Laura Lorenzo Orts
IMB Mainz
I am excited to announce that I will be moving to IMB Mainz next year! The Winter call for the IPP PhD program is now open; if you are interested in maternal #mRNA regulation and #translation in early vertebrate development, please apply! Deadline: 16 October.
More info: www.imb.de/students-pos...
15.09.2025 12:13 — 👍 60 🔁 24 💬 2 📌 3
Love RNA biology?
Join us to explore the piRNA pathway with structural and genetic approaches (see 👇👇).
PhD student/postdoc position co-supervised by Clemens Plaschka & myself.
DM or email us if you’d like to know more!
@vbcscitraining.bsky.social @imbavienna.bsky.social @impvienna.bsky.social
18.09.2025 14:34 — 👍 18 🔁 20 💬 0 📌 1
My first first-author paper is out!🎉
Here we propose a model where a silencing complex, PIWI*, assembles on target RNAs to recruit effectors and shut down transposon activity.
Huge thanks to the Brennecke and Plaschka labs, especially Julius and Clemens, and all co-authors!
17.09.2025 13:00 — 👍 48 🔁 20 💬 3 📌 1

PIWI clade Argonautes are essential for transposon silencing. Without them, animals are sterile due to massive transposon activity.
But how does piRNA-guided target interaction translate into silencing?
PhD student Júlia Portell Montserrat has an intriguing answer
www.cell.com/molecular-ce...
17.09.2025 10:38 — 👍 89 🔁 41 💬 2 📌 4
Close inter-lab collaborations with shared PhD students and postdocs are the future!
Julia is the hero of this work, she is currently looking for postdoc labs …
09.09.2025 10:59 — 👍 17 🔁 4 💬 1 📌 0
New paper by our Plaschka lab with Brennecke’s lab at IMBA!
Researchers at the Vienna BioCenter have solved a 20-year-old mystery in genome biology, revealing how PIWI proteins engage partner molecules to silence “jumping genes” that threaten genetic stability.
08.09.2025 09:10 — 👍 12 🔁 5 💬 1 📌 1
So happy to see the PhD work of Emilio Santillán published—what an amazingly elegant piece of work!!
Working with him and Luisa at the IMP was a great and formative experience.
03.09.2025 19:46 — 👍 1 🔁 0 💬 1 📌 0
1/ How do animals develop immunity against a newly encountered transposable element from scratch? Our study reveals that the mobility of TEs is their Achilles heel, allowing hosts to develop a powerful small RNA-mediated silencing response.
www.biorxiv.org/content/10.1...
14.08.2025 17:09 — 👍 47 🔁 26 💬 4 📌 2
My colleagues Baptise Rafanel et al. investigated how a newly invading transposon can be silenced. They found that not only antisense insertions in piRNA clusters such as flamenco, but also insertions in the 3‘UTR of genes can drive potent silencing.
02.08.2025 20:36 — 👍 21 🔁 9 💬 0 📌 0
Pls. share widely
Calling all transposon fans & lovers of genetic innovation
MOBILE GENOME welcomes you in Heidelberg, Nov. 4–7 2025
→ Vibrant & friendly community
→ Cutting-edge talks from mechanisms to physiology
→ Plenty of surprises (TEs never stop innovating)
submit abstract by July 29
16.07.2025 08:43 — 👍 61 🔁 48 💬 1 📌 1
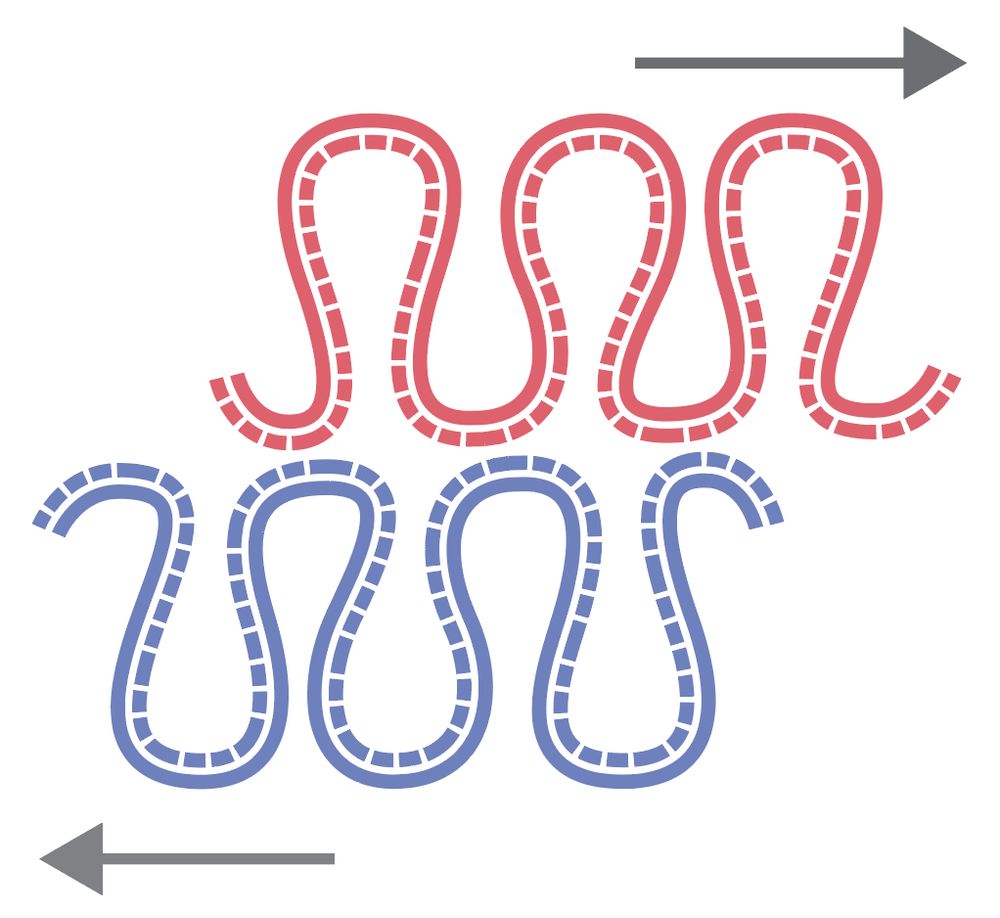
We found a new asymmetry in the large-scale chromosome structure: sister chromatids are systematically shifted by hundreds of kb in the 5′→3′ direction of their inherited strands! The work was led by Flavia Corsi, in close collaboration with the Daniel Gerlich lab.
www.biorxiv.org/content/10.1...
1/
15.07.2025 08:11 — 👍 111 🔁 58 💬 3 📌 7
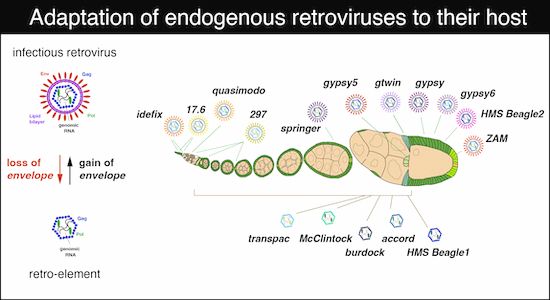
The fascinating work spearheaded by Kirsten Senti on the diversification and adaptation of endogenous retroviruses in the host gonad 'ecosystem'' is officially published.
17.06.2025 20:34 — 👍 44 🔁 13 💬 1 📌 1
I freaking 😍 this paper. The ecology of the genome is one thing. The ecology of the Drosophila ovary another.
19.06.2025 15:26 — 👍 7 🔁 1 💬 0 📌 0

🚨📢 New paper alert!
A study led by Kirsten Senti and Julius Brennecke offers new insights into how ancient endogenous retroviruses diversified to exploit different cell “niches” in the fruit fly ovary, and how the host’s defenses adapted in return. https://imba.science/Brennecke_EMBOJ
16.06.2025 09:40 — 👍 28 🔁 10 💬 0 📌 2
REFEREE 1: “I conclude with a simple, direct statement - this is the best paper I have read all year!”
#TransparentPeerReview
Co-evolving infectivity and expression patterns drive the diversification of endogenous retroviruses
@juliusbrennecke.bsky.social et al
www.embopress.org/doi/full/10....
06.06.2025 14:18 — 👍 102 🔁 33 💬 3 📌 2
The @viennabiocenter.bsky.social PostDoc Program call is now open! We have 16 fully-funded positions for scientists with a PhD in biology, chemistry, physics, bioinformatics etc.
Apply here: training.vbc.ac.at/post-docs/vi...
#VBCPostDoc
02.06.2025 11:28 — 👍 36 🔁 22 💬 2 📌 1
Link to Peter‘s thread:
bsky.app/profile/germ...
25.04.2025 11:16 — 👍 0 🔁 0 💬 0 📌 0
Highly interesting work from Riedelbauch et al. from Peter Andersen‘s Lab @germline.bsky.social
25.04.2025 11:16 — 👍 3 🔁 0 💬 2 📌 0
YouTube video by Johan Gibcus
Rules of Engagement
Earnshaw, Goloborodko, Dekker & Mirny labs are excited to present our latest work, "Rules of engagement for condensins and cohesins guide mitotic chromosome formation" - now accepted!!
www.science.org/doi/10.1126/...
A short clip describing the key results:
www.youtube.com/watch?v=pmvO...
11.04.2025 07:45 — 👍 98 🔁 53 💬 7 📌 6
A thread from the Postdoc who led the work on bacterial Telomeric transposons. A wonderful collaboration with the Barabas lab. Officially out in the journal today!
27.03.2025 19:55 — 👍 15 🔁 2 💬 1 📌 1

IMBA is recruiting a Junior Group Leader! Are you interested in starting your own lab, pursuing curiosity-driven basic research in the life sciences? Apply now to our group leader position. The deadline is May 28. Link is below.
28.03.2025 11:15 — 👍 67 🔁 75 💬 1 📌 4
1. For the past thirty years I've had the best job in the world.
I've had the opportunity to follow my curiosity; explore the workings of nature and society; mentor students and junior colleagues in the same process; and teach generations of students about it all.
19.03.2025 19:32 — 👍 2602 🔁 937 💬 38 📌 238
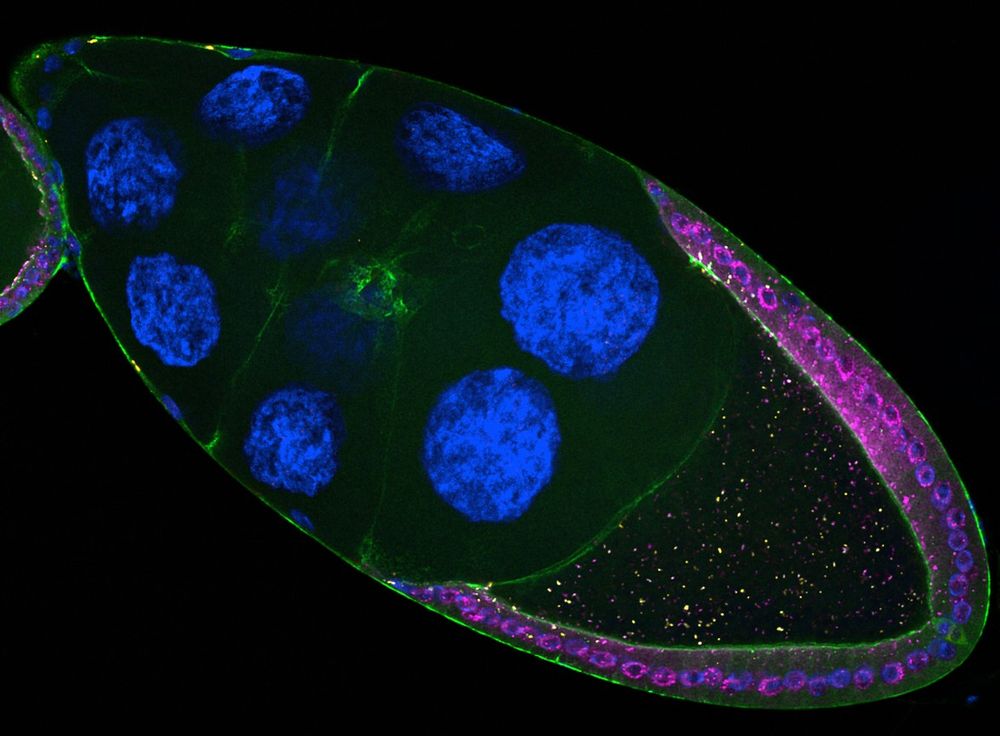
Drosophila follicle showing retrotransposons (pink & yellow) expressed in somatic cells infecting the oocyte
1/ Transposable elements are often called "jumping genes" because they mobilize within genomes. 🧬
But did you know they can also jump 𝘣𝘦𝘵𝘸𝘦𝘦𝘯 cells? 🤯
Our new study reveals how retrotransposons invade the germline directly from somatic cells.
www.biorxiv.org/content/10.1...
A short thread 🧵👇
17.03.2025 11:56 — 👍 544 🔁 259 💬 11 📌 33
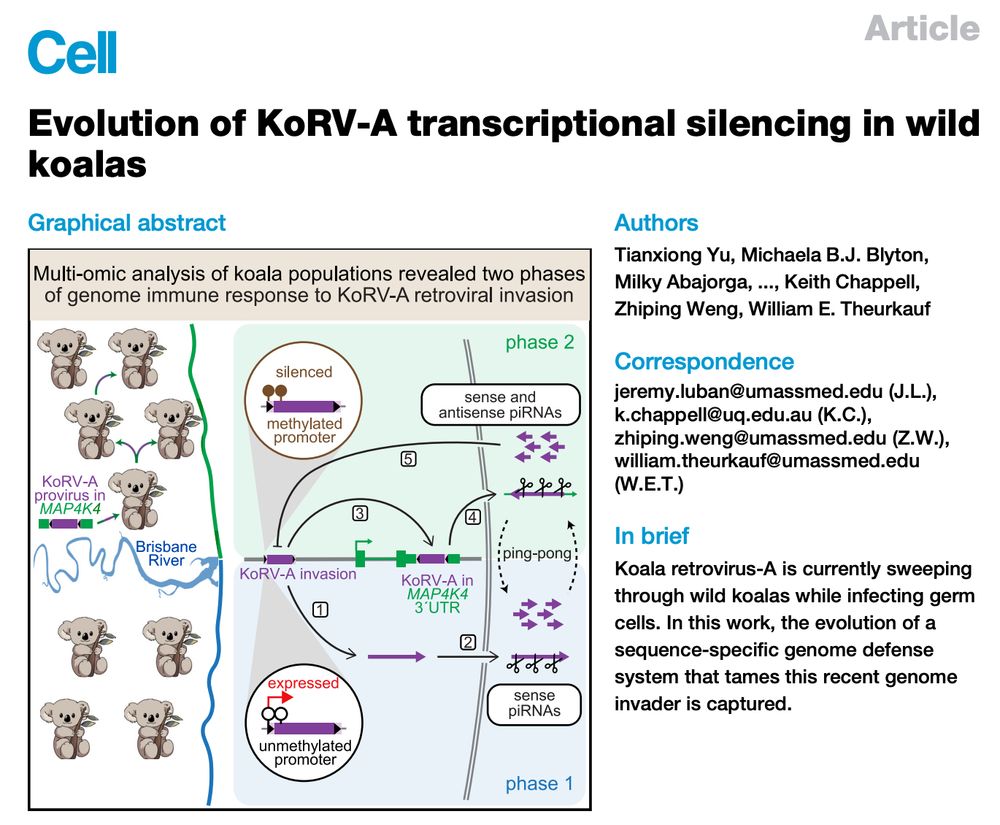
I am thrilled to share our story online at @cellpress.bsky.social . Big thanks to all authors: Zhiping, Bill, @lubanlab.bsky.social, Keith & bluesky-less!
authors.elsevier.com/c/1kjdaL7PXu...
How to tame a genome invader? It takes wild koalas 🐨🐨 to learn it.
#Retrovirus #koala #piRNA
More below 👇
09.03.2025 02:35 — 👍 43 🔁 24 💬 3 📌 3
"Scientists have to organize, they have to protest, they have to make their case to the public. They must never give in." Quote from Paul Nurse (2001 Nobel) from upcoming interview in @annualreviews.bsky.social with him and Bruce Alberts (former President NAS). @standupforscience.bsky.social today
07.03.2025 10:25 — 👍 31 🔁 15 💬 2 📌 0
Beautiful work on the copia retrotransposon by Sven Klumpe (ex-MPI, soon IMBA/IMP) and Kirsten Senti from the Brennecke Lab!
05.03.2025 16:46 — 👍 2 🔁 0 💬 0 📌 0
- EMBL with EMBL outstations
- Max Planck institutes in Germany (e.g. Dresden, Goettingen)
- Vienna BioCenter (IMP, IMBA, GMI, Max Perutz Labs)
- FMI Basel
- ETH Zürich
- CEITEC Brno
- CRG Barcelona
- Hubrecht Institute (Netherlands)
- EPFL Lausanne
01.03.2025 08:44 — 👍 17 🔁 7 💬 6 📌 1
Fascinated about the molecular mechanisms of evolutionary arms races and fast evolving sequences.
Hobby enthusiasts of non-model organisms.
Developmental neurobiologist using cerebral #organoids to study brain size and evolution. Opinions my own.
mRNA & cryo-EM enthusiasts at IMP Vienna. Posts are by lab members.
scientist | pacifist | transposable elements | evo-devo
PhD @trono-lab.bsky.social
Shared PostDoc in Veit Hornung’s lab and Fabian Theis’ lab in Munich. Let‘s go.
PhD student at Aarhus University, Denmark
IMB, Mainz
PhD student studying Argonaute biology in Rene Ketting Lab
Assistant Professor @ Stanford Genetics & BASE Initiative. Mapping the regulatory code of the human genome to understand heart development and disease. www.engreitzlab.org
microRNAs, omics and computational biology. Associate professor at Stockholm University and SciLifeLab
www.friedlanderlab.org
PhD candidate, Bartel Lab @bartellab.bsky.social, MIT/Whitehead Institute. Biochemical + biophysical principles of protein/RNA mechanisms. MicroRNA/siRNAs & AGO proteins.
PhD student at the Brennecke and Plaschka labs, Vienna.
Interested in RNA silencing
Structural biologist interested in all things green. Professor at University of Geneva, Switzerland. https://web.structplantbio.org/
Scientist at IMP in Vienna. Excited about gene expression regulation and its encoding in our genomes - enhancers, transcription factors, co-factors, silencers, AI.
#Mechanics of #zebrafish, #ascidian, and #drosophila #development @Institute of Science and Technology Austria (ISTA)
https://heisenberglab.pages.ist.ac.at
Incoming group leader @FMI, Basel
Postdoc @MPI-MG, Berlin | PhD @IMBA, Vienna
Investigating cell fate and loss in tissue dynamics
https://www.fmi.ch/research-groups/groupleader.html?group=150
PhD candidate in the Kvon Lab @ UC Irvine 👩🏻🔬🧬
Genomics biologist. Group leader at MRC Laboratory of Medical Sciences, Hon. Senior Lecturer (Associate Prof.) at Imperial College London.
Opinions / view my own.
https://functionalgenecontrol.group











