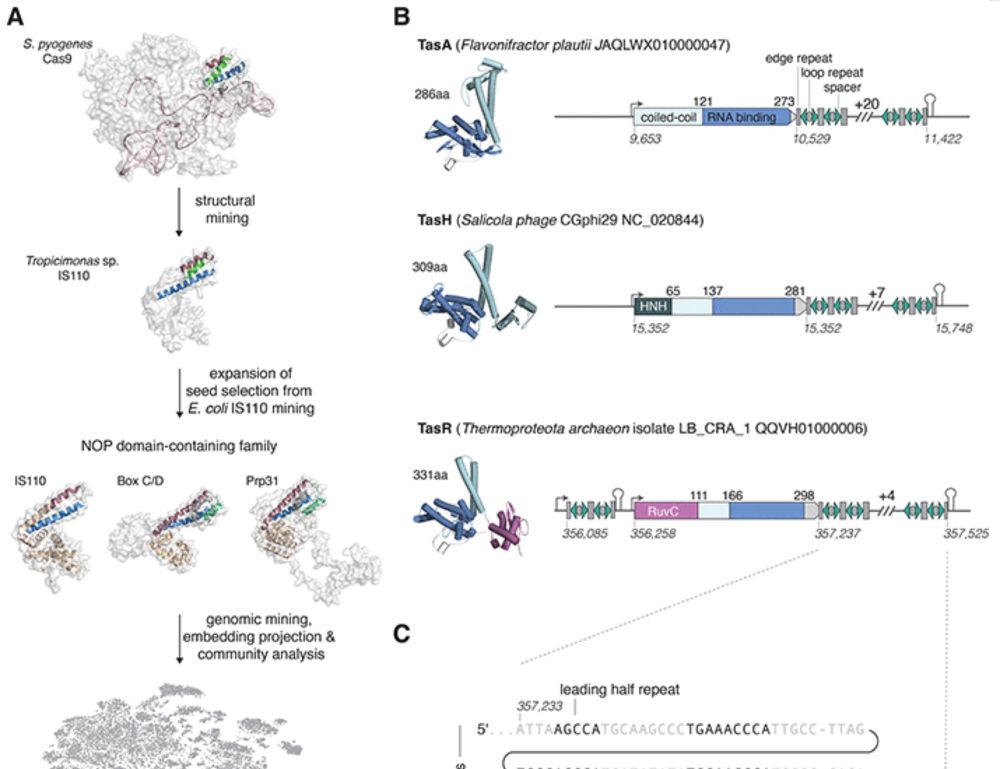
Top: A representative modeling example from Chimpanzee CPEB3 HDV-like ribozyme (PDB ID: 7QR3), with models predicted by four better-performing methods (blue cartoons) overlaid on experimental structure (gray cartoons). Left to right: DRFold2, DeepFoldRNA, AlphaFold3, RhoFold. Bottom: Structural visualization of the example from coxsackievirus B3 cloverleaf RNA (PDBID: 8DP3), showing experimental structure (left), AlphaFold3’s best prediction from 100 models (middle), and 5th model of DRfold2 (right), respectively. Structures are rainbow-colored from 5′ (blue) to 3′ (red) end.
Accurate RNA structure prediction remains a challenge, despite recent computational advances. This study presents DRFold2, a #DeepLearning framework that significantly enhances accuracy of de novo #RNAstructure prediction by increasing contact prediction precision @plosbiology.org 🧪 plos.io/4aoOQiX
18.02.2026 09:37 —
👍 23
🔁 8
💬 0
📌 0

The train station in Qingdao. A light-flooded palace for high-speed trains.

Inside the high-speed train Qingdao-Beijing in China. Riding at more than 300 km/h.
Riding a train in China, returning from the ISPP 2025 and the Green Carbon conference.
26.09.2025 16:03 —
👍 6
🔁 0
💬 0
📌 0
Welcome
To correctly determine a distance there, we used Jensen Shannon Divergence as a metric, which is a symmetrified version of the well-known Kullback–Leibler divergence for comparing distributions.
The GradR data can be searched online at synecho-rapdor.biologie.uni-freiburg.de
@meinbio.bsky.social
26.09.2025 15:51 —
👍 1
🔁 0
💬 0
📌 0
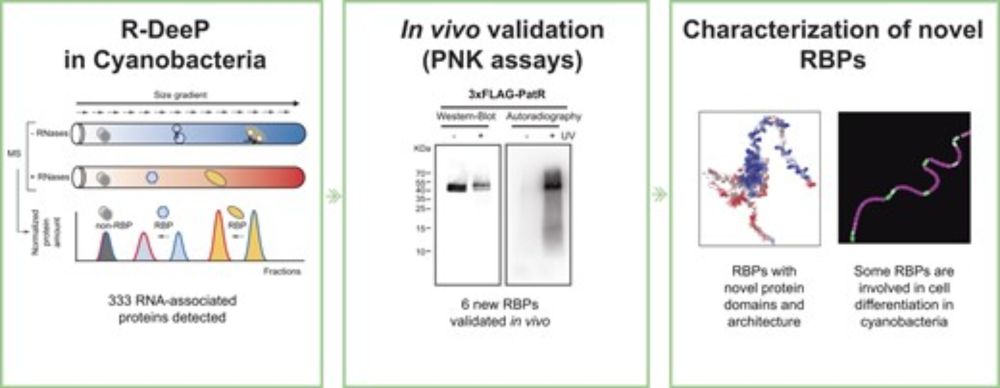
For the intuitive analysis + visualization of such datasets we introduce RAPDOR. In contrast to expression profiles, which are compared e.g. by correlation, the results of GradR and similar approaches are naturally displayed as distributions, as the protein fraction in each compartment is measured.
26.09.2025 15:51 —
👍 1
🔁 0
💬 1
📌 0
We have been searching for novel RNA-binding proteins and validated, among many candidates, Sll1967 (a possible RlmD homolog), Sll0726 (phosphoglucomutase), Ssl2245 (a putative antitoxin), Slr0711 (possible QueF homolog), and Sll0947 (ribosome-associated inhibitor RaiA/LrtA homolog) as binding RNA.
26.09.2025 15:51 —
👍 1
🔁 0
💬 1
📌 0


Figure on track 1.

The train is made to look nice. Forest items everywhere
Riding a train in Lithuania.
14.08.2025 08:00 —
👍 4
🔁 0
💬 0
📌 0
¡Muchas gracias, José!
26.06.2025 16:29 —
👍 1
🔁 0
💬 0
📌 0
Many thanks, Jörg! And it has been a pleasure to visit Göttingen recently!
26.06.2025 16:28 —
👍 1
🔁 0
💬 0
📌 0
RNA-binding proteins and photosynthesis: The RRM domain–containing protein Rbp3 interacts with ribosomes and the 3’ ends of mRNAs encoding photosynthesis proteins | PNAS www.pnas.org/doi/10.1073/...
26.06.2025 13:59 —
👍 6
🔁 3
💬 2
📌 0
They must cope with low CO2 solubility in water, which can limit the CO2 fixation by RubisCO. Moreover, the total pool of inorganic carbon is highly variable in this environment, depending on pH, temperature, and salinity.
23.06.2025 08:04 —
👍 2
🔁 0
💬 1
📌 0
How are fully submerged algae efficiently fixing carbon?
All organisms performing oxygenic photosynthesis fix CO2 in
the Calvin–Benson–Bassham cycle using RubisCO. But aquatic organisms have a challenge.
23.06.2025 08:04 —
👍 2
🔁 0
💬 1
📌 0
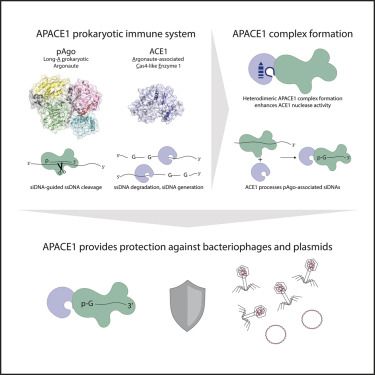
Out now: Cyanobacterial Argonautes and Cas4 family nucleases cooperate to interfere with invading DNA
cell.com/molecular-ce...
Most long-A pAgos interfere with invading DNA solo. Why then are cyanobacterial pAgos co-encoded with a Cas4-like protein?
13.05.2025 07:41 —
👍 32
🔁 12
💬 1
📌 1
15th Workshop on Cyanobacteria. Online registration by Cvent
Just three days left for the deadline for oral abstract submissions for the 15th Workshop on Cyanobacteria at Vanderbilt University (Nashville, TN, USA) - don't miss out!
Workshop dates: 4-7th June 2025. More information: web.cvent.com/event/3d0bd3.... Please share widely!
18.03.2025 18:00 —
👍 8
🔁 10
💬 1
📌 0
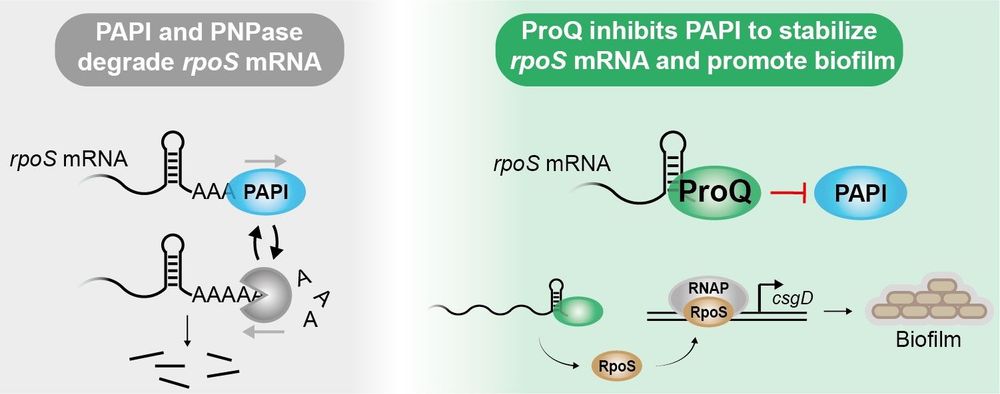
🚨 New #Breakthrough article from @holmqvist-lab.bsky.social: "ProQ prevents mRNA degradation through inhibition of poly(A) polymerase" #RNA #Stability #Degredation #mRNA 📖 Read the full article here: https://doi.org/10.1093/nar/gkaf103
27.02.2025 10:00 —
👍 24
🔁 7
💬 0
📌 1
Annegret Wilde (@freecyano.bsky.social)
cyanobacteriologist from University Freiburg
Our new Opinion article, now out in Trends in Plant Science is the result of a great collaboration with Conrad Mullineaux and Annegret Wilde freecyano.bsky.social @cyanolab.bsky.social
06.02.2025 10:35 —
👍 5
🔁 1
💬 0
📌 0
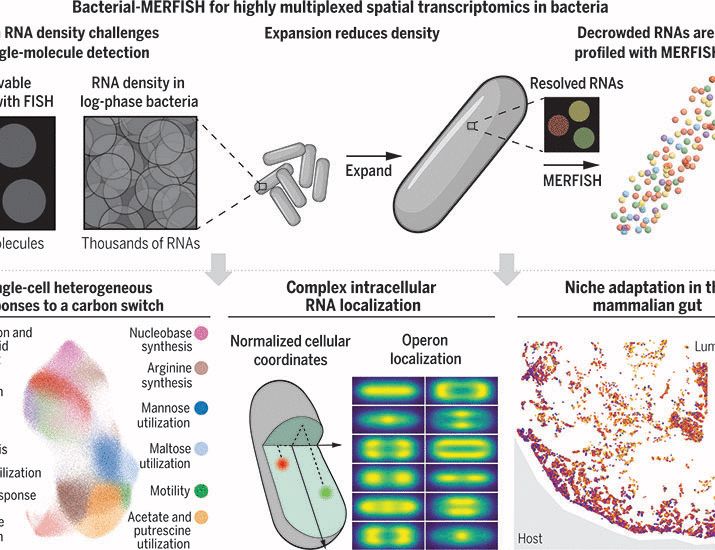
Highly multiplexed spatial transcriptomics in bacteria
Single-cell decisions made in complex environments underlie many bacterial phenomena. Image-based transcriptomics approaches offer an avenue to study such behaviors, yet these approaches have been hin...
🅲🅾🅾🅻 🆃🅴🅲🅷🅽🅾🅻🅾🅶🆈
“Bacterial-MERFISH” provides ~1000-fold volumetric expansion of individual cells, charts gene expression in hundreds of thousands of cells, deciphering bacterial single-cell heterogeneity, intracellular transcriptome organization, and bacterial adaptation to µm-scale niches in vivo
25.01.2025 17:33 —
👍 98
🔁 31
💬 1
📌 3
This review has been put together by 12 authors working on multicellularity in bacteria. Many thanks to the DFG by supporting this research through the SPP priority program 2389 “Emergent Functions of Bacterial Multicellularity”.
26.01.2025 13:46 —
👍 0
🔁 0
💬 0
📌 0

Sharepic showing Date, Time and Place of Prof. Eilon Shani's CIBSS Seminar talk on "A Multi-targeted genome-scale CRISPR Toolbox to Overcome Functional Redundancy and Reveal Hidden Hormone Transport Mechanisms", Tuesday, January 14th 2025, Lecture Hall, Institute of Biology I, Hauptstr. 1, and online per Zoom.
Mark your calendars for next Tuesday: Tel Aviv University's Prof. Eilon Shani will present a #CRISPR -based toolkit that targets multiple genes at a genome-wide scale, addressing functional overlap in genetic systems and to uncover hidden mechanisms behind hormone transport.
kurzlinks.de/fprn
10.01.2025 10:18 —
👍 11
🔁 6
💬 0
📌 1
Not all bacteria, even of a single species, are created equal. Nor are they always unicellular. Some cyanobacteria develop heterocysts and fix nitrogen. GradR/R-DeeP analysis identified new RNA-binding proteins. Some were previously known for other functions. Others were completely uncharacterized.
19.12.2024 17:03 —
👍 5
🔁 0
💬 1
📌 0