Learn about de novo antibody sequencing by mass spec using only one run at poster TP-005 today!
#asms2025 #proteomics
@hecklab.bsky.social @tshamorkina.bsky.social @tkadava.bsky.social @joostsnijder.bsky.social
03.06.2025 15:45 — 👍 5 🔁 0 💬 0 📌 0
Thanks to @cinderbio.bsky.social and @sciex.bsky.social for their contribution and support
30.05.2025 00:49 — 👍 2 🔁 1 💬 0 📌 0
We are excited to have it out: a combination of archaeal proteases and hybrid fragmentation for antibody de novo sequencing. Poster TP-005 #asms2025 #teammassspec
It was a true team effort @lperezpaneda.bsky.social @tkadava.bsky.social @joostsnijder.bsky.social and @hecklab.bsky.social
30.05.2025 00:48 — 👍 16 🔁 8 💬 2 📌 0

Just over a week to go until #ASMS2025 in Baltimore
Come see posters using CinderBio enzymes on the schedule and topics below.
Steven Yannone our Founder and CEO, will be at the conference. Reach out to info@cinderbio.com for more information.
#proteomics #TeamMassSpec
23.05.2025 10:00 — 👍 3 🔁 0 💬 0 📌 0

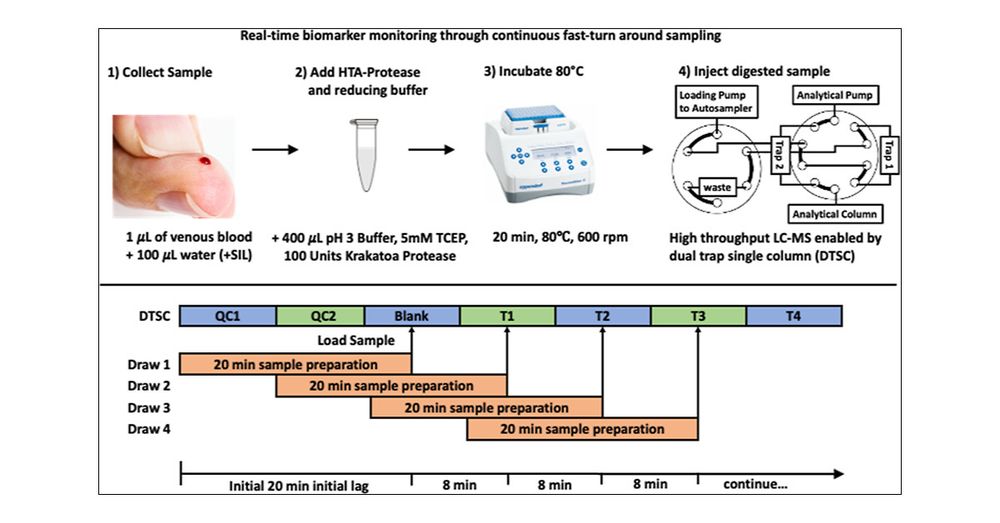
Toward Real-Time Proteomics: Blood to Biomarker Quantitation in under One Hour pubs.acs.org/doi/abs...
---
#proteomics #prot-paper
24.03.2025 09:40 — 👍 4 🔁 1 💬 1 📌 0

At US HUPO, CinderBio technologies were featured in a platform presentation and two posters. Researchers highlighted how the simple sample preparation workflows led to complete and deep sequence reads for mAbs and de novo sequencing for new proteins.
cinderbio.com
#Proteomics #TeamMassSpec
20.03.2025 15:30 — 👍 1 🔁 0 💬 0 📌 0
Thank you!
26.01.2024 01:08 — 👍 0 🔁 0 💬 0 📌 0
@ucdproteomics.bsky.social , @benneely.com - any plans to restart radio hour?
24.01.2024 08:37 — 👍 2 🔁 0 💬 2 📌 0

Create a christmas carol talking about mass spectrometers and proteomics
Created with Bard.
Someone wants to go to a baseball game? Not sure I get the reference. Anyway, here is our ai-enabled contribution to the Christmas carols - enjoy! g.co/bard/share/b...
12.12.2023 15:05 — 👍 2 🔁 0 💬 1 📌 0
@ucdproteomics.bsky.social, @benneely.com - radio hour this week or no?
12.12.2023 02:18 — 👍 0 🔁 0 💬 1 📌 0
We want to give a public "Thank You!” to early adopters of CinderBio's initial HTA-Proteases. We are thrilled to contribute to experiments in clinical proteomics, epigenetics, immunotherapies, cancer drug discovery and more. We enjoy being this kind of busy. Thanks to all of you - Team CinderBio
14.11.2023 10:45 — 👍 0 🔁 0 💬 0 📌 0
@hartl.bsky.social do your numbers match on the $ side as well? If I understood correctly, Michael is saying ~$500 in labor per sample (including lab-performed data analysis) and ~$50 in depreciation and consumables. How do you think the labor breaks out in terms of activities?
13.11.2023 12:04 — 👍 0 🔁 0 💬 0 📌 0
Thanks, next step to address @cashwood.bsky.social original question is how much labor does it take to run 4000 samples/year or roughly 16 samples a wkg day. To make things simple let's assume fractional people are a possibility
12.11.2023 16:50 — 👍 0 🔁 0 💬 1 📌 0
How much of that $50 is depreciation which is dependent on the amount of activity on the LC and MS?
11.11.2023 12:26 — 👍 0 🔁 0 💬 1 📌 0
What’s a reasonable estimate for labor required to do everything
11.11.2023 01:38 — 👍 1 🔁 0 💬 1 📌 0
@lkpino.bsky.social , @scienthusiast.bsky.social Why do you think mass spec has underdelivered? What was missing then and is it here now or at least on the horizon? If not, who is working on it?
25.10.2023 12:20 — 👍 0 🔁 0 💬 1 📌 0

VENNY VENN IS DOWN!
Okay. I'm not going to panic, but VENNY doesn't work. If you aren't familiar, the VENNY website by Juan Carlos Oliveros (2007) was the fir...
Venny has moved! After reading this blog post by @proteomicsnews.bsky.social I did a little digging and found this page: bioinfogp.cnb.csic.es where it says the site is down for maintenance. However, Venny is temporarily available here csbg.cnb.csic.es/BioinfoGP/ve... Happy Venning!
#proteomics
23.10.2023 11:41 — 👍 6 🔁 1 💬 3 📌 1
@ucdproteomics.bsky.social can you clarify the time? Eastern and Pacific seemed to be switched above and they are only 3 hours apart - not 4.
18.10.2023 22:37 — 👍 1 🔁 0 💬 1 📌 0
(2/2)
- Olink’s proven and transformative innovation is highly complementary to our leading mass spectrometry and life sciences platforms.
- [Olink will] meaningfully accelerate discovery and scientific breakthroughs
17.10.2023 13:45 — 👍 0 🔁 0 💬 0 📌 0
It’s a lot for sure. ~15x projected 2024 revenue for something expected to grow revenue in the “mid-teens” in 2025/6.
From the press release on the rationale:
- proteomics is having a profound impact on life science research and precision medicine.
1/2
17.10.2023 13:43 — 👍 1 🔁 0 💬 1 📌 0
Hi Tonya, HTA Proteases get reasonable coverage without reduction. With reduction, coverage improves. IAA is not required at pH3. Happy to answer other questions. Please contact us on the website. cinderbio.com
Our enzymes are strange and fun!
#proteomics #massspec #archaea
05.10.2023 22:56 — 👍 1 🔁 0 💬 0 📌 0
Tell us about yourself….
01.10.2023 17:01 — 👍 1 🔁 0 💬 0 📌 0
Mass Spec|Proteomics|Chemical Biology
Proteins - mainly intact, getting into oligos recently…
Application Scientist | LC-MS | Pharma BioPharma
#TeamMassSpec
Views are my own
I'm a biochemist in the field of mass spectrometry-based proteomics since 2012.
India -> Australia -> UK
Scientist working in the field of clinical proteomics.
https://scholar.google.com/citations?user=jQ_KiOEAAAAJ&hl=en
Proteomics, precision health, mass spectrometry, biomarkers, quantitative, changing medicine
PhD Student in Biochemistry @ UW-Madison
focus on plasma proteomics technologies and applications
website: https://wbeimers.github.io/
Microbial geneticist interested in large and small DNA 🧬 rings in bacteria and archaea.
Living in Brontë country and based at the University of Old York.
https://barillalabyork.weebly.com/
Post doc at the University of Copenhagen in Jesper Olsen’s group
Working on proteomics, phosphoproteomics, palaeoproteomics 🧫🔬🦷
Assoc. prof. at LUMC/@unileiden. Cdiff, DNA replication, AMR, microbiome, anaerobes. Clostpath Steering committee. Education officer ESGCD. Previously: @unigroningen and @mitofficial. https://sites.google.com/view/expbac-lumc/homepage
PI and Lecturer, University of Manchester
https://dannagifford.com @MERManchester.bsky.social
antibodies, viruses, cryo-EM -- habitually curious -- BMB graduate student at Caltech
Doing proteomics until I still can.
https://github.com/41ison
Interested in mass spectrometry, -omics and tasting the best food in every place I visit!
I help SCIEX’s accurate mass customers in the Nordics, UK & Ireland
Chairperson for @lpdg.bsky.social and help the LBMSDG meetings in London
Proteomics 🩸 Mass Spectrometry ✨ Structural Biology 🧬 Network Biology 🕸️ Bioinformatics 💻 Canada 🍁 Columbia University 🗽 @ionsbyfriday 𝕏
Bioinformatics Solutions Inc. is a world leading software provider for mass spectrometry-based proteomics.
https://www.bioinfor.com
🇲🇽🇫🇷🇪🇺 Analytical chemist, specialized in #MassSpec #Proteomics. Leading the Immunopeptidomics Platform at HI-TRON Mainz / DKFZ
Researcher @cea_grenoble 🇫🇷 — dealing with Light, Protein structures, and Mass Spectra - MSCA alumn - Previously 🇳🇱🇮🇹@UniUtrecht@PoliTOnews@UniPadova
PhD student at HeckLab, Utrecht University.
Using mass spectrometry to characterize protein complexes.
https://linktr.ee/TKadava