Thank you!
04.12.2025 19:25 — 👍 0 🔁 0 💬 0 📌 0Bohan Ni
@bohanni.bsky.social
PhD student in CompSci, Johns Hopkins University. I try to find rare and common variants with impacts on traits/health.
@bohanni.bsky.social
PhD student in CompSci, Johns Hopkins University. I try to find rare and common variants with impacts on traits/health.
Thank you!
04.12.2025 19:25 — 👍 0 🔁 0 💬 0 📌 0
Excited to share our work from the Battle Lab! It’s on an innovative deep learning model architecture incorporating biological domain knowledge to predict traits from large scale proteomic data: www.medrxiv.org/content/10.1...
27.11.2025 17:14 — 👍 5 🔁 1 💬 0 📌 1
Really excited to share our new PRS method, developed with @aprilkim.bsky.social and @alexisbattle.bsky.social ! Our approach is to use a lot of recently developed functional annotations to better estimate the weights of the SNPs.
www.medrxiv.org/content/10.1...
Check out the amazing work from my colleague and friend!
15.05.2025 15:27 — 👍 2 🔁 0 💬 0 📌 0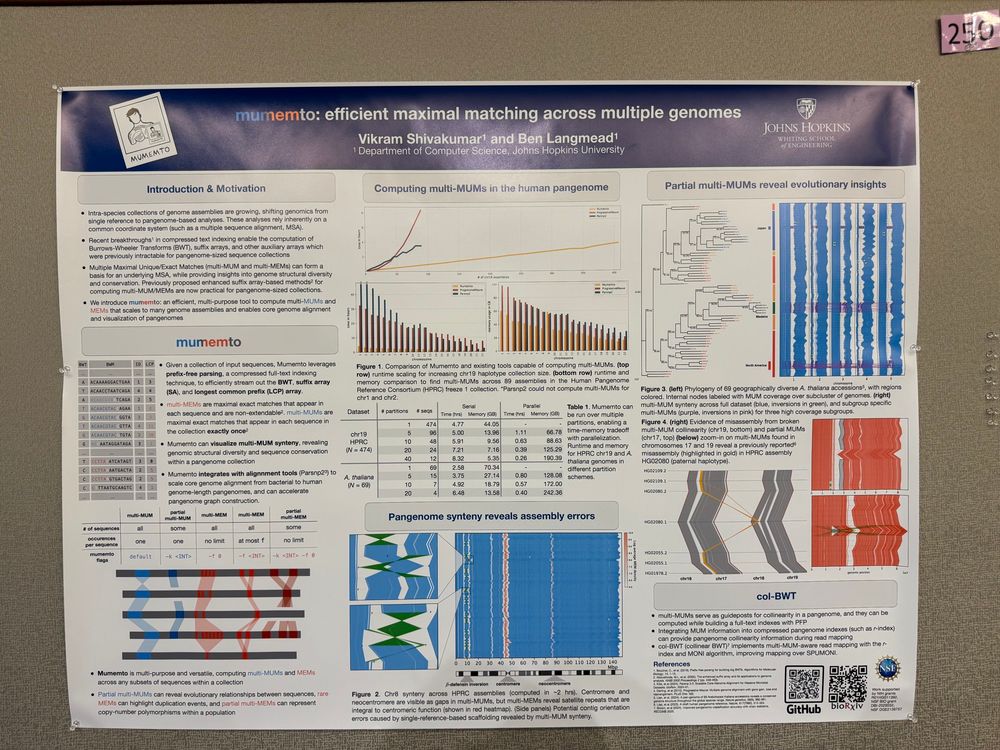
Excited to share our latest work on comparing and visualizing multiple genome assemblies to identify conservation and structural variation in pangenomes with Mumemto! Check out poster 250 at #bog25 if you are here. New preprint coming very soon 👀
09.05.2025 16:26 — 👍 34 🔁 14 💬 0 📌 2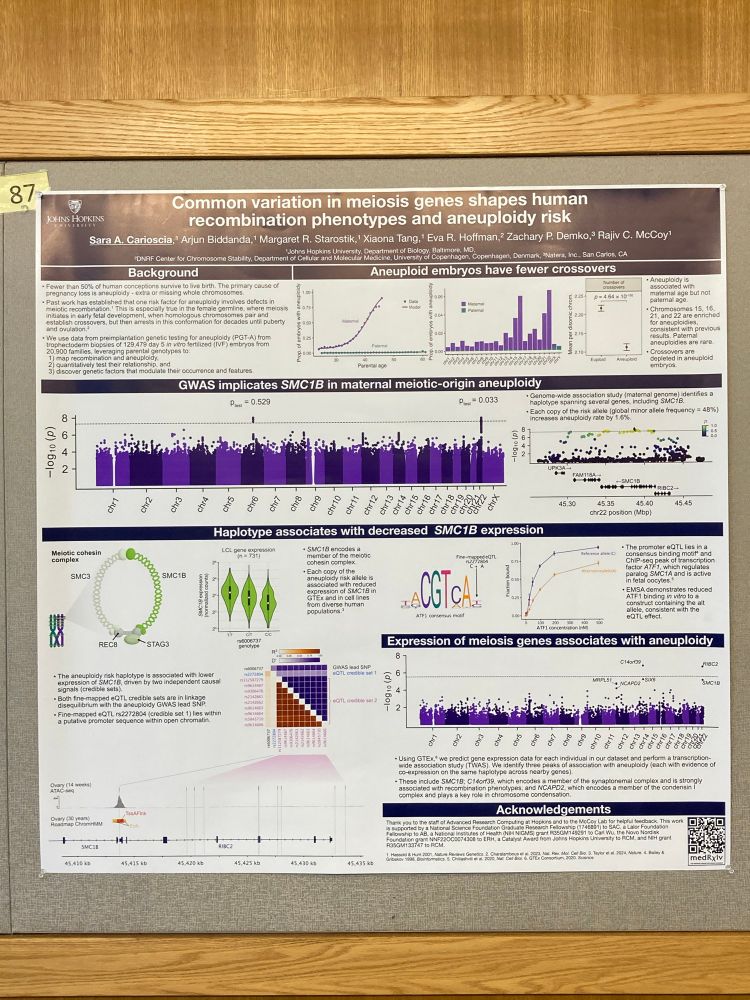
If you are here at #bog25 please check out my poster (number 87) tonight! 😁 Showing our work on common variation associated with aneuploidy in human embryos
07.05.2025 17:26 — 👍 20 🔁 7 💬 0 📌 01/n 🚨Very excited to share our recent work!🚨
To understand gene regulation across diverse environmental conditions and cellular contexts, we treated a broad array of human cell types with three environmental exposures in vitro.
www.biorxiv.org/content/10.1...
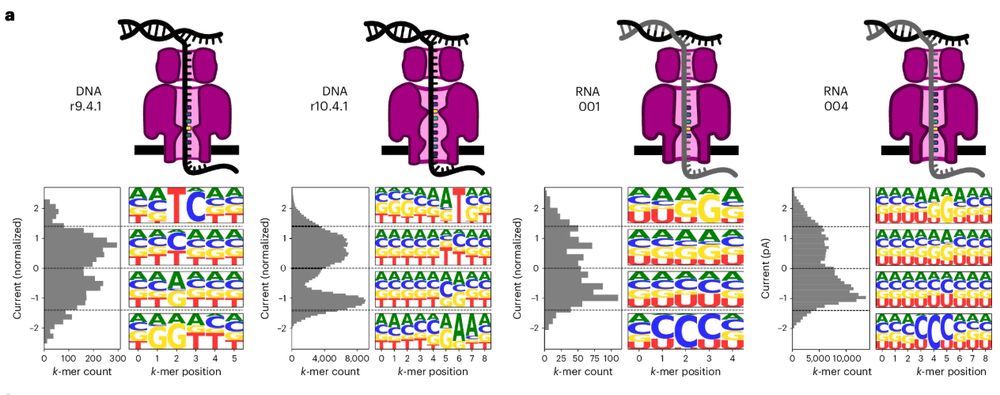
Uncalled4: a toolkit for nanopore signal alignment, analysis and visualization of DNA and RNA modifications.
www.nature.com/articles/s41...
SV prioritization tools in both prioritizing coding and noncoding variant benchmarks. Finally, we were able to prioritize confident rare disease candidate SVs and genes in the Undiagnosed Disease Network data. To learn more, read our paper here: genome.cshlp.org/content/earl....
26.03.2025 14:30 — 👍 0 🔁 0 💬 0 📌 0After extensive analyses of rare SVs near gene expression outliers using long and short-read sequencing, we developed Watershed-SV, extending Watershed, to prioritize functional rare SVs and genes they impact. Our model significantly improves from a baseline model and state-of-the-art
26.03.2025 14:30 — 👍 0 🔁 0 💬 1 📌 0Happy to share our work characterizing functional rare SVs in rare diseases with long-read genome sequencing and transcriptomic outlier data: genome.cshlp.org/content/earl...
26.03.2025 14:30 — 👍 10 🔁 7 💬 1 📌 1
Excited to share a preprint for (w/ @benlangmead.bsky.social) our new tool, Mumemto, on biorxiv! Mumemto finds multi-MUMs across pangenomes (i.e. mummer but for pangenomes). It can rapidly visualize synteny, identify misassemblies, and accelerate core genome and multiple alignment, highlighting SVs.
06.01.2025 15:27 — 👍 34 🔁 29 💬 1 📌 2Mumemto: efficient maximal matching across pangenomes https://www.biorxiv.org/content/10.1101/2025.01.05.631388v1
06.01.2025 03:46 — 👍 22 🔁 12 💬 0 📌 0