable HTTP support for MCP servers
…and so much more. aka.ms/VSCodeRelease
Here are some of the highlights… 🧵
08.05.2025 17:17 —
👍 106
🔁 23
💬 3
📌 5

Open-Source and FAIR Research Software for Proteomics
Scientific discovery relies on innovative software as much as experimental methods, especially in proteomics, where computational tools are essential for mass spectrometer setup, data analysis, and interpretation. Since the introduction of SEQUEST, proteomics software has grown into a complex ecosystem of algorithms, predictive models, and workflows, but the field faces challenges, including the increasing complexity of mass spectrometry data, limited reproducibility due to proprietary software, and difficulties integrating with other omics disciplines. Closed-source, platform-specific tools exacerbate these issues by restricting innovation, creating inefficiencies, and imposing hidden costs on the community. Open-source software (OSS), aligned with the FAIR Principles (Findable, Accessible, Interoperable, Reusable), offers a solution by promoting transparency, reproducibility, and community-driven development, which fosters collaboration and continuous improvement. In this manuscript, we explore the role of OSS in computational proteomics, its alignment with FAIR principles, and its potential to address challenges related to licensing, distribution, and standardization. Drawing on lessons from other omics fields, we present a vision for a future where OSS and FAIR principles underpin a transparent, accessible, and innovative proteomics community.
Fantastic review with an unusual history, growing out of a passionate blog post by @willfondrie.com (willfondrie.com/2024/10/the-...), resulting from a storm (in our teacup) on X during @hupo-org.bsky.social 2024. Great teamwork, authors! pubs.acs.org/doi/10.1021/...
24.04.2025 18:24 —
👍 16
🔁 11
💬 0
📌 1
We provide a new checklist for selecting & using chemical probes—valuable for researchers, journal editors, reviewers, and vendors. Ensure best practices in chemical biology! 🧪✅
🔗 Supplementary material: ars.els-cdn.com/content/imag...
#ChemicalProbes #DrugDiscovery #ChemicalBiology
02.03.2025 09:14 —
👍 9
🔁 5
💬 0
📌 0
A discord server to discuss proteomics, metabolomics, and lipidomics.. #TeamMassSpec
22.02.2025 12:41 —
👍 3
🔁 2
💬 0
📌 0

🚀 Thrilled to share our latest research! With the labs of Yansheng Liu and Junmin Peng we’ve built the largest in vivo protein turnover resource across multiple tissues and brain regions, now published in Cell @cellpress.bsky.social 🔬💡 #Proteomics Check it out! 👇
authors.elsevier.com/c/1koBJL7PXu...
20.03.2025 16:35 —
👍 8
🔁 6
💬 0
📌 0















![A thumbnail with a dark background and the VS Code logo that reads “What’s new in Visual Studio Code April Update [1.100] -Performance improvements in chat -Auto-attach instructions to chat -Reuse prompts for common tasks -Smarter chat responses with GitHub, extensions, and notebook tools -Identify staged changes in editor -Improved multi-window support -Quick create for Python Environments -Image and Streamable HTTP support for MCP servers”](https://cdn.bsky.app/img/feed_thumbnail/plain/did:plc:2xibtqlq4oplg5hkbqw56xgu/bafkreih3cfvsxjemy3ni2symoe77ag7tbbarx77wn4e6uzwzae6eztk73y)