
New preprint from the Trcek lab, with Siran Tian as the lead author - congratulations Siran! doi.org/10.64898/202...
26.12.2025 10:41 — 👍 6 🔁 2 💬 1 📌 0@trceklab.bsky.social

New preprint from the Trcek lab, with Siran Tian as the lead author - congratulations Siran! doi.org/10.64898/202...
26.12.2025 10:41 — 👍 6 🔁 2 💬 1 📌 04) nanos mRNA localization counteracts Vasa-mediated enhancement of Oskar mobility. While germ granule mRNA localization requires Vasa, this process opposes Vasa’s effect on Oskar condensates, providing an RNA-mediated negative feedback mechanism that regulates condensate properties.
26.12.2025 10:41 — 👍 0 🔁 0 💬 0 📌 03) mRNA localization to Oskar condensates requires Vasa and its interaction with Oskar. Although germ granules contain diverse proteins, Vasa and Oskar are necessary and sufficient for robust germ granule mRNA localization to the granules.
26.12.2025 10:41 — 👍 0 🔁 0 💬 1 📌 02) Vasa’s modulation of condensate properties requires direct protein-protein interaction. This activity requires specific binding interfaces between Vasa and Oskar, but is less dependent on Vasa helicase activity.
26.12.2025 10:41 — 👍 0 🔁 0 💬 1 📌 01) Vasa modulates the material properties of ribonucleoprotein condensates in fly oocytes and human cells. Recruitment of Vasa increases the mobility of the germ-granule nucleator Oskar, thereby preventing aggregation of granules.
26.12.2025 10:41 — 👍 0 🔁 0 💬 1 📌 0This new function in regulating protein condensation extends beyond its established function as an RNA helicase. Specifically, Siran shows that:
26.12.2025 10:41 — 👍 0 🔁 0 💬 1 📌 0She investigated the role of a conserved DEAD-box RNA helicase Vasa in condensation of germ granules in Drosophila. Her work reveals a previously unrecognized function of Vasa in regulating the material properties of Oskar ribonucleoprotein condensates through its direct interaction with Oskar.
26.12.2025 10:41 — 👍 0 🔁 0 💬 1 📌 0
New preprint from the Trcek lab, with Siran Tian as the lead author - congratulations Siran! doi.org/10.64898/202...
26.12.2025 10:41 — 👍 6 🔁 2 💬 1 📌 04) Germ granule mRNAs are replete with GC-rich complementary sequences predicted to be competent of engaging in persistent intermolecular base pairing however in vivo these interactions are unrealized and restricted by RNA folding.
30.08.2025 23:14 — 👍 1 🔁 0 💬 0 📌 03) Engineered germ granule mRNAs with exposed GC-rich complementary sequences (CSs) within stem loops induce sustained base pairing in vitro and enhanced intermolecular interactions in vivo. But, the presence of these structures disrupts development, and GC-rich CSs exacerbate this phenotype.
30.08.2025 23:14 — 👍 0 🔁 0 💬 1 📌 0We find that:
1) mRNAs remain structured within germ granules
2) mRNAs base pair intermolecularly without high sequence complementarity or significant melting of secondary structure. Base pairing is driven by scattered and discontinuous stretches of bases appearing on the surface of structured RNAs

New paper from the Trcek lab, with Siran Tian as the lead author - congratulations Siran!: www.nature.com/articles/s41...
30.08.2025 23:13 — 👍 18 🔁 8 💬 1 📌 2
New preprint from the Trcek lab - Jeffrey Liao as the lead author! We studied intermolecular base pairing using AMT crosslinking in mammalian cells during oxidative stress doi.org/10.1101/2025...
28.05.2025 11:43 — 👍 3 🔁 1 💬 1 📌 0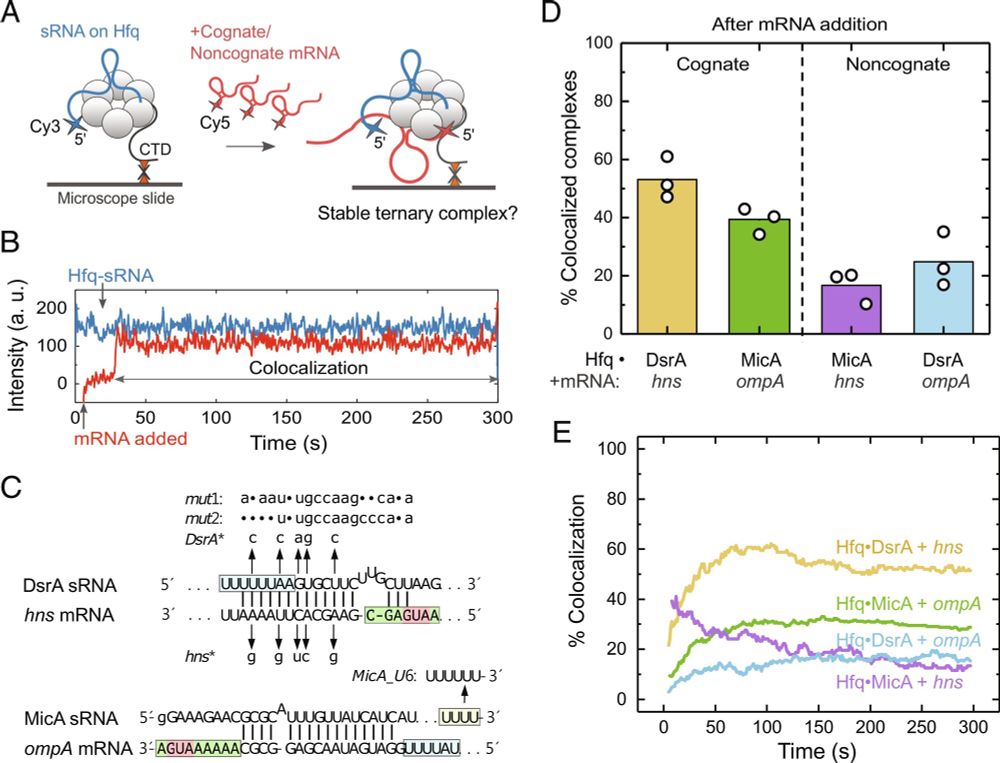
Our new paper is out in PNAS! We show that competition for binding to the Hfq matchmaker disfavors mismatched sRNA-mRNA combinations while allowing matched sRNA-mRNAs to form stable pairs. Kudos to Jorjethe Roca and Yi-Lan Chen for their fantastic work on this problem! doi.org/10.1073/pnas...
19.07.2025 00:36 — 👍 7 🔁 2 💬 0 📌 1Congrats, Jorjethe and Yi-Lan! 👏👏
19.07.2025 10:35 — 👍 1 🔁 1 💬 0 📌 0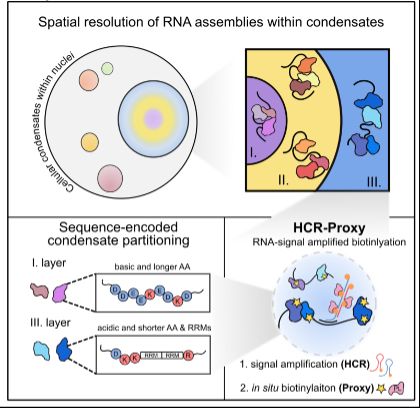
New method drop:
HCR-Proxy, a modular proximity labelling approach to profile local proteome composition around RNA at nanoscale, subcompartmental resolution. Thereby we resolved nested nucleolar layers and uncovered the grammar of spatial protein partitioning.
www.biorxiv.org/content/10.1...

Excited to share our lab's preprint! We found that flexible RNAs have much broadened ion atmospheres than structured RNAs. This leads us to speculate the implications on molecular recognition and phase behavior. Please check full text for other interesting details www.biorxiv.org/content/10.1...
28.05.2025 18:42 — 👍 4 🔁 2 💬 1 📌 0Jeffery is presenting his work at the annual RNA society meeting in San Diego - visit his poster! #RNA25; #RNA2025
28.05.2025 11:55 — 👍 6 🔁 0 💬 0 📌 0Jeffery is presenting his work at the annual RNA society meeting in San Diego - visit his poster! #RNA25; #RNA2025
28.05.2025 11:54 — 👍 1 🔁 0 💬 0 📌 0Characterization of intermolecular base pairing using AMT crosslinking in mammalian cells during oxidative stress https://www.biorxiv.org/content/10.1101/2025.05.21.655305v1
27.05.2025 08:19 — 👍 1 🔁 1 💬 0 📌 0Characterization of intermolecular base pairing using AMT crosslinking in mammalian cells during oxidative stress https://www.biorxiv.org/content/10.1101/2025.05.21.655305v1
27.05.2025 08:19 — 👍 1 🔁 1 💬 1 📌 0We propose that the combined suppression of intermolecular base pairing and the increased RNA structural diversity may serve as a mechanism to preserve normal RNA function during stress, while also facilitating the reversible assembly of stress granules
28.05.2025 11:43 — 👍 0 🔁 0 💬 1 📌 0c) However, these RNAs increase in structural diversity in a manner dependent on oxidative stress and the stress granule nucleating proteins G3BP1 and G3BP2.
d) In the absence of the main stress granule nucleating proteins G3BP1 and G3BP2, mRNAs remain dispersed in cells during oxidative stress
We show that:
a) Oxidative stress downregulates intermolecular base pairing of reporter mRNAs engineered to enhance detection of such interactions.
b) RNA regions within transcripts that enrich in stress granules remain structured during stress.

New preprint from the Trcek lab - Jeffrey Liao as the lead author! We studied intermolecular base pairing using AMT crosslinking in mammalian cells during oxidative stress doi.org/10.1101/2025...
28.05.2025 11:43 — 👍 3 🔁 1 💬 1 📌 0