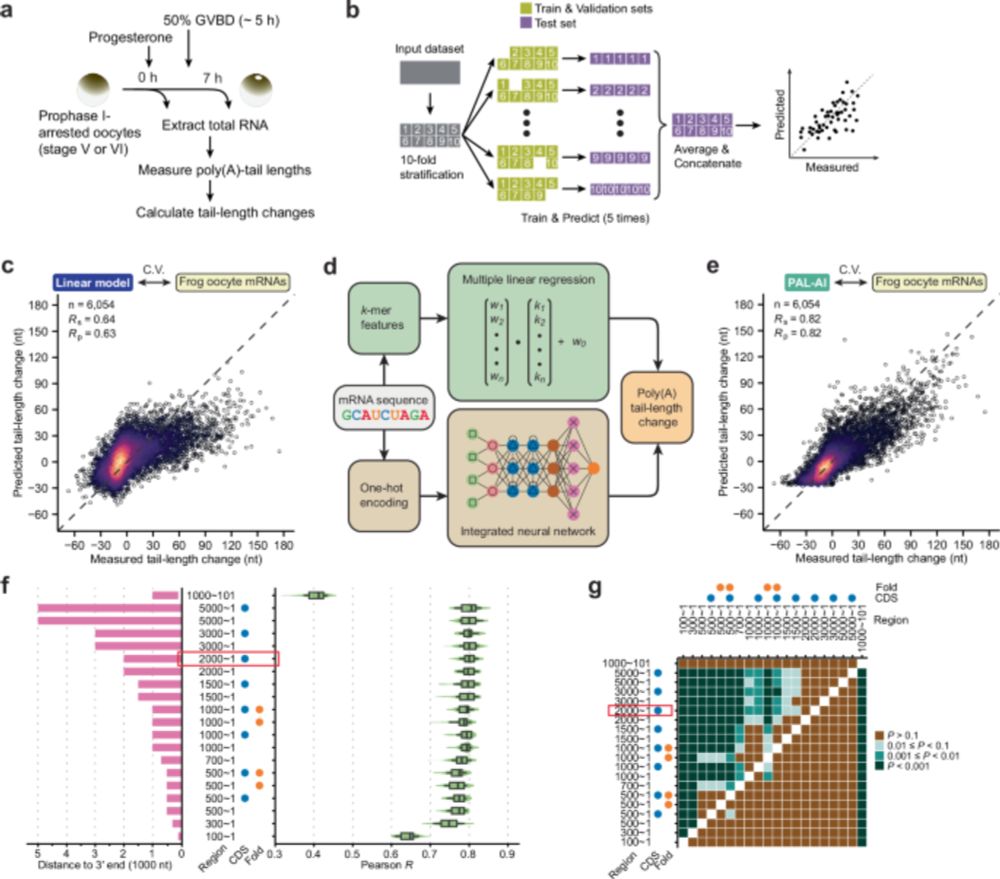
Diverse database and machine learning model to narrow the generalization gap in RNA structure prediction
Length matters—secondary structures of long, diverse RNAs in RNAndria and deep learning model eFold improve prediction accuracy.
Exciting new work from the Rouskin Lab out at @science.org
If you are interested in RNA structure and machine learning, you should probably read it..
Nuevo paper del Rouskin lab!
Si te interesa la estructura de ARN y machine learning, no te lo pierdas..
www.science.org/doi/10.1126/...
25.02.2026 20:06 — 👍 3 🔁 0 💬 0 📌 0
Interesante
09.02.2026 17:47 — 👍 0 🔁 0 💬 0 📌 0
Investigadores del Laboratorio de Anatomía Comparada y Evolución de los Vertebrados del Conicet (Lacev-Conicet) descubrieron un huevo de dinosaurio carnívoro en perfecto estado de conservación y lo anunciaron en un nuevo streaming de paleontología www.lanacion.com.ar/sociedad/inv...
08.10.2025 12:33 — 👍 18 🔁 9 💬 0 📌 1
A must read from Luisa, Emi, and team.
09.09.2025 20:31 — 👍 1 🔁 0 💬 0 📌 0
This work is the result of a collaboration between many wonderful people: @grantyang.bsky.social, Yve, Casper, Matty, Scott, myself, and Silvi from @harvardmed.bsky.social; Dan H from @stanfordpress.bsky.social; and, for the first time as a PI, the great Lauren Hagler from @tamu.bsky.social !!
4/4
31.07.2025 22:06 — 👍 1 🔁 0 💬 0 📌 0


Aside from the app itself, we found that DMS-MaPseq + SEISMICgraph can be used to in vitro validate multiple riboswitches at once using a library, and that many of those riboswitches stop functioning if pseudoU is used instead of regular U (an important finding for those working in RNA tx).
3/4
31.07.2025 22:06 — 👍 1 🔁 0 💬 1 📌 0


The platform offers a plethora of graphs (see pic), and supports data coming from ShapeMapper, RNA framework, and SEISMIC; it should make the life of anybody analyzing DMS/SHAPE-MaP a lot easier :)
2/4
31.07.2025 22:06 — 👍 1 🔁 0 💬 1 📌 0

SEISMICgraph: a web-based tool for RNA structure data visualization
Abstract. In recent years, RNA has been increasingly recognized for its essential roles in biology, functioning not only as a carrier of genetic informatio
Our latest publication (doi.org/10.1093/nar/...) from the Rouskin (www.rouskinlab.com) & Hagler Labs (www.haglerlab.org) is out on @narjournal.bsky.social ! We developed a web interface for easy visualization of RNA structure chemical probing data (from DMS/SHAPE-MaP).
1/4
31.07.2025 22:06 — 👍 4 🔁 1 💬 1 📌 0

Lance 22.1 Mar Del Plata Canyon | SOI Divestream 817
YouTube video by Schmidt Ocean
A wonderful collab between CONICET (Argentina) and the Schmidt Ocean Institute led to the exploration of the Submarine Canyon in Argentina’s deep shores. It is hypnotic to see:
www.youtube.com/live/xg9NLXK...
31.07.2025 18:12 — 👍 0 🔁 0 💬 0 📌 0
A cool one
28.07.2025 12:05 — 👍 1 🔁 0 💬 1 📌 0
Super super cool!
12.06.2025 22:31 — 👍 2 🔁 1 💬 1 📌 0
That looks cool, how did you do it ?!?!
03.06.2025 17:13 — 👍 1 🔁 0 💬 1 📌 0
And on other news. My first contribution to science as a postdoc with @stirlingchurchman.bsky.social is out! We discuss the knowns and unknowns of post-transcriptional splicing. It was really fun to write with the great @karinechoquet
11.04.2025 19:29 — 👍 33 🔁 10 💬 2 📌 0
Everybody should
10.04.2025 14:59 — 👍 1 🔁 0 💬 0 📌 0
Tomorrow at the Systems Virology Journal Club, #AyanoKohlgruber will present her work with #SteveElledge on high-throughput discovery of MHC class I- and II-restricted T cell epitopes using synthetic cellular circuits. pubmed.ncbi.nlm.nih.gov/38956325/
10.04.2025 01:48 — 👍 2 🔁 1 💬 0 📌 0
In vivo folding of nascent RNA! Awesome work from Leo (@leoschaerfen.bsky.social) and team, check it out:
26.03.2025 15:51 — 👍 2 🔁 0 💬 0 📌 0

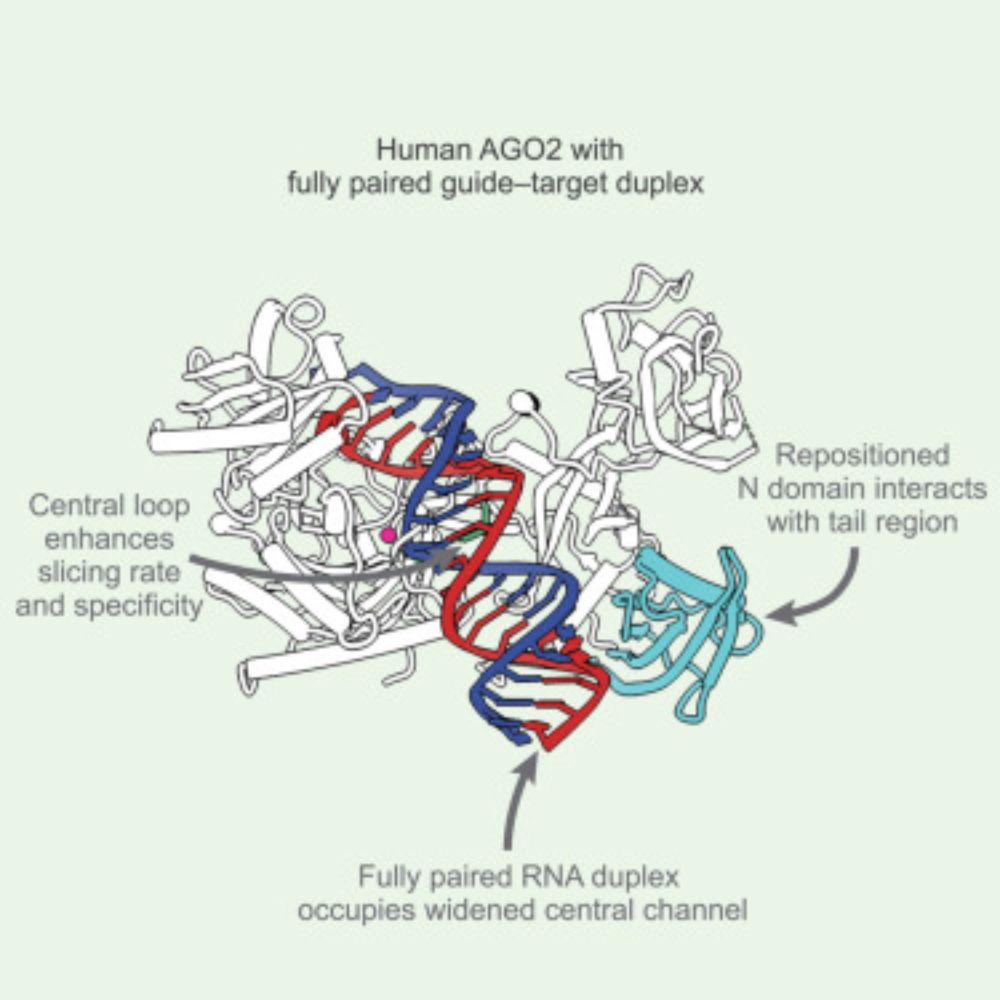
Functional microRNA targeting without seed pairing
MicroRNAs (miRNAs) associate with Argonaute (AGO) proteins to serve as guides, directing binding to partially complementary sites in mRNAs, ultimately causing post-transcriptional repression. Compleme...
Pleased to share my recent work from the Bartel Lab! Some miRNAs recognize novel target sites called 3'-only sites, which pair to the miRNA 3' region but not to the seed. These sites direct mRNA repression in cells and are engaged by different mechanisms to seed sites
www.biorxiv.org/content/10.1...
12.03.2025 21:57 — 👍 6 🔁 2 💬 0 📌 1
I would have to say circRNAs, never ending possibilities (since they have no ends)
02.03.2025 01:26 — 👍 2 🔁 0 💬 1 📌 0
MicroRNAs are just so much fun
01.03.2025 23:20 — 👍 1 🔁 0 💬 1 📌 0

RNA xkcd.com/3056
26.02.2025 14:58 — 👍 17855 🔁 2560 💬 154 📌 171
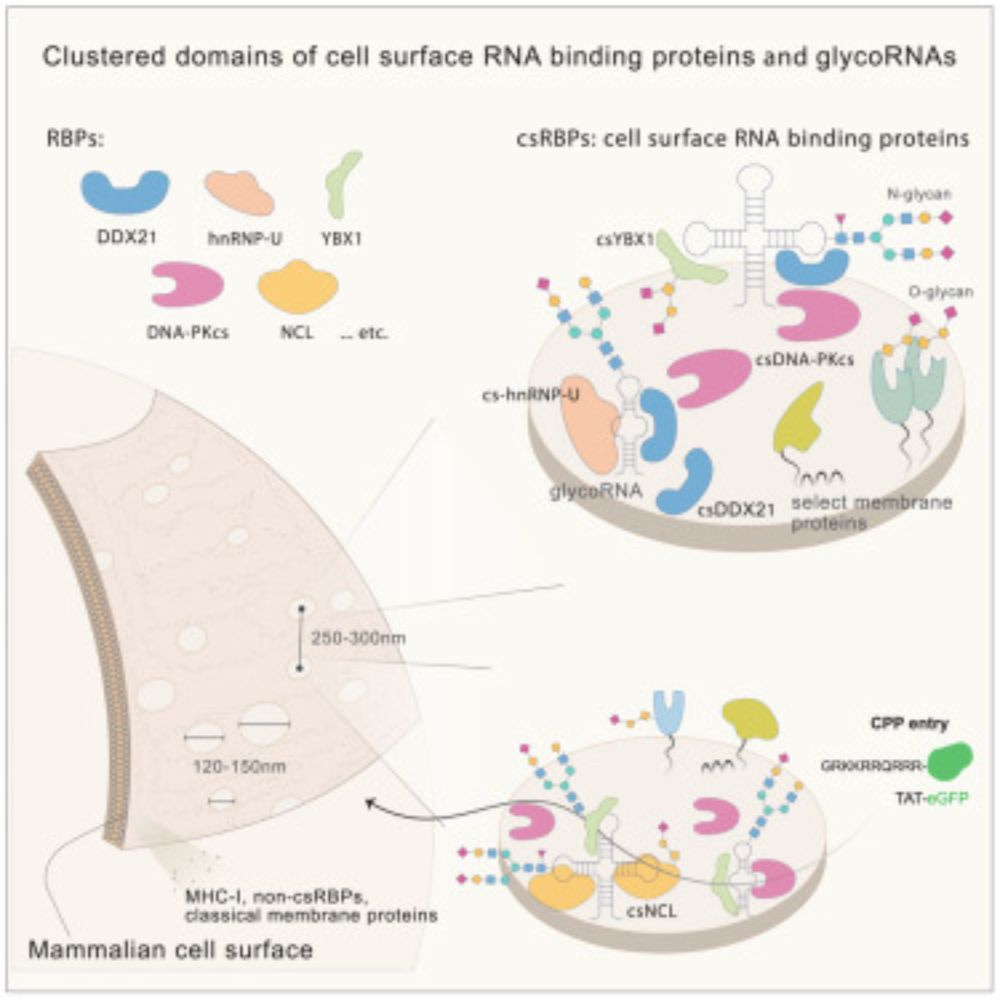
RNA-binding proteins and glycoRNAs form domains on the cell surface for cell-penetrating peptide entry
Mammalian cells present RNA-binding proteins on the cell surface that form clustered
domains containing glycoRNAs.
Excited to share the work of Jonathan Perr where he uncovered a surprisingly common feature of cell surfaces - the presentation and clustering of RNA binding proteins with #glycoRNA.
Critically support by #NIH @cp-cell.bsky.social www.cell.com/cell/abstrac...
27.02.2025 15:48 — 👍 186 🔁 62 💬 11 📌 8

Our very own Scott Grote got into Harvard’s PHD program in Virology! #ProudLab #RouskinLab
08.02.2025 01:00 — 👍 2 🔁 0 💬 0 📌 0

Has Bluesky replaced X for scientists? Take Nature’s poll
The research community has flocked to the social-media platform Bluesky. Tell us about your experience.
We have a feeling you'll be interested in this one.
Nature is keen to find out how scientists are using Bluesky and whether it has become their go-to social-media platform. Take the survey! 🧪
14.01.2025 23:39 — 👍 256 🔁 108 💬 19 📌 9
Felicitaciones Rodri y equipo!
04.12.2024 14:26 — 👍 1 🔁 0 💬 0 📌 0
Assistant Professor at UMich with a focus on computational pharmacology (he/him)
#PeriodismoPuro en tiempo real
Noticias de #Argentina y del mundo 🌎
Lee todas nuestras notas en ⬇
https://linkbe.me/perfilcom
Executive Editor/Team Lead Open Access Science & Medicine Journals Sage Publishing
Opinions = mine
http://linkedin.com/in/jlovick-editor
#oncology #cancerresearch #medicine #biology #cardiology #neurology #microbiology #publichealth #healthcare
Porteña. Me pierdo en la ciudad para encontrarme.
A veces publico sobre edificios antiguos, a veces sobre perritos...
https://www.instagram.com/buenosairesperdida/
News, features, analysis and opinion about Argentina.
buenosairesherald.com
Science integrity consultant and crowdfunded volunteer, PhD.
Ex-Stanford University. Maddox Prize/Einstein F Award winner
NL/USA/SFO.
#ImageForensics
@MicrobiomDigest on X.
Blog: ScienceIntegrityDigest.com
Support me: https://www.patreon.com/elisabethbik
Proud worm researcher. Interested in miRNAs, all RNAs really, and gene regulation in general. A biochemist turned geneticist trying to think about questions of cell biology. Lucky to lead a lab at Johns Hopkins School of medicine. https://cochellalab.org/
Since 1889 🗞️
Sign up for our newsletters and alerts: http://wsj.com/newsletters
Got a tip? http://wsj.com/tips
Follow our staff: https://go.bsky.app/2ppWqxF
El valor de entender lo que pasa 📱💻 Suscribite y accedé sin límites: http://bit.ly/3kYTs3i
David Bartel's lab @WhiteheadInst @MIT @HHMI | microRNAs, mRNAs, and other RNAs
Ex editora de Ciencia y Salud en La Nación. Ahora, en
@eldestapeweb. "Rebelión en el Laboratorio", "Diez preguntas que la ciencia (todavía) no puede contestar". Los viernes de 19 a 20 en "Al infinito y más allá", AM1070,YouTube y Canal 20 de Telecentro
Superpresent is a quarterly magazine of the arts founded in 2020. We place equal emphasis on visual arts and the written word.
Website: www.superpresent.org
The Oxford RNA Club is a dynamic community of researchers focused on all aspects of RNA biology, from gene regulation to disease. We host in-person monthly seminars, and larger events throughout the year. Join us here: https://rna.web.ox.ac.uk/home
Toda la Información deportiva argentina e internacional al instante.
X/Instagram/Facebook: @deportes24ar
Mail: deportes24ar@gmail.com
An RNA lab in Santa Cruz, working on splicing and the spliceosome, asking how splicing capacity is allocated to the primary transcriptome, and how intron gain affects genome evolution.
Nucleic Acids Research (NAR), from Oxford University Press, publishes the results of leading-edge research into physical, chemical, biochemical and biological aspects of nucleic acids and proteins involved in nucleic acid metabolism and/or interactions
Co-lead R&D at MGZ Medical Genetics Center Munich • Former PhD student/postdoc in @NovoaLab at @CRGenomica • she/her
Professor, Molecular & Cellular Biology, Harvard University