
New #MicrobiomeDigest is OUT microbiomedigest.com/2025/10/02/o...
• Shanghai dog microbiome / @annacusco.bsky.social
• microbetag / @hariszaf.bsky.social
• invitation for the #MVIF 42 / @microbiomevif.bsky.social
& more.
@annacusco.bsky.social
Microbiome scientist | Metagenomics | Long-read sequencing Postdoc at Big Data Biology Lab

New #MicrobiomeDigest is OUT microbiomedigest.com/2025/10/02/o...
• Shanghai dog microbiome / @annacusco.bsky.social
• microbetag / @hariszaf.bsky.social
• invitation for the #MVIF 42 / @microbiomevif.bsky.social
& more.

At Handan campus, Fudan University, Shanghai.

Exploring the Great Wall near Beijing
If you made it this far, thanks for reading. I hope you enjoyed it.
I am currently looking for my next research adventure. If you have insights on the microbiome/microbial genomics job market in Europe (academia & industry), or want to chat about this work, please reach out.
That sounds great, thanks for sharing!
25.09.2025 10:05 — 👍 1 🔁 0 💬 0 📌 0
September 2025 updates!
A focus on Anna's preprint, but several other updates too, including Faith Adegoke joining us to work on AMR and several other preprints
bigdatabiology.substack.com/p/bdb-lab-se...
Thanks Steven! :)
24.09.2025 08:09 — 👍 0 🔁 0 💬 0 📌 0
At Handan campus, Fudan University, Shanghai.

Exploring the Great Wall near Beijing
If you made it this far, thanks for reading. I hope you enjoyed it.
I am currently looking for my next research adventure. If you have insights on the microbiome/microbial genomics job market in Europe (academia & industry), or want to chat about this work, please reach out.

The main characters of this story: the Shanghai pet dogs
I would like to thank all co-authors: Yiqian Duan, Fernando Gil, Alexei Chklovski, Nithya Kruthi, Shaojun Pan, Sofia Forslund, Susanne Lau, Ulrike Löber, Xing-Ming Zhao, and especially @luispedrocoelho.bsky.social
And of course, all the dog owners and the main characters of this story🐶

Screenshot of https://sh-dog-mags.big-data-biology.org/
All the generated data and resources are publicly available.
You can play around with the Shanghai dog MAG catalog here: sh-dog-mags.big-data-biology.org
where we added MAG basic information, ARGs annotation, and linked the 16S rRNA genes to MicrobeAtlas.
More data at Zenodo & ENA.

PCoA plot representing beta diversity (Bray-Curtis on log-transformed data).
Using global dog gut microbiome data, we found that the living environment (household, colony, free-roaming) was the strongest factor shaping the gut microbiome composition.
(This is consistent with the mapping results: higher mapping rates for pet dog vs. non-pet dog cohorts)

Comparison of the representative species-level genome assemblies: canine long-read MAGs vs. public database representative (RefSeq or GenBank).
In general, our species-level MAG representatives were more contiguous and had a higher quality than the reference genome assembly in public databases —especially regarding the presence of rRNA and mobile genetic elements, which are often missed in short-read assemblies.
23.09.2025 11:23 — 👍 2 🔁 0 💬 1 📌 0
The Shanghai dog MAG catalog captures the majority of the microbial diversity of other pet dog cohorts living in households (median read mapping of >90%).
Our Shanghai dog catalogs proved to be globally representative🌍: over 90% of reads (median value) from pet dog cohorts in Germany, South Africa, and the USA mapped to them.
23.09.2025 11:23 — 👍 2 🔁 0 💬 1 📌 0

Total number of MAGs per sample, stratified by quality. Almost all the high-quality MAGs fulfilled the MIMAG criteria.
Let's go to some of the main messages:
🧬We recovered 2,676 MAGs from Shanghai dogs, ~72% were near-finished (high-quality regarding MIMAG criteria) & highly contiguous.
⭕We recovered 185 circular extrachromosomal elements like plasmids and viruses from the same dogs.
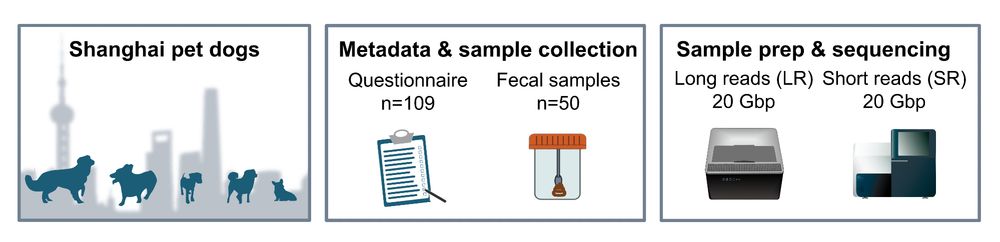

Each stool sample was deeply sequenced 🐶💩: 20 Gbp Illumina short-reads + (at least) 20 Gbp Nanopore long-reads per dog. That's a substantial throughput for assembling MAGs from complex metagenomes🧬!
23.09.2025 11:23 — 👍 3 🔁 1 💬 1 📌 0
Sampling kit material for the Shanghai dog gut microbiome project
We started by sharing questionnaires via WeChat 📲 and then collected 💩 samples from 51 pet dogs across Shanghai 🐶. Along the way, we met many doggies (& their humans)!
23.09.2025 11:23 — 👍 4 🔁 1 💬 1 📌 0
Finally, the results of my postdoc at the
@bigdatabiology.bsky.social lab in Shanghai see the light! The work includes my three favorite things research-wise: 🦠 microbiome, 🧬 long-reads, and 🐶 dogs.
See our new preprint:
www.biorxiv.org/content/10.1...

At Handan Campus, Fudan University, Shanghai.

Exploring the Great Wall near Beijing
If you made it this far, thanks for reading. I hope you enjoyed the thread.
I am currently looking for my next research adventure. If you have insights on the microbiome/microbial genomics🧬🦠 job market in Europe (academia and industry), or want to chat about this work, please reach out!

I would like to thank all co-authors: Yiqian Duan, Fernando Gil, Alexei Chklovski, Nithya Kruthi, Shaojun Pan, Sofia Forslund, Susanne Lau, Ulrike Löber, Xing-Ming Zhao, and especially @luispedrocoelho.bsky.social
And of course, all the dog owners and the main characters of this story 🐶

Screenshot of https://sh-dog-mags.big-data-biology.org/
All the generated data and resources are publicly available.
You can play around with the Shanghai dog MAG catalog here: sh-dog-mags.big-data-biology.org
where we have added MAG basic information, ARGs annotation, and linked 16S rRNA genes to MicrobeAtlas.
More data at Zenodo and ENA.
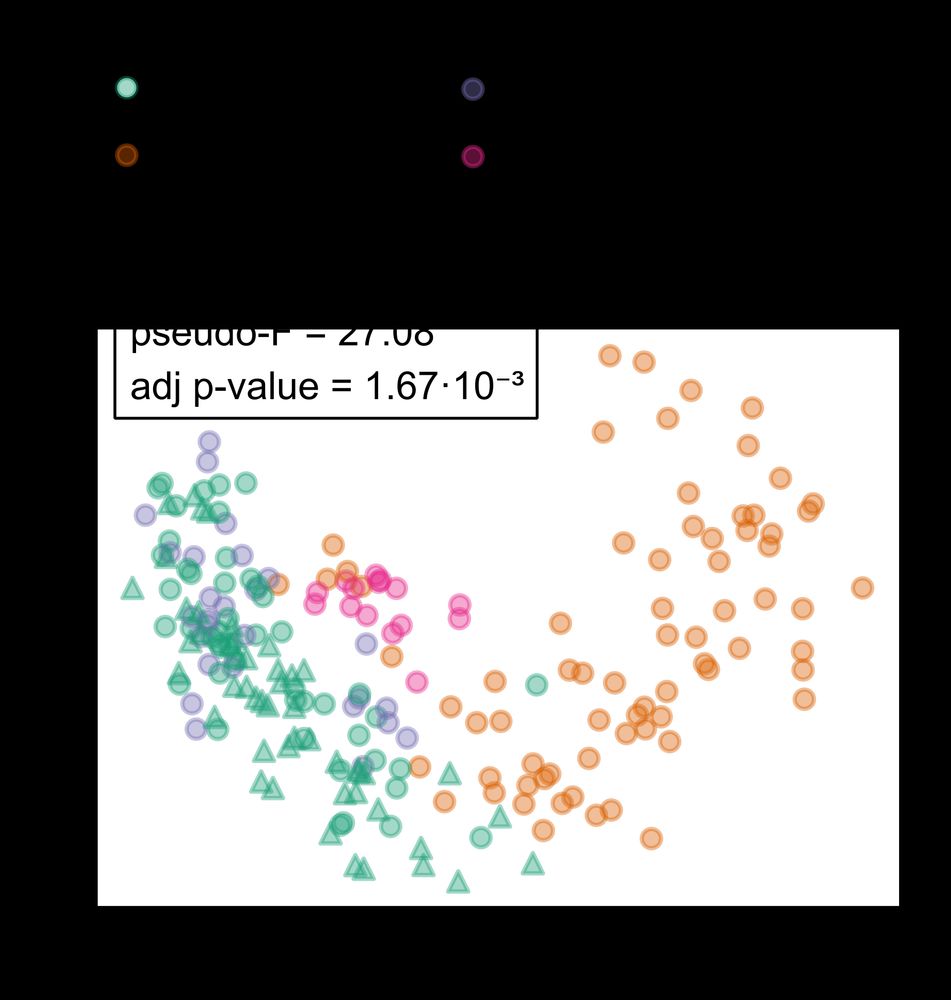
PCoA plot representing beta diversity (Bray-Curtis on log-transformed data). Green triangles indicate pet dogs in this study.
Using global dog gut microbiome data, we found that the living environment (household, colony, free-roaming) was the strongest factor shaping the gut microbiome composition.
(This is consistent with previous mapping results to our catalog: higher mapping rates for pet dog vs. non-pet dog cohorts)

In general, our species-level MAG representatives were more contiguous and had a higher quality than the reference genome assembly in public databases —especially regarding the presence of rRNA and mobile genetic elements, which are often missed in short-read assemblies.
23.09.2025 10:41 — 👍 0 🔁 0 💬 1 📌 0
The Shanghai dog catalogs capture the majority of the microbial diversity of other pet dog cohorts living in households (median read mapping of >90%). The mapping is lower for non-pet cohorts (colony, shelter, or free-roaming dogs).
Our Shanghai dog catalogs proved to be globally representative🌍: over 90% of reads (median value) from pet dog cohorts in Germany, South Africa, and the USA mapped to them.
23.09.2025 10:41 — 👍 0 🔁 0 💬 1 📌 0
Shanghai dog catalogs: metagenome-assembled genomes & extrachromosomal elements

Shanghai dog MAG catalog: total number of MAGs per sample, stratified by quality. Almost all the high-quality MAGs fulfilled the MIMAG criteria.
Let's go to some of the main messages:
🧬We recovered 2,676 MAGs from Shanghai dogs, ~72% were near-finished (high-quality regarding MIMAG criteria) and highly contiguous.
⭕Additionally, we retrieved 185 circular extrachromosomal elements like plasmids and viruses from the same dogs.

Project overview
Each stool sample was deeply sequenced 🐶💩: 20 Gbp Illumina short-reads + (at least) 20 Gbp Nanopore long-reads per dog. That's a substantial throughput for assembling MAGs from complex metagenomes🧬!
23.09.2025 10:41 — 👍 0 🔁 0 💬 1 📌 0
Sampling material
We started by sharing questionnaires via WeChat 📲 and then collected 💩 samples from 51 pet dogs across Shanghai 🐶. Along the way, we met many doggies (& their humans)!
23.09.2025 10:41 — 👍 1 🔁 0 💬 1 📌 0
Full thread will come later, but @annacusco.bsky.social's preprint on the dog pet gut microbiome is out!
Using ONT+Illumina, we get better MAGs than to corresponding species representative in public databases
doi.org/10.1101/2025...
Capturing global pet dog gut microbial diversity and hundreds of near-finished bacterial genomes by using long-read metagenomics in a Shanghai cohort https://www.biorxiv.org/content/10.1101/2025.09.17.676595v1
18.09.2025 06:17 — 👍 6 🔁 5 💬 0 📌 0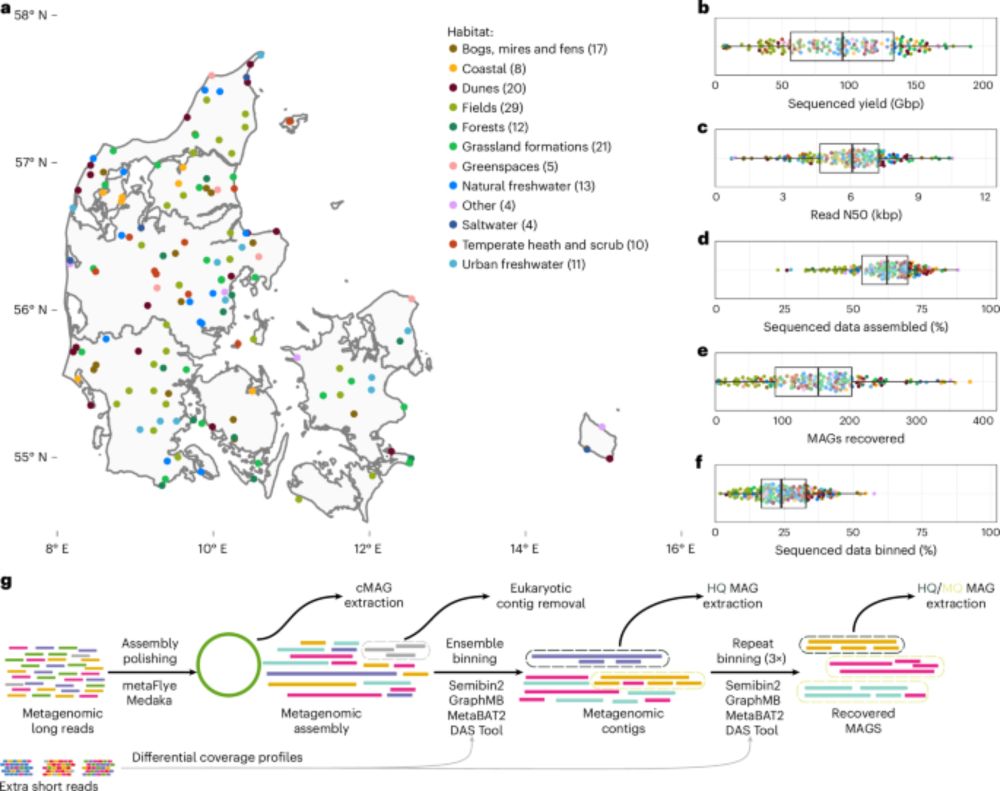
Genome-resolved long-read sequencing expands known microbial diversity across terrestrial habitats www.nature.com/articles/s41... #jcampubs
24.07.2025 12:55 — 👍 36 🔁 13 💬 0 📌 1
Logo for the Sandpiper website
Out in @natbiotech.nature.com: Metagenome taxonomy profilers usually ignore unknown species. SingleM is an accurate profiler which doesn't, even detecting phyla with no MAGs. Profiles of 700,000 metagenomes at sandpiper.qut.edu.au. A 🧵
16.07.2025 21:59 — 👍 131 🔁 71 💬 7 📌 9First code release of "SingleM for dsDNA phage"! Lyrebird scans metagenomic reads for marker genes to give a “phage community profile”. It detects many novel phages, many more than standard contig-centric methods. @benjwoodcroft.bsky.social @emerge-bii.bsky.social wwood.github.io/singlem/Lyrebird
16.05.2025 10:55 — 👍 12 🔁 8 💬 0 📌 1argNorm is published now academic.oup.com/bioinformati...
17.04.2025 04:57 — 👍 27 🔁 14 💬 2 📌 1