
A new gateway to global antimicrobial resistance data
New online portal connects bacterial genomes with experimental resistance data to support antimicrobial resistance research.
Antimicrobial resistance (AMR) is a growing health threat, making infections harder to treat and complicating routine medical care.
EMBL-EBI’s new AMR portal brings together laboratory resistance data and bacterial genomes in one open platform.
#WAAW2025 #ActOnAMR
www.ebi.ac.uk/about/news/t...
🧬💻
18.11.2025 09:59 — 👍 35 🔁 18 💬 1 📌 2
Huge congratulations and thanks to
@sunjaelee.bsky.social, @milot.bsky.social,
@cameron.gilchrist, and @martinsteinegger.bsky.social 👍
16.10.2025 07:29 — 👍 2 🔁 0 💬 0 📌 0

Curate your database. 🧵4/5
- GTDB, NCBI, ICTV, or custom taxonomy supported.
- Add genomes to a pre-built DB to save time (benchmark in figure)
- Expand taxonomy. e.g., integrate ICTV viruses into a GTDB prokaryote DB.
⏱️Building a DB of 8,520 GTDB species took 106 min on a MacBook M2 Pro (32G)
16.10.2025 07:29 — 👍 0 🔁 0 💬 1 📌 0
Explore results with interactive visualization. 🧵3/5
- Generate customized Sankey plots.
- Search for taxa of interest.
- Filter by classified reads or proportion.
- Click taxon nodes for subtree views.
- Extract reads classified to a taxon.
- Access NCBI Taxonomy and genome browser.
16.10.2025 07:29 — 👍 0 🔁 0 💬 1 📌 0

The app runs desktop-optimized Metabuli. 🧵2/5
Classifying 2X22M human gut reads vs. 36K genomes over 8,465 species took:
🖥️46 min on a Windows desktop (i9-9900, 32GB RAM)
💻39 min on a MacBook M2 Pro (32GB RAM)
16.10.2025 07:29 — 👍 0 🔁 0 💬 1 📌 0
Easy and interactive taxonomic profiling with Metabuli App.
It integrates database curation, read QC, taxonomic profiling, and visualization right on your desktop.
No command line, server, or internet required.
Now published in Bioinformatics! 🧵1/5
doi.org/10.1093/bioi...
github.com/steineggerla...
16.10.2025 07:29 — 👍 24 🔁 11 💬 1 📌 1

Illustration of Burrows-Wheeler Transform and many auxiliary structures from the input string how$now$brown$cow$#
New tool "bwt-svg" for making illustrations of the BWT and the many auxiliary arrays and other structures related to it. Pyodide-based no-installation-necessary interface here: benlangmead.github.io/bwt-svg/. (H/t to @robert.bio for pointing me to pyodide!) Full repo: github.com/benlangmead/....
14.10.2025 20:48 — 👍 40 🔁 21 💬 4 📌 1
Finally we got an end-to-end structural annotation tool for phages!
08.08.2025 08:26 — 👍 11 🔁 1 💬 0 📌 1

The Em Dash Responds to the AI Allegations
“In recent months, a curious fixation has emerged in corners of academia: the em dash. More specifically, the apparent moral panic around how it is...
"Writers have been using me long before the advent of AI. I am the punctuation equivalent of a cardigan—beloved by MFA grads, used by editors when it’s actually cold, and worn year-round by screenwriters. I am not new here."
17.07.2025 18:20 — 👍 417 🔁 171 💬 10 📌 26
Folddisco finds similar (dis)continuous 3D motifs in large protein structure databases. Its efficient index enables fast uncharacterized active site annotation, protein conformational state analysis and PPI interface comparison. 1/9🧶🧬
📄 www.biorxiv.org/content/10.1...
🌐 search.foldseek.com/folddisco
07.07.2025 08:21 — 👍 152 🔁 70 💬 8 📌 3
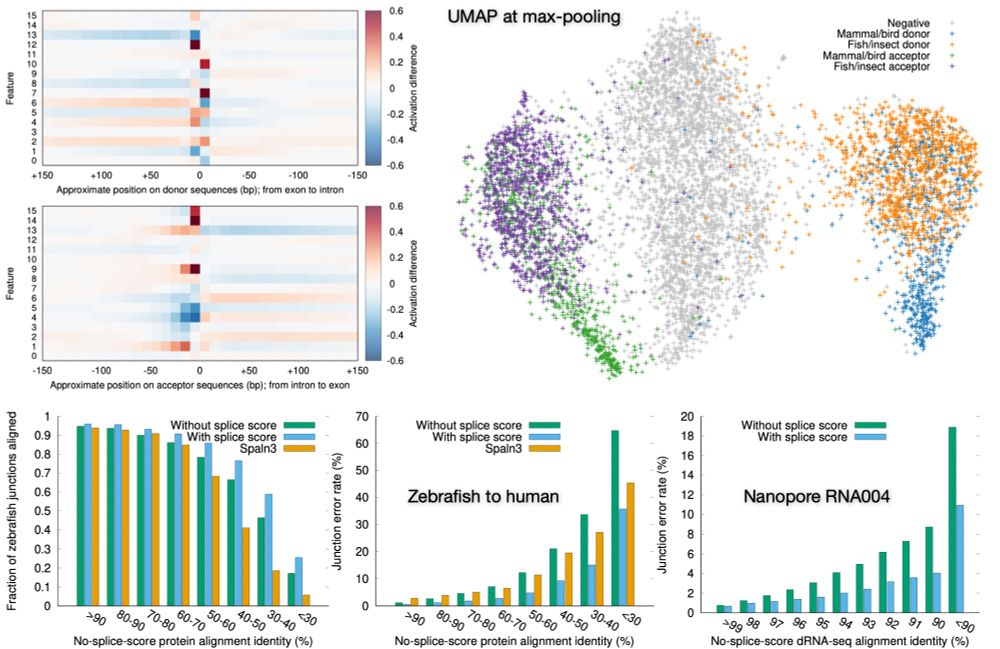
Preprint on "Improving spliced alignment by modeling splice sites with deep learning". It describes minisplice for modeling splice signals. Minimap2 and miniprot now optionally use the predicted scores to improve spliced alignment.
arxiv.org/abs/2506.12986
17.06.2025 01:48 — 👍 112 🔁 54 💬 0 📌 2
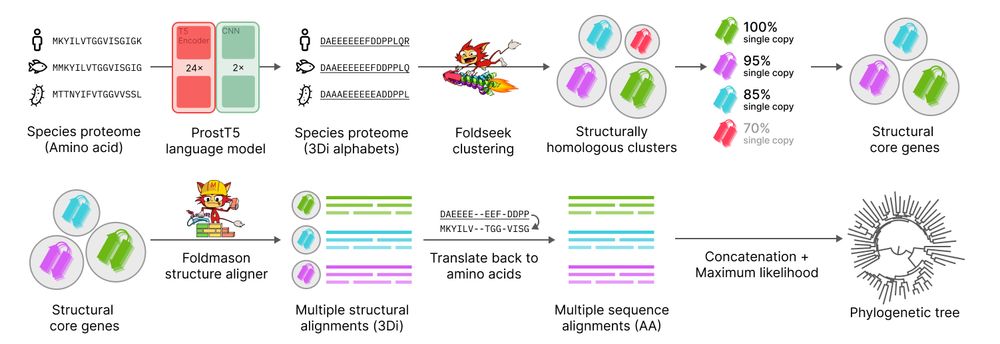
Unicore is now published on GBE 🚀
Unicore rapidly identifies structural single-copy core genes from input species proteomes for phylogenetic analysis. Powered by Foldseek and ProstT5, Unicore enables linear-scale structure-based phylogeny of any given set of taxa. 🧵1/n
📃 doi.org/10.1093/gbe/evaf109
03.06.2025 06:54 — 👍 68 🔁 31 💬 3 📌 2

Introducing our invited speaker for the session on 'Viral Dark Matter' we have Rachel Seongeun Kim from the Seoul National University!!!!
The registrations for on-site & remote participation are still open! More info: RdRp.io
#RdRpSummit2025
02.05.2025 14:55 — 👍 26 🔁 12 💬 1 📌 0
I'm presenting a poster about Metabuli, a metagenomic taxonomic classifier leveraging both DNA and protein sequences, at #RECOMB2025! Please come and share yout thoughts!
25.04.2025 07:52 — 👍 4 🔁 0 💬 0 📌 0
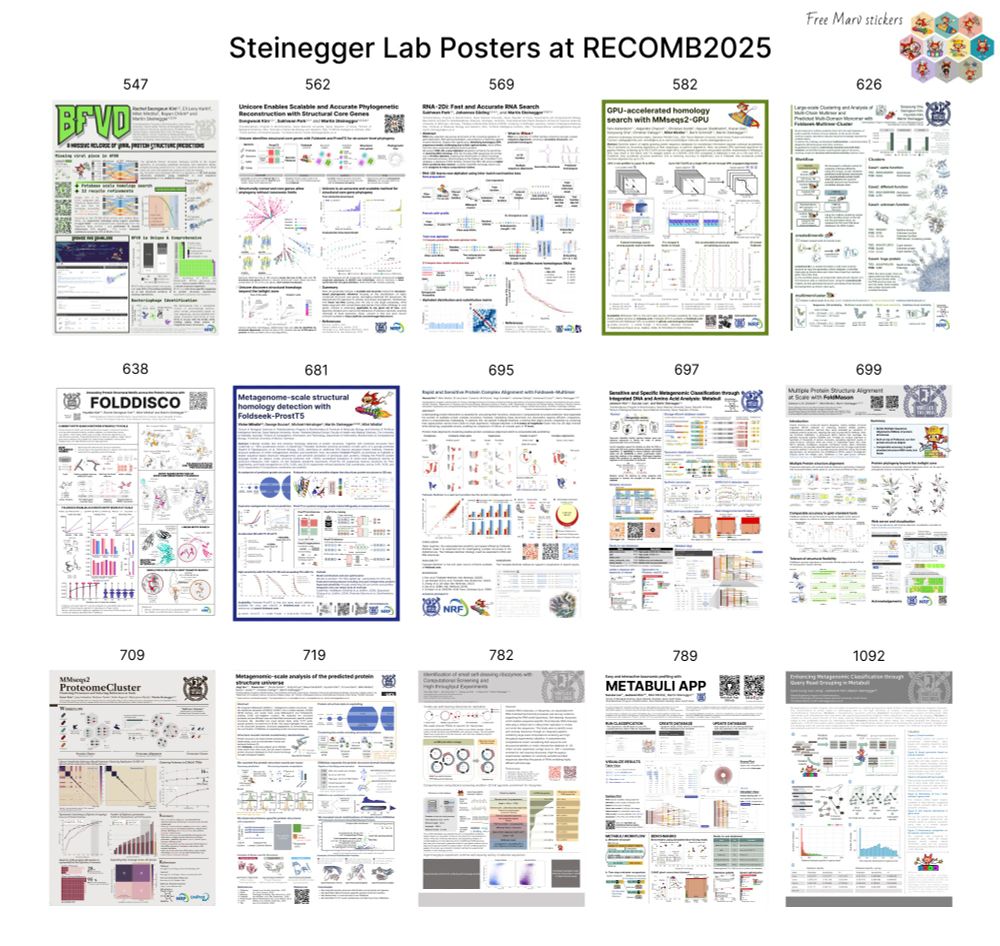

Visit our posters at #RECOMB2025 for:
Structural: MSAs, Virus DB, Core Genes, Motif Discovery, Multimer Clustering & Search, pLM Foldseek, Environmental analysis
Metagenomics: Classification & Metabuli App
GPU-based & RNA search, Proteome clustering, Novel Ribozyme discovery
& get Marv stickers!
25.04.2025 07:45 — 👍 64 🔁 19 💬 2 📌 4

@eunbelivable.bsky.social presented our viral protein structure database BFVD, including the new V2 update with improved predictions using 12 recycles for higher quality structures. Check out the paper and data here:
📄 academic.oup.com/nar/article/...
🌐 bfvd.foldseek.com
#RECOMB2025
25.04.2025 05:02 — 👍 22 🔁 8 💬 1 📌 0

RECOMB2025 Things to do - Google My Maps
RECOMB2025 Things to do
I also updated the main #RECOMB2025 things-to-do map to include more tourist attractions and some of the standout vegan places I have visited myself (except the two places next to Yonsei, which I didn't have a chance to visit yet):
www.google.com/maps/d/edit?...
21.04.2025 13:48 — 👍 7 🔁 2 💬 1 📌 0
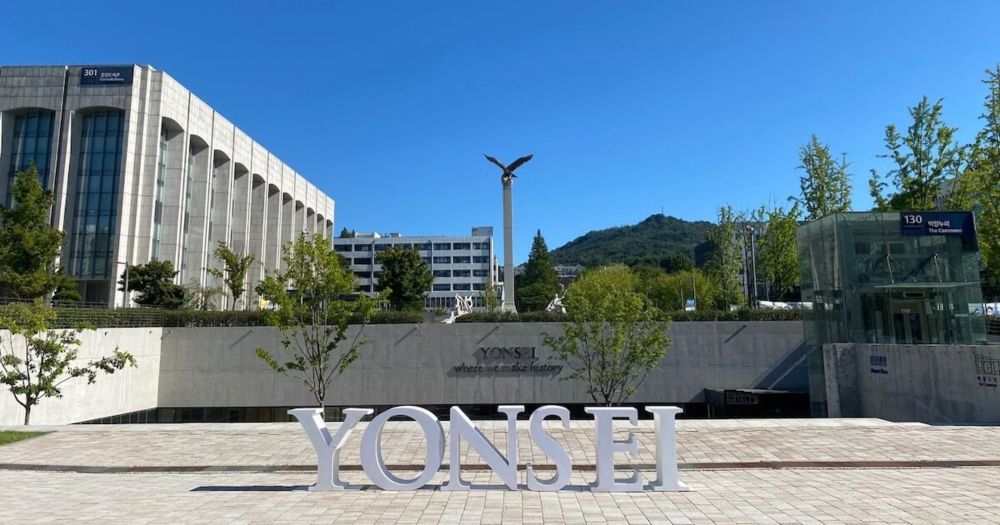

RECOMB 2025 - Things to do
RECOMB 2025 - Seoul, South Korea
Finding vegetarian and vegan food in Korea can be tricky. I added a mini-guide to the #RECOMB2025 things-to-do site with some resources. Let me know if you want more recommendations!
recomb.org/recomb2025/t...
21.04.2025 04:42 — 👍 17 🔁 2 💬 2 📌 0
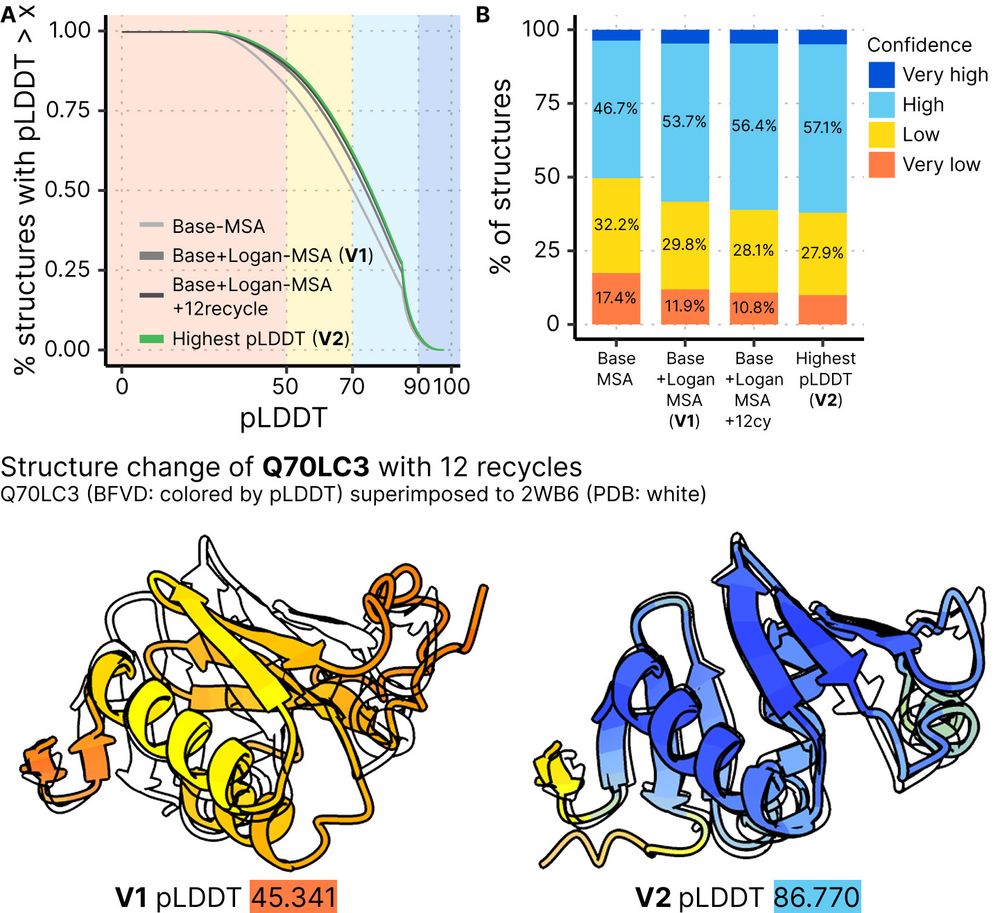
Big Fantastic Virus Database (BFVD) version 2 improves 31% of predictions through 12 ColabFold recycles. PAEs and MSAs now also available for download and in the webserver.
🌐https://bfvd.foldseek.com
💾https://bfvd.steineggerlab.workers.dev/
1/3
31.03.2025 05:07 — 👍 66 🔁 24 💬 2 📌 2
죽고 싶지만 떡뽂이는 먹고싶어!
29.03.2025 12:26 — 👍 0 🔁 0 💬 1 📌 0
Metabuli Databases
🚀 New Metabuli DB is out!
It includes quality-filtered GTDB R220, RefSeq viruses, and the human T2T.
Prokaryotes follow the GTDB taxonomy; viruses and the human genome use NCBI taxonomy.
Download it and expand with genomes of your choice using "updateDB" command.
metabuli.steineggerlab.workers.dev
27.03.2025 07:43 — 👍 3 🔁 0 💬 0 📌 0
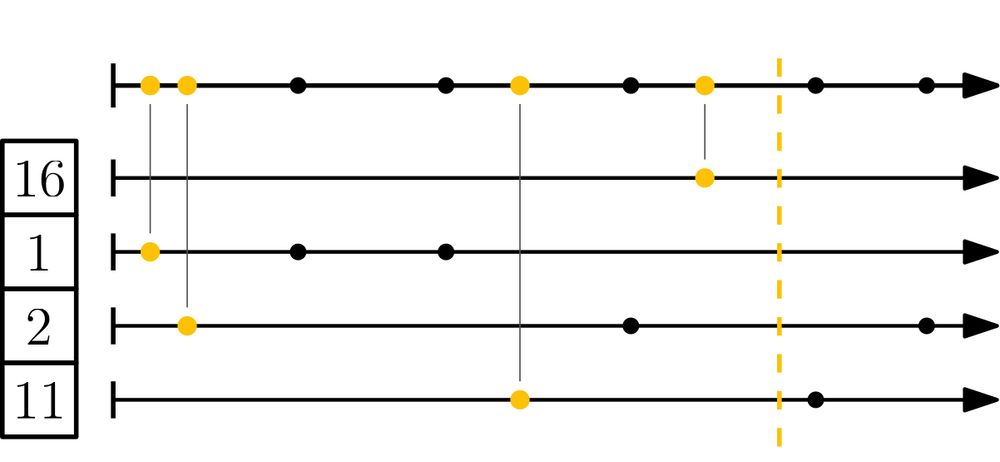
Schematic of the mod-bucket algorithm: all k-mer hashes are partitioned into s buckets via their remainder mod s. Then, in each bucket the smallest hash is selected.
Just published simd-sketch, a crate for fast bucket sketches.
It's 7x to 30x faster than BinDash, by using the simd-minimizers crate for fast hashing, and a nearly branch-free implementation.
Here's a blogpost with a survey of minhash history & methods, and evals:
curiouscoding.nl/posts/simd-s...
14.03.2025 00:35 — 👍 12 🔁 9 💬 1 📌 0
Metabuli App is a team effort! Sunny led the interface design and implementation, while Milot and Martin provided expert guidance throughout the project! @sunjaelee.bsky.social @milot.bsky.social @martinsteinegger.bsky.social Huge thanks and congrats everyone🎉
13.03.2025 08:50 — 👍 2 🔁 0 💬 0 📌 0

⚡Metabuli is now even faster on Linux servers and Windows/macOS computers.
Annotating 22M paired-end human gut reads vs. 36K genomes (8,465 GTDB species) took:
🖥️57 min on a Windows desktop (i9-9900, 32GB RAM)
💻39 min on a MacBook M2 Pro (32GB RAM) 🧵4/5
13.03.2025 08:50 — 👍 2 🔁 0 💬 1 📌 0

🛠️Curate your database.
- Build DBs with GTDB, NCBI, ICTV, or custom taxonomy.
- Add genomes to pre-built DBs to save time (benchmark in figure)
- Expand taxonomy—e.g., integrate ICTV viruses into a GTDB prokaryote DB.
⏱️Building a DB of 8520 GTDB species took 104 min on a MacBook M2 Pro (32G). 🧵3/5
13.03.2025 08:50 — 👍 1 🔁 0 💬 1 📌 0
🔍Explore results with interactive visualization.
- Generate customized Sankey plots.
- Search for taxa of interest.
- Filter by classified reads or proportion.
- Click taxon nodes for subtree views.
- Extract reads classified to a taxon.
- Access NCBI Taxonomy and genome browser. 🧵2/5
13.03.2025 08:50 — 👍 1 🔁 0 💬 1 📌 0
Metabuli App preprint is out!
💻Taxonomic classification & interactive visualization—right on your laptop
🛠️Create new databases or update existing ones with new sequences. 🧵1/5
github.com/steineggerla...
www.biorxiv.org/content/10.1101/2025.03.10.642298v1
13.03.2025 08:50 — 👍 20 🔁 16 💬 1 📌 1
Postdoc at Dana-Farber and Harvard Med with Heng Li (@lh3lh3.bsky.social). Prev: UBC / UofT.
I like thinking about biological sequence analysis and its applications to metagenomics / microbial genomics.
https://jim-shaw-bluenote.github.io
Incoming PI at Institut Pasteur 🇫🇷
Current: Evolutionary Virologist based at the University of Tokyo 🇯🇵
💻🦠🏳️🌈
Trailrunner, Bioinfo Researcher at HKU D24H
linktr.ee/carloshk
Research group in Lille, Fr.
Postdoc on high troughput bioinformatics @ KIT Karlsruhe;
IMO, ICPC, Xoogler, Rust, road-cycling, hiking, wild camping, photography
Associate Professor at Department of Biology, Aarhus University, Denmark 🇩🇰 working on microbial genomics, bioinformatics, cable bacteria, and other sediment microorganisms. https://www.au.dk/en/ianpgm@bio.au.dk/ 🇦🇺
‖ Permanent Researcher / INRIA Start. Faculty @ INRIA Rennes 🇫🇷 ‖
BioInfo/CompBio: algorithms, genomics, pathogens & rapid diagnostic of antibiotic resistance《 https://brinda.eu | https://github.com/karel-brinda 》
Hiring PhDs/Postdocs CGRLab.github.io | Incoming DDLS Assistant Professor, Data Science and AI division, Chalmers University | Pangenomics for studying genome variation and evolution | | Formerly at JHU-LangmeadLab, UNIL-DessimozLab, WUR
Ph.D student in Computer Science and Engineering at the University of Washington working with Su-In Lee.
Computer Science PhD student @ GC, CUNY (Xie Research Group)
Group leader @igmm-montpel.bsky.social studying viruses - ERC Stg- Amateur athlete and musician- Husband and father- Europtimist @biolum-montpellier.bsky.social
https://www.igmm.cnrs.fr/en/team/rna-viruses-and-host-factors/
Doctoral researcher @mpi-bio-fml.bsky.social
The smallest things in life hold so much mystery - I research them 💻, and I love talking about them 💬!
BioDesign, Machine Learning, Drug Discovery | Rosenkranz Award 2021 | Dad | Polyglot | Capybarist | plissonf.github.io
Founding ingeniebio.com
ORCID 0000-0003-224
We’re on a mission to end the headaches of ordering synthetic DNA and give researchers what they really want: reliable DNA, on time, exactly as ordered. #DNASynthesis 🧪
ansabio.com
Machine learning for molecular biology. ELLIS PhD student at Fabian Theis lab. EPFL alumnus.






















