📽️ G&D Tapes 📽️
G&D Author, Marion Solbach Mouginot tells us about their recent study in #genesdev describing how meta-domains control neuronal gene expression.
Learn more here:
➡️ bit.ly/45wGi5M
#TAD #chromatin #transcription #neuron #drosophila #3Dgenome
14.08.2025 19:13 — 👍 2 🔁 2 💬 0 📌 0

Join us in person or online for the 24th edition of the Gene Regulation Workshop at Uni Lausanne on Sept 1st, with an again fantastic group of speakers 🥳
Edouard Bertrand
Antonio Giraldez
Phillip Grote @phillipgrote.bsky.social
Susan Mango
Danny Nedialkova
Lucia Strader @luciastrader.bsky.social
04.08.2025 11:28 — 👍 17 🔁 8 💬 0 📌 0
Beautiful 😎
06.06.2025 09:49 — 👍 1 🔁 0 💬 0 📌 0
This work was led by Marion Mouginot, who notably established a locus-specific proteomics approach that identified Lola-I as a meta-loop-associated factor,
together with Sahar Hani (supported by Arianna Ravera, UNIL’s image analysis expert!), who uncovered hierarchical meta-loop formation. 👏
18.04.2025 18:50 — 👍 1 🔁 0 💬 0 📌 0

Lola-I was, however, only required to form a single meta-domain. This highlights a dichotomy between factors like Lola-I, which may control neuronal fate largely independently of inter-TAD interactions, and structural proteins like Cp190, which may enable ultra-long range gene regulation.
18.04.2025 18:50 — 👍 0 🔁 0 💬 1 📌 0

We developed an unbiased proteomics approach (inspired by Baldi..Becker 2018) to identify proteins bound to meta-loop anchors. This identified a specific isoform of the axon guidance protein Longitudinals lacking (Lola-I) as enriched at several meta-loop anchors, in our pull-downs and in vivo.
18.04.2025 18:50 — 👍 1 🔁 0 💬 1 📌 0
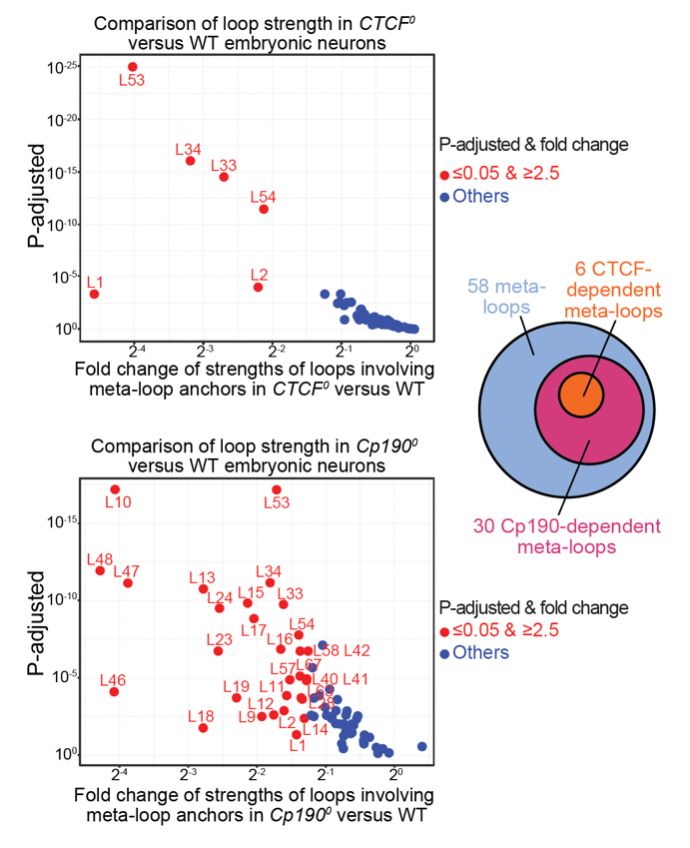
Many meta-loop anchors coincide with TAD boundaries. We found that a major determinant of TAD boundary formation in flies (called Cp190) is required to form a large fraction of meta-loops, including CTCF-dependent meta-loops.
18.04.2025 18:50 — 👍 1 🔁 0 💬 1 📌 0

How do meta-loops form?
Most meta-domains contain 2 or more meta-loops, with some of these forming earlier during neurogenesis than the other(s) in the same meta-domain.
We found that in many meta-domains, specific meta-loops (often the earlier forming ones) nucleate others.
18.04.2025 18:50 — 👍 0 🔁 0 💬 1 📌 0

1 meta-domain we studied contains a meta-loop connecting promoters of Glutamate receptors 1A and 1B. Disrupting the meta-domain uncoupled their co-transcription (similar to Levo..Levine 2022 for intra-TAD loops connecting paralogous gene promoters).
18.04.2025 18:50 — 👍 0 🔁 0 💬 1 📌 0
We precisely disrupted 3 independent meta-domains and found transcriptional defects in all 3 cases.
2 meta-domains we studied contain a meta-loop connecting a neuronal gene promoter to a distant intergenic element. Disrupting the meta-domain decreased neuronal gene transcription by 2-to-3-fold.
18.04.2025 18:50 — 👍 0 🔁 0 💬 1 📌 0

We recently showed that in fly neurons, specific pairs of TADs megabases apart specifically come together to form meta-domains (Mohana et al. 2023).
Within meta-domains, meta-loops connect accessible chromatin peaks present at promoters of specific neuronal genes or in non-coding intergenic DNA.
18.04.2025 18:50 — 👍 0 🔁 0 💬 1 📌 0
Sign In
A biweekly scientific journal publishing high-quality research in molecular biology and genetics, cancer biology, biochemistry, and related fields
Our recent paper reports extremely long-range (up to 5.1 Mb 😲) regulation of certain genes in fly neurons and provides new insights into how they form (see short thread below 👇).
genesdev.cshlp.org/gca?gca=gene...
18.04.2025 18:50 — 👍 35 🔁 11 💬 1 📌 0
and thank you to the reviewers whose feedback greatly improved the review 😎
13.02.2025 12:27 — 👍 0 🔁 0 💬 0 📌 0
Fun to do this with talented Sanyami Zunjarrao in our lab!
Thank you @andersshansen.bsky.social and Marcelo Nollmann for including our review in the 2025 Genome Architecture & Expression edition of COGEDE 😀
13.02.2025 12:26 — 👍 1 🔁 0 💬 1 📌 0

We highlighted new research from the past two years on examples of long-range gene regulation - within large TADs, across TADs, and across chromosomes - comparing mechanisms in flies and mammals 🪰❤️🐁.
www.sciencedirect.com/science/arti...
13.02.2025 12:22 — 👍 29 🔁 8 💬 1 📌 0
🛑CANCELLED—“Research on Women’s Health” funding website of the NIH.
01.02.2025 05:23 — 👍 2407 🔁 1348 💬 99 📌 109
yes indeed, BEAF-32 mat- zyg- mutants are not happy...
15.11.2024 14:03 — 👍 1 🔁 0 💬 0 📌 0
Thank you to the outstanding efforts of notably Anastasiia Tonelli and Pascal Cousin and another great collaboration with Aleksander Jankowski @ajank@mastodon.online 💪
15.11.2024 13:59 — 👍 0 🔁 0 💬 0 📌 0

Exciting findings using insulator-seq are:
Not all insulator-binding protein sites act as enhancer-blockers.
Not all TAD boundaries act as enhancer-blockers.
Both a CTCF motif and its surrounding sequence context are essential for enhancer-blocking.
15.11.2024 13:59 — 👍 0 🔁 0 💬 1 📌 0

(2) The insulator activity of CTCF binding sites depends on an intact CTCF motif.
15.11.2024 13:59 — 👍 0 🔁 0 💬 1 📌 0

Here’s some proof that insulator-seq works:
(1) It identifies biologically meaningful insulators in genomic DNA (both known and novel).
15.11.2024 13:59 — 👍 0 🔁 0 💬 1 📌 0
That’s right, insulators works on transiently transfected plasmids! This allows us to test the enhancer-blocking activities of up to 1 million DNA test fragments at a time.
15.11.2024 13:59 — 👍 0 🔁 0 💬 1 📌 0

Anastasiia Tonelli in the lab established insulator-seq - a massively parallel reporter assay for enhancer-blockers in Drosophila cells
15.11.2024 13:59 — 👍 0 🔁 0 💬 1 📌 0
G&D publishes high-quality research in molecular biology, cancer biology, development, neuroscience and related fields. In addition, G&D also publishes comprehensive Review Articles, Perspectives and Outlooks written by leading scientists.
Plant Biologist at the Salk Institute
Chromatin and transcription aficionado. Postdoc in Bickmore lab.
Asst Prof at University of California, Irvine.
Genetics, Genomics, Gene Regulation, Development. Views are my own.
https://www.kvonlab.org/
Postdoc - Duboule Lab - @college-de-france.fr @cirbcdf.bsky.social
Formerly PhD student - Furlong Lab - @EMBL.org
Interested in gene expression and genome topology regulation during embryogenesis
From 🇮🇹 , then Heidelberg 🇩🇪, now Paris 🇫🇷
Nuclear cell biologist with a passion for super-resolution imaging
We study transcriptional regulation and chromosome folding using an interdisciplinary approach combining wet- and dry-lab methods.
https://giorgettilab.org
@fmiscience.bsky.social
Scientist — Biophysicist
https://tglab.princeton.edu/
https://research.pasteur.fr/en/team/physics-of-biological-functions/
Postdoc @UZH. Interested in Genome stability, Cellular heterogeneity & Imaging.
Senior editor @naturesmb.bsky.social. Views and opinions are personal.
We study developmental epigenetics using fruit flies
https://www.su.se/english/research/research-groups/group-mannervik
Institute for Quantitative Biosciences, UTokyo. Mechanism of transcriptional regulation in the early embryo.
https://sites.google.com/view/fukaya-lab-iqb/home
Group leader at iGReD Clermont-Ferrand, drosophila & epigenetics, food & mushrooms
https://www.igred.fr/en/team/ero-epigenetic-regulations-ontogenesis/
Biophysicist & computational biologist | Genome organization, chromatin modeling
Postdoc at Institut Curie, Paris (Leonid Mirny group) | Formerly at IMBA, Vienna (Anton Goloborodko group)
Evolution, ecology & biological clocks - with a focus on lunar rhythms
in development & reproduction of the marine insect Clunio.
+ Genomics | Biodiversity | Behaviour | NeuroBio | MolBio | SciCom
bit.ly/KaiserLab
@mpi-evolbio.bsky.social @maxplanck.de
Sensory biology, genetics, behavior, and evolution (especially in the Twilight Zone) | Incoming Assistant Professor, NJIT Biological Sciences (Fall '26) | Current postdoc, University of Lausanne, Benton Group
https://himmellab.org
👩🏻🔬 Geneticist and molecular biologist
Single molecule enthusiast 🔍
EMBO postdoctoral fellow @CIG Lausanne🇨🇭
Previously BioSPC PhD student @IJM Paris🇨🇵
Tenure-Tracking @ NICHD_NIH
#gene_regulation #nuclear_organization #development | handkerchief user | views are mine alone, probably for the best
rochalab.nichd.nih.gov
Multiscale Genome Organisation subgroup of the Biophysical Society https://www.biophysics.org/subgroups/multiscale-genome-organization














