2 PhD candidates in my group: Michal @malszycki.bsky.social and Lisa led this work. We are very thankful to Ay lab ( @ferhatay.bsky.social ) and Alev lab (ASHBi, Kyoto), who helped make our story more complete. This work was funded by @dfg.de in the framework of @spp2202.bsky.social
25.02.2026 16:01 —
👍 4
🔁 1
💬 0
📌 0

Analysis of the evolution of this gene architecture and the presence of GC-rich isochores revealed that these features are specific to amniotes. By using the pA-RNA signal as a proxy for speckles we show the speckles are present in amniotes and absent in fish and invertebrates.
25.02.2026 16:01 —
👍 5
🔁 1
💬 1
📌 0

The short & GC-rich introns emerged in amniotes and were once long and GC-poor (in fish). The emergence of this gene architecture coincides with the expansion of SON’s IDRs. IDRs of SON are necessary and sufficient to form functional speckles.
25.02.2026 16:01 —
👍 1
🔁 1
💬 1
📌 0

Problems in splicing, due to SF3 complex dispersal, cause changes in gene expression, but only a small set of exons-introns are sensitive to speckles loss. These genes have introns with short & GC-rich architecture and are found in GC-rich isochores.
25.02.2026 16:01 —
👍 1
🔁 1
💬 1
📌 0

Differential gene expression analysis upon speckles loss reveals only genes within high GC chromatin domains are susceptible. The distance of the chromosomes 18 (lowest GC content) and 19 (highest GC content) from the speckles also indicate their speckle dependency.
25.02.2026 16:01 —
👍 1
🔁 1
💬 1
📌 0

Speckles take a large space in the human cell nucleus and form viscoelastic boundaries that restrict random chromatin movements. Upon speckles loss, the chromatin mobility increases and active(A) and inactive(B) chromatin compartments mix as shown by the Pore-C analysis by @ferhatay.bsky.social lab
25.02.2026 16:01 —
👍 3
🔁 1
💬 1
📌 0

Nuclear speckles are protein and RNA rich condensates that are proximal to GC-rich regions of the human genome. We dissolved speckles by rapidly depleting the 2 core proteins; SON & SRRM2 and investigated the role of speckles in: 1. 3D genome folding 2. transcription & splicing.
25.02.2026 16:01 —
👍 2
🔁 1
💬 1
📌 0
Redirecting
Our most recent work on the “function and evolution” of #nuclear-speckles is now online at Cell @cp-cell.bsky.social
doi.org/10.1016/j.ce...
Read the thread👇 for the highlights of our findings.
25.02.2026 16:01 —
👍 114
🔁 56
💬 9
📌 5
RECOMB-RSG 2026 | Regulatory Genomics Satellite
Thanks to
@ferhatay.bsky.social
and Aly Khan, we’re excited to announce a new chapter for RECOMB-RSG 2026. After years with ISCB/DREAM, we are transitioning to an official RECOMB satellite meeting (May 25 in Thessaloniki). recomb-rsg.github.io
27.01.2026 04:21 —
👍 4
🔁 4
💬 1
📌 0

From RIME to reason
In this Research Letter by Hal M. Hoffman @ferhatay.bsky.social & team, scRNA-seq analysis before & after treatment of recurrent infectious mucocutaneous eruption (RIME) reveals dysregulation of TNF/IFN signaling & classical monocytes: doi.org/10.1172/jci....
14.01.2026 14:02 —
👍 3
🔁 3
💬 1
📌 0
It did fly! Happy new year Jiao, thank you!
08.01.2026 07:22 —
👍 1
🔁 0
💬 0
📌 0

10th year anniversary of Ay lab at LJI! What a journey so far! Thank you to all lab members, collaborators, LJI family, who made this enjoyable and exciting. Here is a photo that summarizes how my lab sees me now 😂😂😢😢
06.01.2026 23:50 —
👍 7
🔁 0
💬 1
📌 0
Happy to see this out @insight.jci.org The first paper of hopefully many with Reid Oldenburg, Croker lab and others at UCSD utilizing BD Rhapsody platform for single-cell profiling of patient samples for severe conditions such as RIME.
25.11.2025 22:27 —
👍 4
🔁 1
💬 0
📌 0
Our source code, utility scripts and links to processed data and results are all available on our lab's GitHub: lnkd.in/g7EdGYuu
Hope you enjoy reading it and reach out if you have any questions or feedback!
Big thanks to Cell Reports Methods and their editorial team for the efficient review
04.11.2025 22:58 —
👍 1
🔁 1
💬 0
📌 0
Our results highlight the importance of distance stratification in capturing differences in long-range loops, differences in sensitivity across different statistical models and provides overall best practices for differential HiChIP analysis.
04.11.2025 22:58 —
👍 1
🔁 0
💬 1
📌 0
good collaborator Katia Georgopoulos in annotation of the results, we implemented a unified framework with all different approaches to date, developed performance metrics and systematically evaluated tools/tests utilized by us and others on multiple different HiChIP datasets.
04.11.2025 22:58 —
👍 1
🔁 0
💬 1
📌 0
We and others have worked on this problem but realized the variability in effectiveness of different approaches across different data sets. In this work, led by Sourya Bhattacharyya (now at Empirico) and Daniela Salgado Figueroa (UCSD Bioinformatics PhD student) in my lab and with help from our
04.11.2025 22:58 —
👍 1
🔁 0
💬 1
📌 0
Our latest paper on comparative analysis of HiChIP data is now online! HiChIP is one of the most useful/practical assays to profile 3D genome organization and chromatin loops but has its challenges in the data analysis especially when it comes to comparative analysis.
04.11.2025 22:58 —
👍 12
🔁 3
💬 1
📌 0
I have
NIGMS R35, impact score 12
NIHGRI R21, 4th percentile
NHGRI R01, 7th percentile (co-I)
and it seems like none will be funded. 0/3.
PO (who has been very helpful) said "Unfortunately, I do not expect this application will be selected for funding in FY25."
😭
19.08.2025 23:30 —
👍 159
🔁 43
💬 42
📌 16
Oh my god 😱😱 good luck Anders. These are amazing scores..We have multiple single digit %ile grants with collaborators.. one is a resubmission after multiple submissions finally making “the cut” just to have the goal post moved 😥 fingers crossed for some last minute miracle for all of us
21.08.2025 06:17 —
👍 1
🔁 0
💬 0
📌 0

RegSys Day 2!
Starting the day with a keynote by Roser Vento-Tormo on the cellular components of the placenta and uterus interaction
#ismbeccb2025
24.07.2025 08:25 —
👍 4
🔁 1
💬 0
📌 0

Today marks the start of the #RegSys @iscb-regsys.bsky.social track at ##ISMBECCB2025! Very much looking forward to listening to the exciting science that will be presented.
Join us in room 11BC today and tomorrow.
23.07.2025 07:41 —
👍 5
🔁 5
💬 0
📌 0
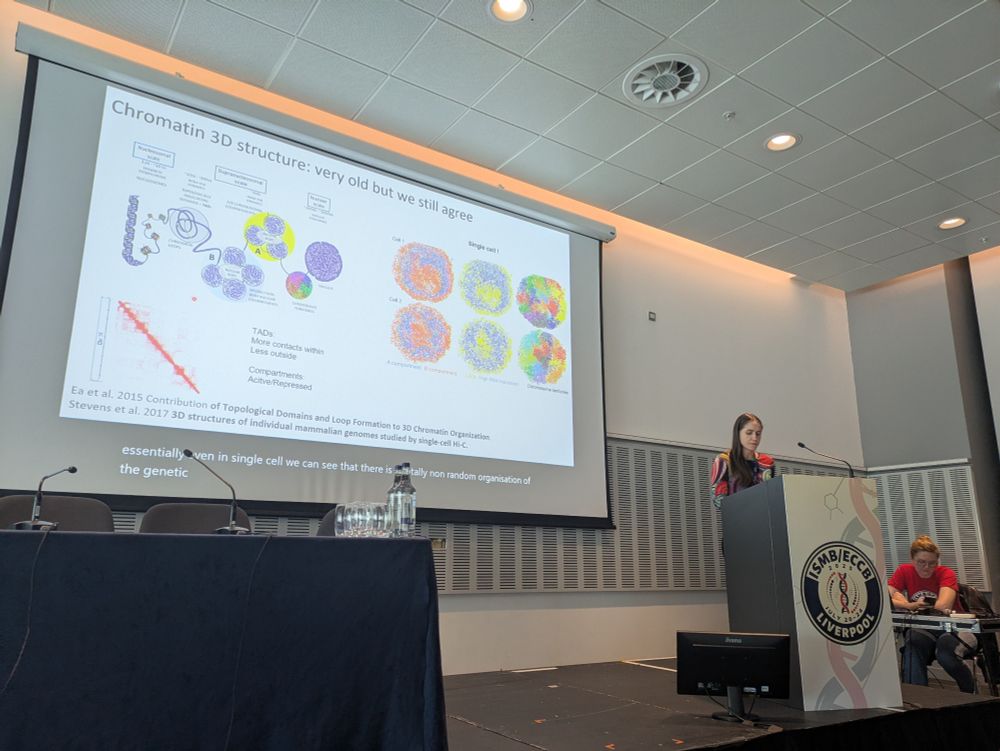
#RegSys is starting with the keynote from @verapancaldi.bsky.social who talks about the changes of chromatin structure and the epigenome in cancer patients #ismbeccb2025
23.07.2025 10:34 —
👍 8
🔁 3
💬 0
📌 0

Laura Hinojosa talks about TFs that regulate replication timing
#RegSys #ismbeccb2025 @ferhatay.bsky.social
23.07.2025 11:55 —
👍 4
🔁 2
💬 0
📌 0
3 years ago we decided to compile these datasets in one place. We were fortunate to get NIH support for this, which transformed it from a local resource for our lab to a comprehensive data resource. What we named Loop Catalog is now a web-based database featuring loop calls from
23.07.2025 08:52 —
👍 1
🔁 0
💬 1
📌 0
The third one is work led by Joaquin Reyna (former UCSD Bioinformatics PhD student) and Kyra Fetter (former UCSD undergraduate student) with help from multiple members of our lab. Seeing the increase in the number and quality of HiChIP datasets and having developed tools for its analysis,
23.07.2025 08:52 —
👍 1
🔁 0
💬 1
📌 0














