super cool
14.02.2026 11:44 — 👍 0 🔁 0 💬 0 📌 0Guifeng Wei
@guifengwei.bsky.social
RNA modifications, X-inactivation, epigenetics, bioinformatics https://sites.google.com/view/guifengwei/
@guifengwei.bsky.social
RNA modifications, X-inactivation, epigenetics, bioinformatics https://sites.google.com/view/guifengwei/
super cool
14.02.2026 11:44 — 👍 0 🔁 0 💬 0 📌 0
Raymond stands at the centre of the picture next to the entrance to the seminar room. He is surrounded by five family members including his wife.
Yesterday we came together to honour Professor Raymond Dwek through the naming of the Raymond Dwek Seminar Room. Raymond's family (pictured) joined the celebration and Raymond was visibly touched, saying: "Thank you for this amazing gesture. I'm humbled by it, but I will treasure it."
06.02.2026 14:28 — 👍 20 🔁 8 💬 0 📌 0
Just out @natrevimmunol.nature.com
RNA-binding proteins and ribonucleoproteins as determinants of immunity
www.nature.com/articles/s41...
gives our perspective on this growing area of research. Special thanks to the editor Alexandra, for streamlining the manuscript.
an interesting work on how RNA regulatory elements control immune systems.
24.01.2026 11:17 — 👍 2 🔁 0 💬 0 📌 0
differential contribution of H3K9 methyltransferases to boundaries at satellites
A new and fascinating story from @bencarty.bsky.social and the group, with crucial help from the teams of @naltemose.bsky.social, Simona Giunta, and @dfachinetti.bsky.social. Many thanks to all for a fantastic collaboration.
www.nature.com/articles/s41...

I’m a fair-weather poster on here to say the least. However, I'm pleased to present our latest paper. A great effort from all authors and an interesting insight into transcriptional regulation (we think!).
www.cell.com/molecular-ce...

⚠️ Paper alert: Using a novel CRISPR screening approach, we mapped the entire regulatory network controlling Xist—key for X-chromosome inactivation.
👉 We discover how sex and development signals are decoded at a single gene locus.
www.nature.com/articles/s41...
👇 Bluetorial
Thanks Joe.
09.09.2025 13:24 — 👍 0 🔁 0 💬 0 📌 0
Thrilled to share our new Brockdorff lab paper at NSMB: we show that the primary role of m6A RNA modification on Xist is to promote RNA degradation. Xist turnover is mediated by the NEXT complex, not PAXT, and is independent of the m6A reader YTHDC1. www.nature.com/articles/s41...
09.09.2025 10:49 — 👍 13 🔁 0 💬 2 📌 2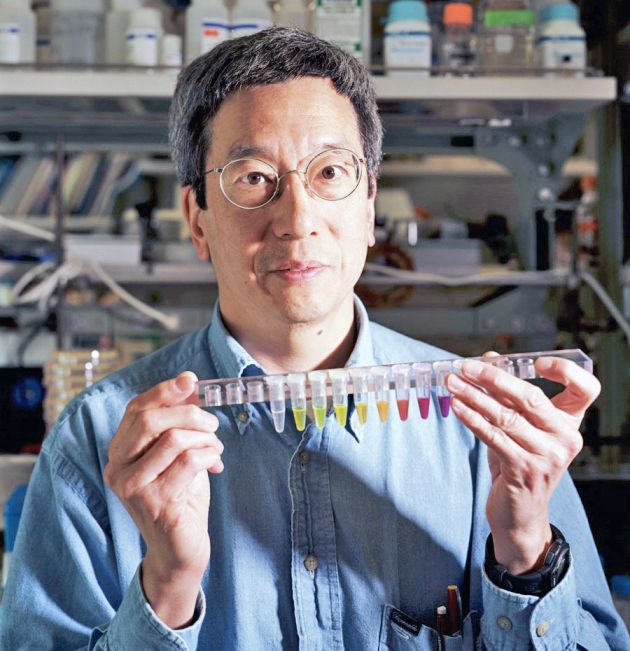



Today, I would like to honor the memory of Roger Y. Tsien, who died on August 24, 2016. His legacy lives with all who use his technologies, including calcium sensors, fluorescent proteins, the acetoxymethyl (AM) ester, & many more! #FluorescenceFriday
www.nature.com/articles/nme...
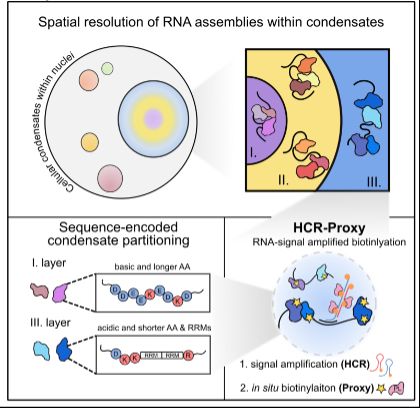
New method drop:
HCR-Proxy, a modular proximity labelling approach to profile local proteome composition around RNA at nanoscale, subcompartmental resolution. Thereby we resolved nested nucleolar layers and uncovered the grammar of spatial protein partitioning.
www.biorxiv.org/content/10.1...
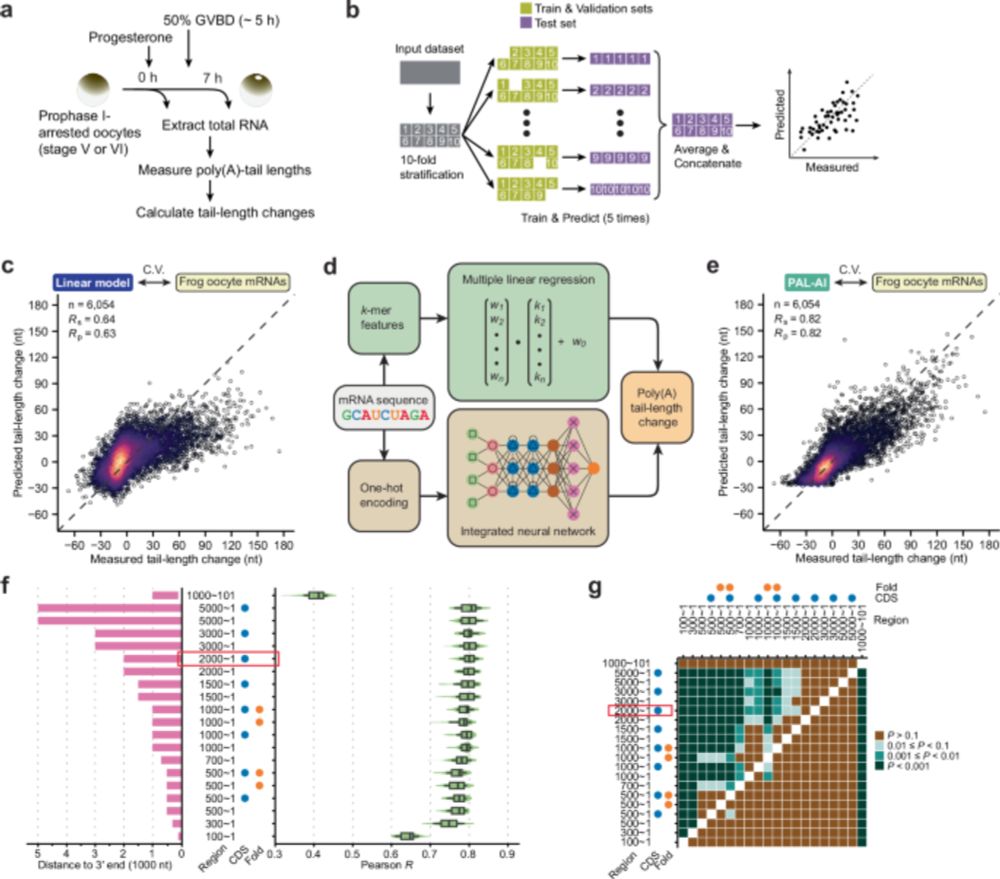
Check out the latest study from our lab, led by Coffee Xiang (@coffeebond007.bsky.social) www.nature.com/articles/s41... (1/2)
02.08.2025 14:59 — 👍 32 🔁 14 💬 1 📌 0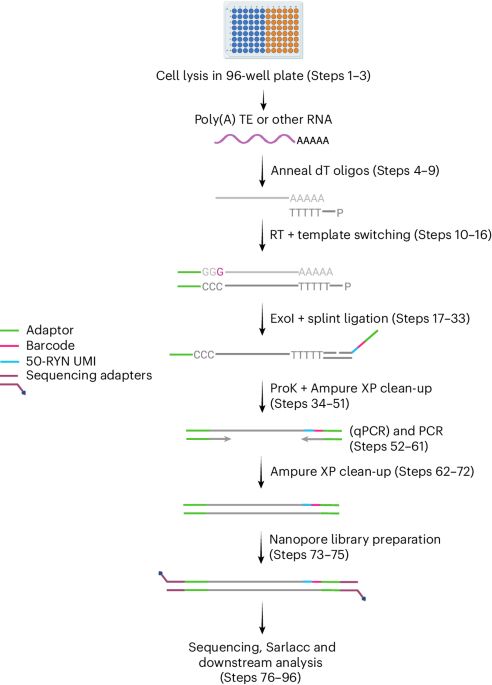
Very happy to share our protocols paper for CELLO-seq. This will make single cell long read RNA-seq more accessible and provides analysis guidelines. We hope this helps the #transposon #TEsky community and folks working on #singleCell isoform and allelic #gene expression. doi.org/10.1038/s415...
16.07.2025 16:55 — 👍 105 🔁 38 💬 8 📌 1
Fresh preprint by @flavia-con.bsky.social from our lab uncovers how SMCHD1 finds & binds chromatin using live-cell single-molecule imaging🔬
She reveals how SMCHD1 dynamically engages chromatin, including the inactive X chromosome, to maintain gene silencing.
www.biorxiv.org/content/10.1...
Excited to share our new paper on Nature - how Ku accommodates Alu expansion in primates by binding to dsRNA, providing a clue for both the high levels of Ku and its essentiality in human cells. Thank @chaolinzhang @hchung03 @LenaSteckelberg More to come www.nature.com/articles/s41...
16.05.2025 23:11 — 👍 49 🔁 25 💬 1 📌 1
An emotional day today @oxfordbiochemistry.bsky.social where leaders in the cell cycle field gathered to celebrate the life of Bela Novak and the huge impact he had on their work! www.bioch.ox.ac.uk/article/prof...
09.05.2025 15:36 — 👍 18 🔁 7 💬 0 📌 1Long-range massively parallel reporter assay reveals rules of distal enhancer-promoter interactions
www.biorxiv.org/content/10.1...
I can’t believe this. He is so kind and friendly.
25.02.2025 14:54 — 👍 1 🔁 0 💬 0 📌 0
Prime editing is precise and flexible, but it can require a lot of optimization. This post will give you some tips to get started making your prime edit as effective as possible!
blog.addgene.org/des...

I am thrilled to share our latest work, released today 🤩
www-nature-com.insb.bib.cnrs.fr/articles/s41...
It results from years of hard work by the team and long-time collaborators, with special thanks to PhD student Clara Roidor and collaborator Laurène Syx.
Preprint alert. Excited to share our latest work. We demonstrate that the primary function of m6A on Xist is to promote RNA degradation. www.biorxiv.org/content/10.1...
10.01.2025 17:58 — 👍 4 🔁 1 💬 0 📌 1Here it is! Congratuations to @jesskelley.bsky.social @edimitrova.bsky.social @neilblackledge.bsky.social @aleksszczurek.bsky.social and the rest of the team (Maciej, Amy, and Hieu). This was a fab collaboration with Arminja Kettenbach. Also see a lovely complementary paper from Fei Chen's group.
26.11.2024 17:03 — 👍 100 🔁 36 💬 8 📌 6Great thread on our paper from Jan👇
22.11.2024 18:39 — 👍 13 🔁 5 💬 0 📌 0
(1st post @BlueSky) Preprint alert🚨a long thread. Cautions in the use of @nanopore sequencing to map DNA modifications: officially reported “accuracy” ≠ reliable mapping in real applications. We performed a critical assessment of nanopore sequencing (across different versions of models) for the 1/n
20.11.2024 11:41 — 👍 115 🔁 44 💬 6 📌 10I've been meaning to write a lengthy thread about our experiences with DNA methylation editing in mouse ESCs if that would be useful (or interesting?) for a niche audience here. tl;dr: it was frustrating but we learned a lot, and it ended with our shiny new pub (1/17) www.nature.com/articles/s41...
12.11.2024 21:38 — 👍 129 🔁 51 💬 7 📌 4