Research Assistant - Tonkin-Hill Lab
Research Assistant - Tonkin-Hill Lab
It's a bit last minute, but we have an RA/junior postdoc position available in my lab (closing Sunday). Please reach out if you're interested in developing and applying methods to analyse microbial genomes and metagenomes! careers.petermac.org/job/MELBOURN...
20.02.2026 00:16 — 👍 4 🔁 7 💬 0 📌 0
At long last, my final PhD chapter is out: we developed a novel evolutionary simulator of bacterial pangenomes, Pansim, fitting it to data from >600K genomes using a likelihood-free framework, PopPUNK-mod, to explore neutral and adaptive pangenome dynamics www.biorxiv.org/content/10.6...
07.02.2026 10:08 — 👍 45 🔁 18 💬 2 📌 1
Proud to announce SimPhyNI, a new tool for bacterial GWAS with higher precision and scalability than existing tools. Try it out and let us know what you think!!
05.01.2026 14:55 — 👍 66 🔁 32 💬 5 📌 1

🧬 Come check out my poster on agtools, an open-source Python framework for analysing and manipulating assembly graphs at #ABACBS2025 Poster #106
•
💻 Github: github.com/Vini2/agtools
📄 Preprint: biorxiv.org/content/10.110…
25.11.2025 23:21 — 👍 27 🔁 15 💬 1 📌 0

PeterMac @petermaccc.bsky.social representing at #abacbs2025 What a community 😍
26.11.2025 01:03 — 👍 35 🔁 9 💬 0 📌 0
Happy to share our new AMR resource which has phenotypic AMR (usually MIC data) collected from publications and databases. This is paired with assemblies and annotations
We're excited for users who might train new models, find phenotype/genotype mismatches, or any other use
19.11.2025 12:27 — 👍 62 🔁 36 💬 1 📌 0

Logo of Bin Chicken (Australian white ibis) on a rubbish bin, pulling out a strand of DNA
“Bin Chicken” is now published in Nature Methods! It substantially improves genome recovery through rational coassembly 🧬🖥️. Applied to public 🌍 metagenomes, we recovered 24,000 novel species 🦠, including 6 new phyla.
doi.org/10.1038/s415...
@benjwoodcroft.bsky.social @rhysnewell.bsky.social
🧵1/6
13.11.2025 10:08 — 👍 73 🔁 38 💬 2 📌 4
Metalog
Metalog is a repository of manually annotated metadata (or contextual data) for metagenomic sequencing data from across the globe.
We're very happy to release our new database Metalog metalog.embl.de ! It offers manually curated and harmonised contextual data for 110k metagenomics samples across the globe, incl. precomputed taxonomic profiles, for interactive browsing and for download 🧵 1/7
#microsky
15.08.2025 08:07 — 👍 73 🔁 45 💬 3 📌 2

Delighted to see our paper studying the evolution of plasmids over the last 100 years, now out! Years of work by Adrian Cazares, also Nick Thomson @sangerinstitute.bsky.social - this version much improved over the preprint. Final version should be open access, apols.
Thread 1/n
25.09.2025 21:28 — 👍 299 🔁 154 💬 14 📌 8
Happy to see that Spacedust is now published on Nature Methods!
It combines sensitive Foldseek structure search and conserved neighborhood detection to discover functionally-associated gene clusters in prokaryotic & viral genomes.
1/6🧵
16.09.2025 20:26 — 👍 21 🔁 12 💬 1 📌 0

Efficient sequence alignment against millions of prokaryotic genomes with LexicMap - Nature Biotechnology
LexicMap uses a fixed set of probes to efficiently query gene sequences for fast and low-memory alignment.
Sometimes you meet absolutely incredible bioinfo-magicians.
It was a huge privilege when @shenwei356.bsky.social
joined our group for a year on an @embl.org sabbatical.
While here, he developed a new way of aligning to
millions of bacteria, called LexicMap 1/n
www.nature.com/articles/s41...
10.09.2025 09:12 — 👍 190 🔁 99 💬 5 📌 4

Large AI models are reported to achieve high accuracy (AUROC) predicting pathogenic variants across the genome.
A preprint reports that the predictions are based on splice variants. Using only this info (no sequences, no AI) achieves AUROC=0.944 across noncoding variants.
1/2
09.09.2025 23:01 — 👍 15 🔁 7 💬 1 📌 0

🌎👩🔬 For 15+ years biology has accumulated petabytes (million gigabytes) of🧬DNA sequencing data🧬 from the far reaches of our planet.🦠🍄🌵
Logan now democratizes efficient access to the world’s most comprehensive genetics dataset. Free and open.
doi.org/10.1101/2024...
03.09.2025 08:39 — 👍 218 🔁 118 💬 3 📌 16

The view of the Adelaide Oval, site of the 2025 Australian Biology and Computational Biology Society (ABACBS) conference, from across the River Torrens. The conference will be held in Adelaide from 24 to 28 November and celebrates 10 years of ABACBS.
The 10th anniversary #ABACBS conference will be held in Adelaide from Nov 24-28. If you're thinking of coming, get in quick, as it's going to be a big week: A fantastic #bioinformatics conference with outstanding national and international speakers, and we're wrapping up the ABACBS week an AC/DC gig
17.07.2025 05:43 — 👍 9 🔁 5 💬 0 📌 0

Logo for the Sandpiper website
Out in @natbiotech.nature.com: Metagenome taxonomy profilers usually ignore unknown species. SingleM is an accurate profiler which doesn't, even detecting phyla with no MAGs. Profiles of 700,000 metagenomes at sandpiper.qut.edu.au. A 🧵
16.07.2025 21:59 — 👍 131 🔁 71 💬 7 📌 9

ABACBS 2025 Conference
Adelaide, South Australia. Nov. 24-
The day is finally here! 🎉 We’re releasing the invited speaker line-up, key dates, and lots more info for ABACBS 2025.
Check it out and share widely: www.abacbs.org/abacbs2025
Registrations and abstract submissions open next week, with abstracts due in August!
04.07.2025 04:34 — 👍 8 🔁 6 💬 0 📌 1
FuGACI – JPIAMR
JPIAMR is a global collaborative organisation and platform, engaging 28 nations to curb antimicrobial resistance (AMR) with a One Health approach.
www.bath.ac.uk/jobs/Vacancy...
Post doc available to work on Candida genomics with me at the Milner Centre for Evolution (U Bath, UK) as part of the JPIAMR funded project Fugaci www.jpiamr.eu/projects/fug...
For queries, email me!
20.05.2025 12:39 — 👍 1 🔁 6 💬 0 📌 0
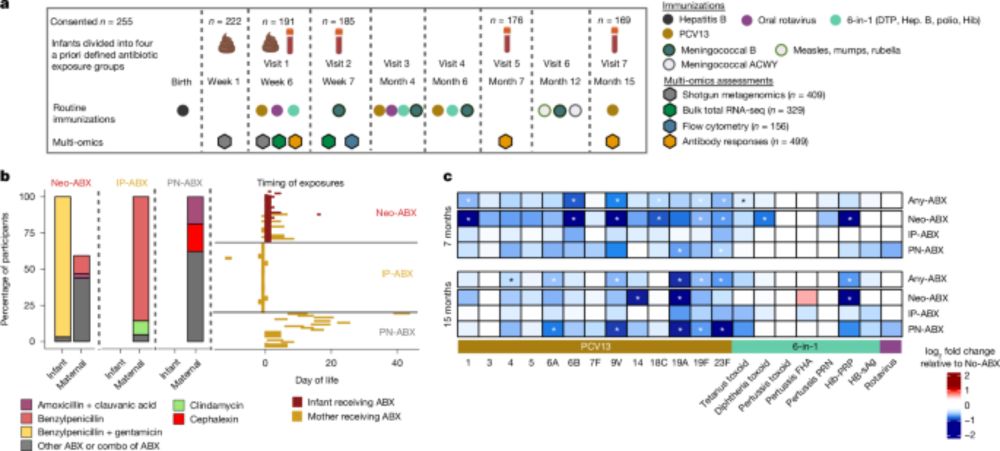
Bifidobacteria support optimal infant vaccine responses - Nature
Neonatal antibiotic use is shown to reduce immune response to infant vaccines, accompanied by reduced abundance of Bifidobacteria in the gut microbiota, with experiments in mice indicating that probio...
Delighted our study (7+ years in the making) is out today in Nature. In a nutshell, 48 hours of antibiotics in week 1 led to impaired vaccine responses up to 15 months later in infants. We show (in-vivo) a probiotic can fix it #microbiome #science #immunology
www.nature.com/articles/s41...
02.04.2025 23:33 — 👍 136 🔁 58 💬 7 📌 4
Being able to data plasmid acquisition, and to see the diversity of plasmids being gained and lost from lineages, was really eye opening! It was a pleasure to contribute my tiny bit to this huge paper and the data has so much more to give! @arredondo.bsky.social @gerrythill.bsky.social et al.
03.04.2025 11:34 — 👍 22 🔁 7 💬 0 📌 0
Group Leader
Group Leader
We are recruiting a group leader in laboratory research at Peter Mac Cancer Centre in
Melbourne, Australia 🔬🦘💥
Work in a cutting edge research environment in a vibrant world city!
We value research excellence, creativity and diversity.
Join us!
careers.petermac.org/job/MELBOURN...
05.03.2025 08:04 — 👍 30 🔁 34 💬 0 📌 5
Home
A tool for generating consensus long-read assemblies for bacterial genomes - rrwick/Autocycler
New year, new assemblies!
I'm excited to announce Autocycler, my new tool for consensus assembly of long-read bacterial genomes!
It's the successor to Trycycler, designed to be faster and less reliant on user intervention.
Check it out: github.com/rrwick/Autoc...
(1/5)
31.12.2024 23:43 — 👍 156 🔁 97 💬 2 📌 3
TNA documentation — TNA 0.0.1 documentation
I just made a first release of TNA - new tool to compare 2 bacterial genomes. Is shamelessly inspired by ACT, but should be simpler to use. Drag-n-drop two genomes: it runs BLAST for you and then you can visualize. For Mac, Windows 11, Linux tna.readthedocs.io
29.11.2024 11:38 — 👍 175 🔁 69 💬 9 📌 8
Associate Professor at Department of Biology, Aarhus University, Denmark 🇩🇰 working on microbial genomics, bioinformatics, cable bacteria, and other sediment microorganisms. https://www.au.dk/en/ianpgm@bio.au.dk/ 🇦🇺
Neurogeneticist interested in the relations between genes, brains, and minds. Author of INNATE (2018) and FREE AGENTS (2023)
Genome annotation geek at University of Greifswald, always interested in friendly scientific collaboration. Developer of BRAKER, GALBA, MakeHub and other Gaius-Augustus tools.
Solve Biology
Head of Generative Genomics, Wellcome Sanger Institute, Cambridge, UK
Systems + Synthetic Biology, CRG, Barcelona
http://barcelonacollaboratorium.com http://allox.bio https://www.sanger.ac.uk/programme/generative-and-synthetic-genomics/
Confess your sins anonymously - will the internet absolve you?
Buy show tickets 2025/6: sites.google.com/view/fesshole
Add confession b3ta.com/addfess
Buy book amazon.co.uk/s?k=very+best+of+fesshole&tag=b3ta-21
Run @robmanuel.b3ta.com
Bacterial genomes and whatnot. University of Bath.
Evolutionary genomics of bacteria
professor at Institut Pasteur. Bacterial genomics, vaccine-preventable diseases, antimicrobial resistance, bioinformatics and public heath applications. International capacity building and teaching. Klebsiella diphtheria whooping cough genomic librairies
Research Fellow @ Flinders University | #Bioinformatics, #Algorithms and #Metagenomics 🧬🦠 | Blog at http://vijini.medium.com
PhD Student in bioinformatics
PhD student | Tonkin-Hill Lab
Peter MacCallum Cancer Centre
PhD candidate at Peter Doherty Institute, University of Melbourne | interested in Pathogen Genomics | Antimicrobial Resistance | Public Health | One Health
Bioinformatician at Peter Mac, Melbourne.
Spatial 'omics w/ R, Python, Nextflow.
Official account of the Biosciences Area of @BerkeleyLab
Solving challenges in energy, environment, health, and biomanufacturing. Re-post ≠ Endorsement
Official account of Lawrence Berkeley National Laboratory (LBNL), a U.S. Department of Energy national lab. newscenter.lbl.gov
Director of the US Department of Energy Joint Genome Institute. Head of JGI’s Secondary Metabolites Science Program. Research interests are natural products, synthetic biology, microbiome
Posts about jobs, conferences, etc. from the Evolution Directory (EvolDir) mailing list https://evol.mcmaster.ca/evoldir.html, run by Brian Golding. This bot is run by @rdmpage.bsky.social. Problems: https://github.com/rdmpage/evoldir-bluesky/issues
Our group develops and applies computational approaches to study molecular variations and their phenotypic consequence. We are part of DKFZ and EMBL.
Website: https://steglelab.org/






















