
Features reconstruction errors between the latent spaces of several universal MLIPs
No day goes by without a new universal #ML potential. But how different they really are? Sanggyu and Sofiia tried to give a quantitative answer by comparing the reconstruction errors between their latent-space features. If you are curious, check out the #preprint arxiv.org/html/2512.05...
09.12.2025 07:16 — 👍 11 🔁 3 💬 0 📌 0

Congrats to 🧑🚀 Sergey Pozdnyakov who received a distinction (best 8% of theses at @materials-epfl.bsky.social) for his PhD thesis "Advancing understanding and practical performance of machine learning interatomic potentials". Поїхали 🚀! infoscience.epfl.ch/entities/pub...
10.12.2025 12:50 — 👍 10 🔁 2 💬 0 📌 0

👏 Congratulations to Prof. @micheleceriotti.bsky.social (@labcosmo.bsky.social) and Prof. Roland Logé (LMTM)
on their promotion to Full Professor of Materials Science in the School of Engineering!
More info: actu.epfl.ch/news/appoint...
19.09.2025 09:15 — 👍 3 🔁 1 💬 2 📌 0

A cartoon explaining how mild finite-temperature conditions induce disorder and dynamical reconstruction on the surfaces of lithium thiophosphates
📢 Now out on @physrevx.bsky.social energy, journals.aps.org/prxenergy/ab... from 🧑🚀 @dtisi.bsky.social and Hanna Türk, our #PET -powered study of the dynamic reconstruction of LPS surfaces, and how it affects their structure, stability and reactivity.
27.08.2025 06:54 — 👍 9 🔁 4 💬 1 📌 0
Just before my last week in @labcosmo.bsky.social. Our metatensor and metatomic paper is out! A collection of the hard work we’ve done at @labcosmo.bsky.social to make atomistic machine learning easier to use for experts and not alike.
22.08.2025 09:56 — 👍 1 🔁 0 💬 0 📌 0
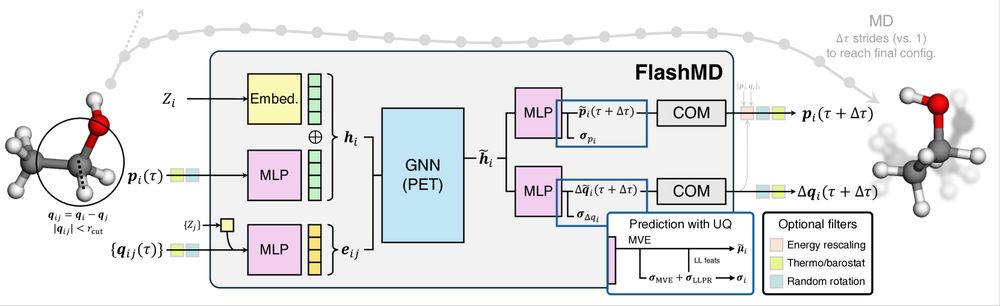
Scheme of the GNN architecture of the FlashMD method.
📢 Running molecular dynamics with time steps up to 64fs for any atomistic system, from Al(110) to Ala2? Thanks to 🧑🚀 Filippo Bigi and Sanggyu Chong, with some help from Agustinus Kristiadis, this is not as crazy as it sounds. Let us briefly introduce FlashMD⚡ arxiv.org/html/2505.19...
27.05.2025 07:02 — 👍 37 🔁 12 💬 1 📌 1
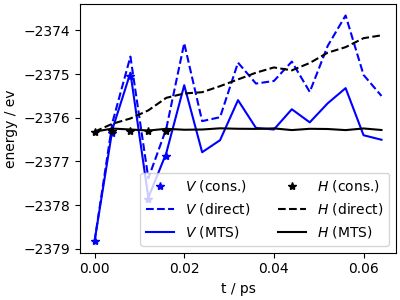
Plot of the potential and total energy trends along a MD trajectory, showing drift for non-conservative forces, and how that is fixed using multiple time stepping.
The PET-MAD universal forcefield mingled with the dark side, and got twice as fast 🚀. Read on, or head to the 🧑🍳📖 atomistic-cookbook.org/examples/pet..., if you are curious of what this is all about. #atomistic-cookbook #compchem #machinelearning #mlip🧵
07.05.2025 05:24 — 👍 10 🔁 3 💬 1 📌 0

A dynamical surface morphology implies changing electronic structure properties of the surface. We found large fluctuations of the electrostatic potential during dynamics, which will be crucial for the surface reactivity.
21.04.2025 09:25 — 👍 0 🔁 0 💬 0 📌 0

We studied the evolution of the SOAP descriptors of the tetrahedra on the surface, for an atomistic analysis of their dynamics. We time-averaged the SOAP vector to rule out the effects of fast-moving Li atoms
21.04.2025 09:25 — 👍 1 🔁 0 💬 1 📌 0

The surface reconstruction increases the atom density and reduces the Li diffusion
21.04.2025 09:25 — 👍 0 🔁 0 💬 1 📌 0

The main takehome is: static calculation are not enough!!
After MD relaxation, the effect of temperature reduces the differences in surface energy and coordination number of the different surface cuts, in agreement with a general amorphisation of the surfaces.
21.04.2025 09:25 — 👍 0 🔁 0 💬 1 📌 0
PET-MAD, a universal interatomic potential for advanced materials modeling
Find (many) more benchmarks and tests in the preprint arxiv.org/html/2503.14..., try PET-MAD for yourself, and let us know if you manage to break it - we're already working towards PET-MAD-2 😅 #compchem @nccr-marvel.bsky.social @materials-epfl.bsky.social @erc.europa.eu @cscsch.bsky.social
19.03.2025 07:23 — 👍 8 🔁 1 💬 0 📌 0
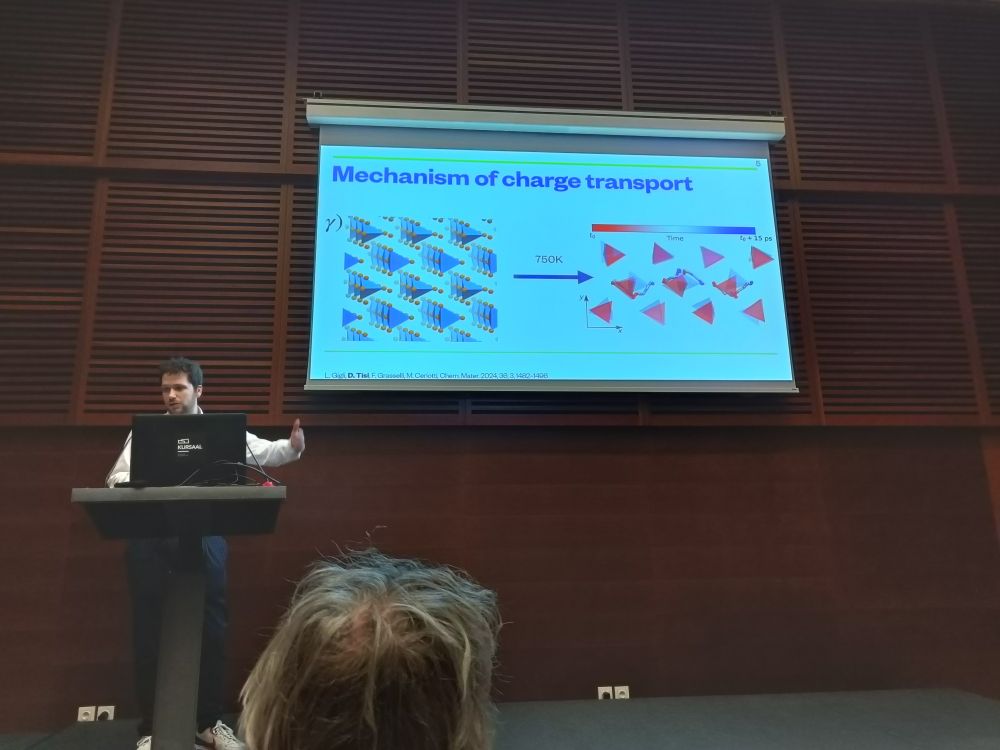


@labcosmo.bsky.social continuing its strong performance with Davide Tisi's talk on transport mechanisms in solid-state electrolytes!
09.04.2025 14:35 — 👍 4 🔁 2 💬 0 📌 0
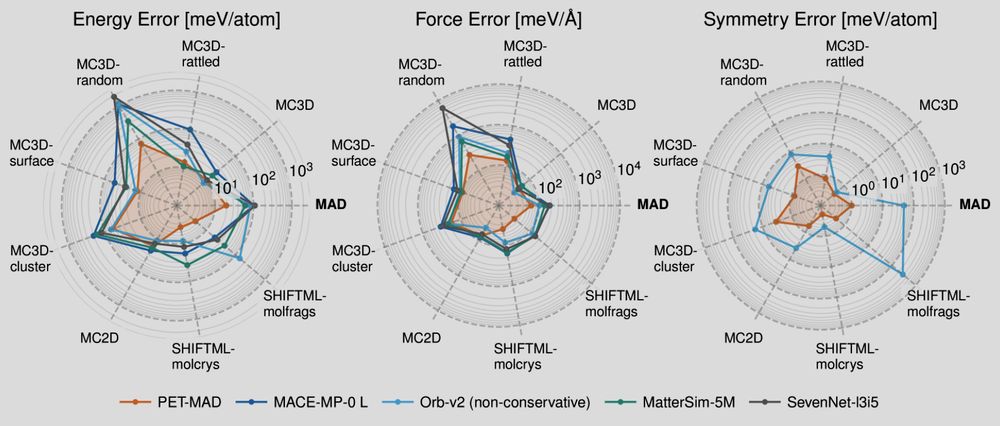
Polar plot showing the errors of several machine-learning potential of different test sets. Smaller is better here!
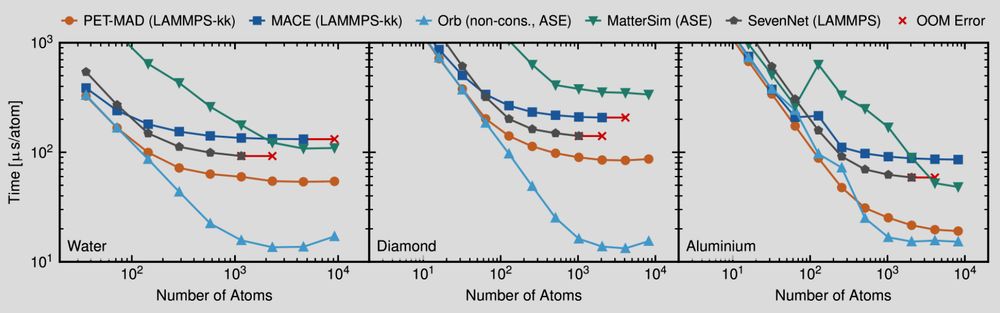
Plots showing the evaluation time per atom for several machine-learning potentials as a function of the number of atoms in a simulation. Smaller is better
📢 PET-MAD has just landed! 📢 What if I told you that you can match & improve the accuracy of other "universal" #machinelearning potentials training on fewer than 100k atomic structures? And be *faster* with an unconstrained architecture that is conservative with tiny symmetry breaking? Sounds like 🧑🚀
19.03.2025 07:23 — 👍 28 🔁 9 💬 1 📌 3
Science. Humanity.
IMDEA Nanoscience, Madrid
Mathematics Sorceror (sensory alchemist) at the Arctangent Transpetroglyphics Algra Laboratory (ATAL), I transflarnx mathematics into living rainbows. http://owen.maresh.info https://github.com/graveolensa
Psoeppe-Tlaxtlal, (an undreamt splendour?)
Computational Chemist. 🦀🐍 Scientist at BAM
+ Professor FSU Jena. Only private opinions here.
Last name: German pronunciation. 🏳️🌈 she/they (sie).
@JaGeo on github
We focus on quantitative #modeling of #material properties and functions, especially in #catalysts and #energy conversion devices.
Based in Berlin Dahlem.
Website: https://www.fhi.mpg.de/th-department
Molecular modeling, simulations, metadynamics, machine learning
Computational chemist with a Bayesian taint. Working on foundational cross-language tools at @labcosmo.bsky.social.
More @ https://rgoswami.me
PhD student in @labcosmo.bsky.social
Theoretical chemist studying the quantum properties of molecules.
@royalsociety.org University Research Fellow at @uclchemistry.bsky.social.
Previously @downingcollege.bsky.social and @newcollegeoxf.bsky.social.
www.hughburton.com | x.com/HughGABurton
Bioinformatics Scientist / Next Generation Sequencing, Single Cell and Spatial Biology, Next Generation Proteomics, Liquid Biopsy, SynBio, AI/ML in biotech // http://albertvilella.substack.com
DeepMind Professor of AI @Oxford
Scientific Director @Aithyra
Chief Scientist @VantAI
ML Lead @ProjectCETI
geometric deep learning, graph neural networks, generative models, molecular design, proteins, bio AI, 🐎 🎶
I'm a theoretical physicist at Durham University
Associate professor at the University of Milano.
Biophysics and computational structural biology.
http://compsb.unimi.it
Developer @plumed.org
https://orcid.org/0000-0002-9923-8590
Assistant professor (RTDb) in Theoretical Physics of Matter
Department of Applied Science and Technology (DISAT)
Politecnico di Torino, Italy
https://www.gmpavanlab.com
Full professor at SISSA, msb.sissa.it group | Group leader @bussilab.org | Founder and developer @plumed.org
We are the biomolecular and pharmaceutical modelling laboratory directed by Prof. Gervasio at Uni Geneva and UCL London. We develop enhanced sampling algorithms and perform simulations on systems bio-pharmaceutical interest (GPCRs, kinases, ...)
Lab Head & Associate Director at @IITalk. Exec Editor of JCTC (@ACSPublications). Director http://www.cecamsimul.eu. Co-Founder @IAMABiotech. Passionate about science!















