
Schematic of cell differentiation prediction (a) and drug response prediction (b).
New paper out in @natmethods.nature.com from @elhamazizi.bsky.social, Kam Leong & @jameszou.bsky.social! The team developed Squidiff, a diffusion #AI model to predict cellular responses to environmental cues and accelerate #PrecisionMedicine.
Learn more: bit.ly/3WRPNsx
10.11.2025 13:59 —
👍 8
🔁 5
💬 1
📌 0
YouTube video by Computational Spatial Biology seminar
Learning single-cell spatial context through integrated spatial multiomics with CORAL - Siyu He
Missed last week’s #CompSpatialBio seminar @justjhong.bsky.social with @siyuhe.bsky.social on CORAL—a deep graph model for spatial-omics? The recording is now available: youtu.be/BV5soaUVUNA
📅 Stay tuned for the next session! csbseminar.github.io
13.05.2025 14:54 —
👍 3
🔁 1
💬 0
📌 0
Thanks @justjhong.bsky.social for the invitation! Looking forward to the discussion.
24.04.2025 17:07 —
👍 2
🔁 0
💬 0
📌 0
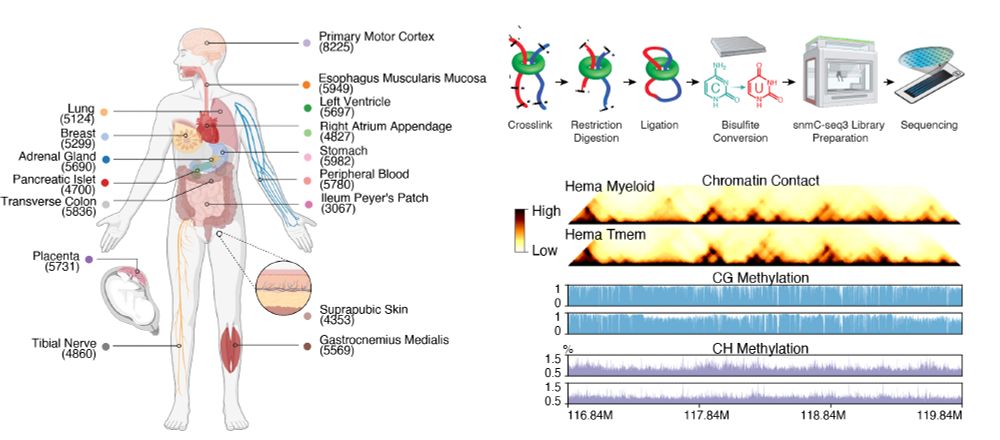
1/10 Excited to share our latest - the first whole-body map of both DNA methylation and 3D genome at single-cell resolution.
25.03.2025 15:49 —
👍 8
🔁 5
💬 1
📌 1

Check out our new collaborative work with Dr. Kam Leong @columbiauniversity.bsky.social now out in Advanced Functional Materials! We developed oral nanoparticles to help mitigate hematopoietic acute radiation syndrome, especially in cases of deep space exploration:
t.co/1jQwcU1pDB
24.02.2025 17:40 —
👍 1
🔁 2
💬 1
📌 0
Applying #CORAL on paired CODEX and Visium data from HCC, we delineated tissue ecosystems and identified macrophages interacting with CD4+ T cells as key players hindering responses.
For details, check our preprint & GitHub!😀
#Immunotherapy #HCC #SpatialOmics #CancerResearch
08.02.2025 16:00 —
👍 1
🔁 0
💬 0
📌 0

#CORAL enables the prediction of cell-cell interactions, as demonstrated in the mouse thymus. The latent variables in the model capture spatial features that transition from the medulla to the cortex, associating with thymic development and T cell maturation. 🧬5/🪸
08.02.2025 16:00 —
👍 0
🔁 0
💬 1
📌 0

Given paired spatial proteomics and downsampled ST data (such as Visium), #CORAL performs comparably to #SpatialGlue in detecting functional domains, while also achieving high-res ST🔬4/🪸
08.02.2025 16:00 —
👍 0
🔁 0
💬 1
📌 0
By leveraging deep generative model and graph attention mechanisms, #CORAL offers key benefits:
🔹 Spatial domain detection
🔹 Single-cell modality imputation
🔹 Inferring cell-cell interactions
Unlocking new possibilities in analyzing #spatialmultiomics integration! 3/🪸
08.02.2025 16:00 —
👍 0
🔁 0
💬 1
📌 0
Big thanks to our collaborators Matthew Bieniosek,
SongDongyuan,zhou_jingtian, benchidester, ZhenqinWu, Joseph Boen, Padmanee Sharma! And a special thanks to my advisors @jameszou.bsky.social and Alex Trevino for their invaluable guidance and support. 2/🪸
08.02.2025 15:59 —
👍 0
🔁 0
💬 1
📌 0
Thrilled to introduce #CORAL🪸—a deep probabilistic graphical model that integrates spatial multiomics data!
🔹Resolves spatial resolution discrepancies across modalities
🔹Deconvolutes low-res data to achieve single-cell granularity
🔹Enables parallel integrative analysis 1/🪸
08.02.2025 15:58 —
👍 14
🔁 4
💬 1
📌 0
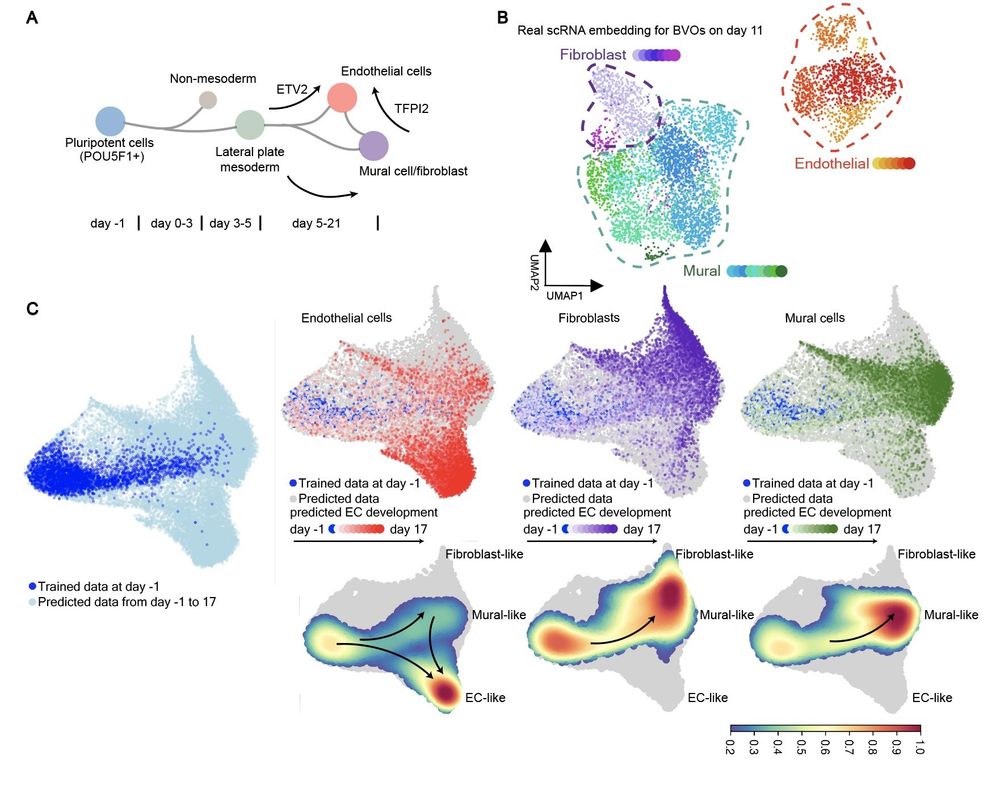
We applied Squidiff to model blood vessel #organoids using single-cell RNA sequencing to examine their response to neutron irradiation and pro-regenerative G-CSF treatment, simulating cosmic irradiation anticipated in #deepspace missions🧑🚀. 5/🦑
09.12.2024 14:13 —
👍 2
🔁 2
💬 1
📌 0
Squidiff effectively predicts non-additive gene perturbation and cell type-specific drug responses, as demonstrated in its application to the analysis of temporal cell states in the glioblastoma dataset in response to novel drug combinations. 4/🦑
09.12.2024 14:13 —
👍 2
🔁 0
💬 1
📌 0

Given the scRNA data on day 0 and day 3, can we predict the data on day 1, day 2 and further? Squidiff predicts this differentiation of induced pluripotent stem cells (iPSCs) into mesendoderm and endoderm, guided by stimuli vectors. 3/🦑
09.12.2024 14:13 —
👍 4
🔁 3
💬 1
📌 0
Thank you so much for the long-term collaboration: Yuefei, @naveedtavakol.bsky.social , Haotian, and Sima. A special thanks to my advisors, James Zou, @elhamazizi.bsky.social , and Kam W. Leong, whose guidance and support have been invaluable. 2/🦑
09.12.2024 14:13 —
👍 3
🔁 0
💬 1
📌 0








